

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146721.7 - phase: 0
(188 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
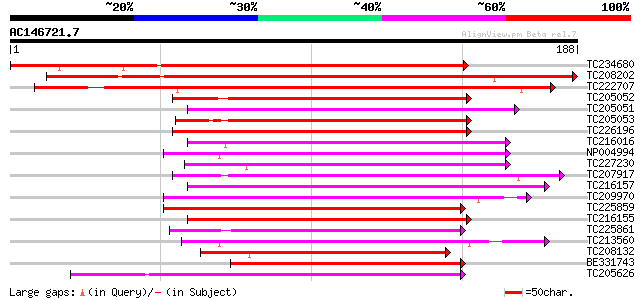
Score E
Sequences producing significant alignments: (bits) Value
TC234680 similar to UP|Q9AT29 (Q9AT29) BZip transcription factor... 238 1e-63
TC208202 similar to UP|Q9AT29 (Q9AT29) BZip transcription factor... 228 1e-60
TC222707 homologue to UP|Q9AT29 (Q9AT29) BZip transcription fact... 175 1e-44
TC205052 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor... 90 5e-19
TC205051 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor... 84 5e-17
TC205053 homologue to UP|Q8L5W2 (Q8L5W2) BZIP transcription fact... 81 2e-16
TC226196 UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2, comp... 80 6e-16
TC216016 similar to UP|P93839 (P93839) G/HBF-1, partial (61%) 74 3e-14
NP004994 G/HBF-1 73 9e-14
TC227230 similar to UP|P93839 (P93839) G/HBF-1, partial (72%) 71 3e-13
TC207917 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor... 71 3e-13
TC216157 weakly similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription... 70 4e-13
TC209970 weakly similar to UP|Q8GTZ1 (Q8GTZ1) BZIP transcription... 68 3e-12
TC225859 similar to UP|O22677 (O22677) BZIP DNA-binding protein,... 68 3e-12
TC216155 weakly similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription... 67 6e-12
TC225861 similar to UP|Q941L6 (Q941L6) BZIP transcription factor... 66 8e-12
TC213560 similar to UP|Q9FUD3 (Q9FUD3) BZIP protein BZO2H2, part... 66 1e-11
TC208132 homologue to UP|Q94KA7 (Q94KA7) BZIP transcription fact... 63 9e-11
BE331743 similar to GP|24460973|gb| bZIP transcription factor {C... 62 1e-10
TC205626 similar to UP|Q9FWS7 (Q9FWS7) F1B16.8 protein (At1g7539... 60 5e-10
>TC234680 similar to UP|Q9AT29 (Q9AT29) BZip transcription factor, partial
(49%)
Length = 571
Score = 238 bits (607), Expect = 1e-63
Identities = 122/157 (77%), Positives = 136/157 (85%), Gaps = 5/157 (3%)
Frame = +2
Query: 1 MLPSNTSPYLQNYNN---MIQNNNISSYQLQKFSNQIYG--NYLNTPHQQFPDFNYSPQS 55
+LPSN PY N NN M QNN ++Q Q+FSNQIYG N NTP+ + PD +SPQS
Sbjct: 104 LLPSNPCPYPPNNNNYTTMFQNNINPTFQFQRFSNQIYGYNNINNTPYHKVPDL-FSPQS 280
Query: 56 SCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNEN 115
SCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQV+WLRNEN
Sbjct: 281 SCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVVWLRNEN 460
Query: 116 HQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKD 152
HQL++KLNHVSE+HDQV+QENAQLKE+ALELRQMI+D
Sbjct: 461 HQLMDKLNHVSESHDQVMQENAQLKEQALELRQMIRD 571
>TC208202 similar to UP|Q9AT29 (Q9AT29) BZip transcription factor, partial
(51%)
Length = 758
Score = 228 bits (582), Expect = 1e-60
Identities = 121/177 (68%), Positives = 138/177 (77%), Gaps = 1/177 (0%)
Frame = +1
Query: 13 YNNMIQNNNISSYQLQKFSNQIYGNYLNTPHQQFPDFNYSPQSSCISSNSTSDEADEQNL 72
+N + N I ++QL K SNQ YG P Q DF+ PQSSCISSNSTSDEADEQ
Sbjct: 43 FNYSMVQNTIPTFQLHKLSNQFYG-LQKPPPQVLADFS-PPQSSCISSNSTSDEADEQQQ 216
