

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
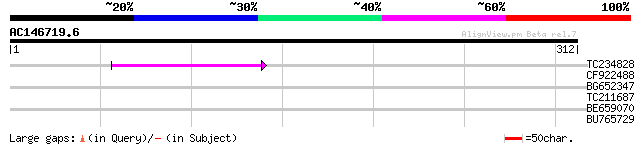
Query= AC146719.6 - phase: 0 /pseudo
(312 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC234828 46 2e-05
CF922488 29 2.4
BG652347 28 5.3
TC211687 similar to UP|Q7VNN2 (Q7VNN2) Conserved probable membra... 28 7.0
BE659070 homologue to SP|O17449|TBB1 Tubulin beta-1 chain (Beta-... 27 9.1
BU765729 27 9.1
>TC234828
Length = 857
Score = 46.2 bits (108), Expect = 2e-05
Identities = 34/85 (40%), Positives = 40/85 (47%)
Frame = +1
Query: 57 CLLRIKQ*LISYRLYTIFQKFFLMKFPMCLQREKLNF*LILFLERSQCL*HLIVCQLLS* 116
C+ + L+ YR FQKFF M F C REK N L L R QC LI C +*
Sbjct: 598 CMWKRTLKLVVYRWCQNFQKFFRMMFVNCHLREKWNSLLTWCLGRIQCRLRLIECLRWN* 777
Query: 117 LS*RNN*RICFRKSLLEQVFHLGER 141
*R+ R +L Q H GER
Sbjct: 778 QR*RHKYRTF*ANNLFVQAHHRGER 852
>CF922488
Length = 741
Score = 29.3 bits (64), Expect = 2.4
Identities = 25/78 (32%), Positives = 41/78 (52%)
Frame = +1
Query: 148 RKMVA*DCALIIDS*TK*RERIGIHFQE*MT*WISWWVHVCSARYI*DRVITRLK*KMKI 207
++M +CA I+ *TK +RI ++ WI+ V S ++ +VITR +* +I
Sbjct: 10 KRMGRCECAWTIEI*TKPVQRISFLYRISTFSWITRPVFPNSPSWMDFQVITR*R*HQRI 189
Query: 208 CRRQLLERGMHIMSIKLC 225
+RQL +I+LC
Sbjct: 190 WKRQLSLLYGEPSAIRLC 243
>BG652347
Length = 356
Score = 28.1 bits (61), Expect = 5.3
Identities = 13/48 (27%), Positives = 22/48 (45%), Gaps = 2/48 (4%)
Frame = +1
Query: 214 ERGMHIMSIKLCLSVLQMHLVCLWST*IA--FFMHFWTVSWLFLLMTF 259
E M + + KLC + H +C+ + + H+W W FL + F
Sbjct: 7 ETSMGLSNQKLCSDSKRSHTICICCMLSS*TIYNHYWL*GWFFLHLAF 150
>TC211687 similar to UP|Q7VNN2 (Q7VNN2) Conserved probable membrane protein,
partial (5%)
Length = 632
Score = 27.7 bits (60), Expect = 7.0
Identities = 14/36 (38%), Positives = 20/36 (54%), Gaps = 3/36 (8%)
Frame = -1
Query: 50 YFL*WHPCLLRIKQ*LISYRLYTIFQK---FFLMKF 82
YF+ W C+L L+S+ LY++F FFL F
Sbjct: 410 YFMLWVRCILCASLCLLSFSLYSVFLHCACFFLFSF 303
>BE659070 homologue to SP|O17449|TBB1 Tubulin beta-1 chain (Beta-1 tubulin).
[Tobacco hawkmoth Tobacco hornworm] {Manduca sexta},
partial (30%)
Length = 705
Score = 27.3 bits (59), Expect = 9.1
Identities = 11/20 (55%), Positives = 15/20 (75%)
Frame = +1
Query: 275 WFCKCSKRRSFMLNYLSVNS 294
W+C S+ +SFML+ SVNS
Sbjct: 346 WYCWYSETKSFMLDSASVNS 405
>BU765729
Length = 425
Score = 27.3 bits (59), Expect = 9.1
Identities = 14/44 (31%), Positives = 22/44 (49%), Gaps = 4/44 (9%)
Frame = +2
Query: 225 CLSVLQMHLVCLWS----T*IAFFMHFWTVSWLFLLMTF*FTPR 264
CL ++ CL+ T + FF SW+FLL+ F ++ R
Sbjct: 134 CLRIMNNTCSCLYQRSCFTGVFFFFFLQIFSWVFLLLVFMWSER 265
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.378 0.167 0.656
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,894,817
Number of Sequences: 63676
Number of extensions: 257442
Number of successful extensions: 5227
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 2920
Number of HSP's successfully gapped in prelim test: 197
Number of HSP's that attempted gapping in prelim test: 2226
Number of HSP's gapped (non-prelim): 3252
length of query: 312
length of database: 12,639,632
effective HSP length: 97
effective length of query: 215
effective length of database: 6,463,060
effective search space: 1389557900
effective search space used: 1389557900
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146719.6