Query: 73 SLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQV 132
SLINERKHRRMISNRESARRSRMRKQKHLDELWSQV+WLRNENHQL++KLNHVSE+HD+V
Sbjct: 217 SLINERKHRRMISNRESARRSRMRKQKHLDELWSQVVWLRNENHQLMDKLNHVSESHDKV 396
Query: 133 VQENAQLKEEALELRQMIKDMQIHSPL-IPSFSPLDDTYLRDDSSNNSISSSMDLLG 188
QEN QL+EEA ELRQMI DMQ+HSP P SP+DD S++SI+ S+DLLG
Sbjct: 397 AQENVQLREEASELRQMICDMQLHSPYHPPPLSPIDDDVSPYVKSDSSITDSLDLLG 567
>TC222707 homologue to UP|Q9AT29 (Q9AT29) BZip transcription factor, complete
Length = 671
Score = 175 bits (444), Expect = 1e-44
Identities = 101/186 (54%), Positives = 124/186 (66%), Gaps = 13/186 (6%)
Frame = +3
Query: 9 YLQNYNNMIQNNNISSYQLQKFSNQIYGNYLNTPHQQFPDFNYSPQ--------SSCISS 60
YL N + N S Q + I +LNT FP+ ++ P SSC+SS
Sbjct: 102 YLAPENPFLVPPNFSLLQ-----SDIPNLHLNTLLSNFPNCHFPPSGHEFVVPPSSCLSS 266
Query: 61 NSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIE 120
NSTSDEADE ++I+ERKHRRMISNRESARRSRMRKQKHLDELWSQV+ LR ENH LI+
Sbjct: 267 NSTSDEADEIQFNIIDERKHRRMISNRESARRSRMRKQKHLDELWSQVVRLRTENHNLID 446
Query: 121 KLNHVSENHDQVVQENAQLKEEALELRQMIKDMQIHSPLIPSFSPLDD-----TYLRDDS 175
KLNHVSE+HD+V+QENA+LKEEA LRQM+ DMQI + + L+D + L+ D
Sbjct: 447 KLNHVSESHDRVLQENARLKEEASALRQMLADMQIGTAFACTMEDLEDLPCNTSQLKPDP 626
Query: 176 SNNSIS 181
N SI+
Sbjct: 627 LNQSIT 644
>TC205052 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2,
partial (58%)
Length = 953
Score = 90.1 bits (222), Expect = 5e-19
Identities = 48/99 (48%), Positives = 69/99 (69%)
Frame = +1
Query: 55 SSCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNE 114
SS + +NS S+E + +L+ +RK +RMISNRESARRSRMRKQKHLD+L SQV LRNE
Sbjct: 541 SSMLQNNSGSEEELQ---ALMEQRKRKRMISNRESARRSRMRKQKHLDDLASQVTQLRNE 711
Query: 115 NHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDM 153
NHQ++ +N ++ + V EN+ L+ + EL ++ +
Sbjct: 712 NHQILTSVNLTTQKYLAVEAENSVLRAQVNELSHWLESL 828
>TC205051 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2,
partial (63%)
Length = 773
Score = 83.6 bits (205), Expect = 5e-17
Identities = 46/110 (41%), Positives = 66/110 (59%)
Frame = +1
Query: 60 SNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLI 119
S+ TS + ++++RK +RMISNRESARRSRMRKQKHLD+L SQ+ LR++N QL+
Sbjct: 406 SSGTSSGTSSELQGMMDQRKRKRMISNRESARRSRMRKQKHLDDLASQLTQLRSQNQQLL 585
Query: 120 EKLNHVSENHDQVVQENAQLKEEALELRQMIKDMQIHSPLIPSFSPLDDT 169
+N S + V EN+ L+ + EL + + L+ F P T
Sbjct: 586 TSVNLTSHKYLAVEAENSVLRAQVNELSHRLDSLNQIIHLLNFFKPNTST 735
>TC205053 homologue to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2,
partial (89%)
Length = 1432
Score = 81.3 bits (199), Expect = 2e-16
Identities = 46/98 (46%), Positives = 67/98 (67%)
Frame = +2
Query: 56 SCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNEN 115
S + NS S+E D Q +++++RK +RMISNRESARRSRMRKQKHLD+L SQV LR EN
Sbjct: 746 SSLLQNSGSEE-DLQ--AVMDQRKRKRMISNRESARRSRMRKQKHLDDLVSQVAQLRKEN 916
Query: 116 HQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDM 153
Q++ +N ++ + V EN+ L+ + EL ++ +
Sbjct: 917 QQILTSVNITTQQYLSVEAENSVLRAQVGELSHRLESL 1030
>TC226196 UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2, complete
Length = 1518
Score = 80.1 bits (196), Expect = 6e-16
Identities = 42/99 (42%), Positives = 62/99 (62%)
Frame = +2
Query: 55 SSCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNE 114
S +S S ++ + ++RK +RMISNRESARRSRMRKQKHLD+L SQV LR E
Sbjct: 761 SGSLSLLQNSGSEEDLQAMMEDQRKRKRMISNRESARRSRMRKQKHLDDLVSQVAQLRKE 940
Query: 115 NHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDM 153
N Q++ +N ++ + V EN+ L+ + EL ++ +
Sbjct: 941 NQQILTSVNITTQQYLSVEAENSVLRAQVGELSHRLESL 1057
>TC216016 similar to UP|P93839 (P93839) G/HBF-1, partial (61%)
Length = 1764
Score = 74.3 bits (181), Expect = 3e-14
Identities = 42/117 (35%), Positives = 67/117 (56%), Gaps = 10/117 (8%)
Frame = +3
Query: 60 SNSTSDEADEQ----------NLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVL 109
S S+ +++D++ N++ ++ ++ RRM+SNRESARRSR RKQ HL EL +QV
Sbjct: 651 SGSSGEQSDDEEAEGEINMTGNMTPVDAKRVRRMLSNRESARRSRRRKQAHLTELETQVS 830
Query: 110 WLRNENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDMQIHSPLIPSFSPL 166
LR+EN L+++ VS+ + +N LK + LR +K + I SP+
Sbjct: 831 QLRSENSSLLKRFTDVSQKYSNAAVDNRVLKADVETLRAKVKMAEETVKRITGLSPM 1001
>NP004994 G/HBF-1
Length = 1137
Score = 72.8 bits (177), Expect = 9e-14
Identities = 44/120 (36%), Positives = 65/120 (53%), Gaps = 5/120 (4%)
Frame = +1
Query: 52 SPQSSCISSNSTSDEAD-----EQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWS 106
S S S DEA+ Q + + ++ RRM+SNRESARRSR RKQ HL EL +
Sbjct: 430 STTSGSSREQSDDDEAEGEAETTQGMDPADAKRVRRMLSNRESARRSRRRKQAHLTELET 609
Query: 107 QVLWLRNENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDMQIHSPLIPSFSPL 166
QV LR EN L+++L +S+ +++ +N LK + LR +K + + +PL
Sbjct: 610 QVSQLRVENSSLLKRLTDISQKYNEAAVDNRVLKADVETLRTKVKMAEETVKRVTGLNPL 789
>TC227230 similar to UP|P93839 (P93839) G/HBF-1, partial (72%)
Length = 1250
Score = 71.2 bits (173), Expect = 3e-13
Identities = 43/115 (37%), Positives = 64/115 (55%), Gaps = 7/115 (6%)
Frame = +2
Query: 59 SSNSTSDEADEQNLSLINE-------RKHRRMISNRESARRSRMRKQKHLDELWSQVLWL 111
SS SD+ D + + +N+ ++ RRM+SNRESARRSR RKQ HL +L +QV L
Sbjct: 425 SSREQSDDEDIEGETSMNDNTDPADVKRVRRMLSNRESARRSRRRKQAHLTDLETQVSQL 604
Query: 112 RNENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDMQIHSPLIPSFSPL 166
R EN L+++L VS+ + +N LK + LR +K + I +P+
Sbjct: 605 RGENSTLLKRLTDVSQKYSDSAVDNRVLKADVETLRAKVKMAEETVKRITGLNPM 769
>TC207917 similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor ATB2,
partial (75%)
Length = 869
Score = 70.9 bits (172), Expect = 3e-13
Identities = 48/131 (36%), Positives = 78/131 (58%), Gaps = 1/131 (0%)
Frame = +2
Query: 55 SSCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNE 114
S+ I S+ E D Q L+ +RK +R SNRESARRSRMRKQKHLD+L +QV L+ +
Sbjct: 14 STSILLQSSGSEEDLQ--LLMEQRKKKRKQSNRESARRSRMRKQKHLDDLIAQVDLLKKQ 187
Query: 115 NHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDMQIHSPLIPSFSPLD-DTYLRD 173
+K+N +++ +V EN+ L + EL Q ++ + LI + + D + Y +
Sbjct: 188 KSLSFKKVNITTQHCLKVEAENSILGAQKTELTQSLQSLNDIINLINTTTTSDHNNYYYN 367
Query: 174 DSSNNSISSSM 184
+++NN+ ++ M
Sbjct: 368 NNNNNNNNNFM 400
>TC216157 weakly similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor
ATB2, partial (68%)
Length = 875
Score = 70.5 bits (171), Expect = 4e-13
Identities = 40/120 (33%), Positives = 70/120 (58%)
Frame = +2
Query: 60 SNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLI 119
S+S + E + ++ +RK +RM+SNRESARRSRMRKQ+HL+ L +Q+ L+ EN Q+
Sbjct: 128 SSSLQNSGSEGDRDIMEQRKRKRMLSNRESARRSRMRKQQHLEGLSAQLDQLKKENTQMN 307
Query: 120 EKLNHVSENHDQVVQENAQLKEEALELRQMIKDMQIHSPLIPSFSPLDDTYLRDDSSNNS 179
+ ++ + V ENA L+ + EL + + + LI S + ++ + D++ +
Sbjct: 308 TNIGISTQLYLNVEAENAILRAQMEELSKRLNSLNEMISLINSTTTTNNCLMFDEAQETT 487
>TC209970 weakly similar to UP|Q8GTZ1 (Q8GTZ1) BZIP transcription factor,
partial (62%)
Length = 1041
Score = 67.8 bits (164), Expect = 3e-12
Identities = 41/124 (33%), Positives = 72/124 (58%), Gaps = 2/124 (1%)
Frame = +2
Query: 52 SPQSSCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWL 111
S SS + + +S + + + +ERK++R SNRESARRSRMRK+ HLD+L Q+ L
Sbjct: 266 SGNSSISTKSQSSGSEGDLQVVITDERKNKRKQSNRESARRSRMRKRNHLDQLTKQLSQL 445
Query: 112 RNENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIKDMQ--IHSPLIPSFSPLDDT 169
N +++ ++ ++++ V EN+ L+ + EL Q ++ + +H +I + + T
Sbjct: 446 AKNNGEILATIDITTQHYLNVEAENSILRAQMGELSQRLQSLNDIVHDIIINTTT----T 613
Query: 170 YLRD 173
Y RD
Sbjct: 614 YERD 625
>TC225859 similar to UP|O22677 (O22677) BZIP DNA-binding protein, partial
(59%)
Length = 1132
Score = 67.8 bits (164), Expect = 3e-12
Identities = 38/100 (38%), Positives = 60/100 (60%)
Frame = +1
Query: 52 SPQSSCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWL 111
SP S +TS ++ + +I+ERK +RM+SNRESARRSRMRKQK L++L +V L
Sbjct: 424 SPIQQQQRSTTTSSGSEGGDPHIIDERKRKRMLSNRESARRSRMRKQKQLEDLTDEVSRL 603
Query: 112 RNENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIK 151
++ N +L E + E + N+ L+ + +EL ++
Sbjct: 604 QSANKKLAENIEAKEEACVETEAANSILRAQTMELADRLR 723
>TC216155 weakly similar to UP|Q8L5W2 (Q8L5W2) BZIP transcription factor
ATB2, partial (66%)
Length = 1264
Score = 66.6 bits (161), Expect = 6e-12
Identities = 35/94 (37%), Positives = 59/94 (62%)
Frame = +1
Query: 60 SNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLI 119
S+S + E + ++ +RK +RM+SNRESARRSR+RKQ+HL+ L +Q+ L+ EN Q+
Sbjct: 622 SSSLQNSGSEGDRDIMEQRKRKRMLSNRESARRSRIRKQQHLEGLSAQLDQLKKENAQIN 801
Query: 120 EKLNHVSENHDQVVQENAQLKEEALELRQMIKDM 153
++ ++ + V ENA L+ + EL + +
Sbjct: 802 TNISITTQMYLNVEAENAILRAQMGELSNRLNSL 903
>TC225861 similar to UP|Q941L6 (Q941L6) BZIP transcription factor BZI-3,
partial (43%)
Length = 1239
Score = 66.2 bits (160), Expect = 8e-12
Identities = 38/98 (38%), Positives = 58/98 (58%)
Frame = +3
Query: 54 QSSCISSNSTSDEADEQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRN 113
Q S +S+ + D Q +I+ERK +RM+SNRESARRSRMRKQK L++L +V L+
Sbjct: 459 QRSTATSSGSEGGGDPQ---MIDERKRKRMLSNRESARRSRMRKQKQLEDLTDEVSRLQG 629
Query: 114 ENHQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIK 151
N +L E + E + N+ L+ + +EL ++
Sbjct: 630 ANKKLAENIKAKEEACVETEAANSILRAQTMELADRLR 743
>TC213560 similar to UP|Q9FUD3 (Q9FUD3) BZIP protein BZO2H2, partial (44%)
Length = 794
Score = 65.9 bits (159), Expect = 1e-11
Identities = 48/127 (37%), Positives = 68/127 (52%), Gaps = 5/127 (3%)
Frame = +2
Query: 58 ISSNSTSDEAD--EQNLSLINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNEN 115
+S DEA EQ+ + I+ ++ RR +SNRESARRSR RKQ HL +L QV LR EN
Sbjct: 380 VSHLMRDDEAGPCEQSTNAIDMKRLRRKVSNRESARRSRRRKQAHLADLEWQVERLRLEN 559
Query: 116 HQLIEKLNHVSENHDQVVQENAQLKEEALELRQMIK---DMQIHSPLIPSFSPLDDTYLR 172
L ++L S+ N LK + LR +K DM L +P+++ L+
Sbjct: 560 ANLFKQLTDASQQFRDADTNNRVLKSDVEALRAKVKLAEDMVTRGTL----TPINNQILQ 727
Query: 173 DDSSNNS 179
+ SS N+
Sbjct: 728 NQSSLNT 748
>TC208132 homologue to UP|Q94KA7 (Q94KA7) BZIP transcription factor 3,
partial (33%)
Length = 706
Score = 62.8 bits (151), Expect = 9e-11
Identities = 37/86 (43%), Positives = 53/86 (61%), Gaps = 3/86 (3%)
Frame = +1
Query: 64 SDEADEQNLSLINER---KHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIE 120
S ++ + L L +ER + RR SNRESARRSR+RKQ DEL + L+ EN L
Sbjct: 133 SRDSAQSQLWLQDERELKRQRRKQSNRESARRSRLRKQAECDELAQRAEALKEENASLRS 312
Query: 121 KLNHVSENHDQVVQENAQLKEEALEL 146
++N + +++Q++ ENA LKE EL
Sbjct: 313 EVNRIRSDYEQLLSENAALKERLGEL 390
>BE331743 similar to GP|24460973|gb| bZIP transcription factor {Capsicum
chinense}, partial (35%)
Length = 292
Score = 62.4 bits (150), Expect = 1e-10
Identities = 32/78 (41%), Positives = 49/78 (62%)
Frame = +2
Query: 74 LINERKHRRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVV 133
+I+ERK +RM+SNRESARRSRMRKQK L++L +V L+ N L E + + +
Sbjct: 26 MIDERKRKRMLSNRESARRSRMRKQKQLEDLTDEVSRLQGANKMLAENIKAKEDACVETE 205
Query: 134 QENAQLKEEALELRQMIK 151
N+ L+ + +EL ++
Sbjct: 206 AANSILRAQTMELADRLR 259
>TC205626 similar to UP|Q9FWS7 (Q9FWS7) F1B16.8 protein (At1g75390), partial
(34%)
Length = 1184
Score = 60.5 bits (145), Expect = 5e-10
Identities = 40/131 (30%), Positives = 69/131 (52%)
Frame = +3
Query: 21 NISSYQLQKFSNQIYGNYLNTPHQQFPDFNYSPQSSCISSNSTSDEADEQNLSLINERKH 80
N S++ + NQI ++ Q+ P +P +S S+ + + ++ERK
Sbjct: 315 NSSNFPIFFLINQICKILISLS*QK-PQIQTNPMASIPRPASSGSSEGGGDPAAMDERKR 491
Query: 81 RRMISNRESARRSRMRKQKHLDELWSQVLWLRNENHQLIEKLNHVSENHDQVVQENAQLK 140
+RM SNRESARRSRM+KQK L++L L+ EN +L + + E + ++ N L+
Sbjct: 492 KRMESNRESARRSRMKKQKLLEDLSDVASRLQGENVRLAQSIKAKEEAYVEIEAANDILR 671
Query: 141 EEALELRQMIK 151
+ +EL ++
Sbjct: 672 AQTMELADRLR 704
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.126 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 8,677,082
Number of Sequences: 63676
Number of extensions: 116987
Number of successful extensions: 754
Number of sequences better than 10.0: 107
Number of HSP's better than 10.0 without gapping: 732
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 747
length of query: 188
length of database: 12,639,632
effective HSP length: 92
effective length of query: 96
effective length of database: 6,781,440
effective search space: 651018240
effective search space used: 651018240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146721.7


