

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146719.12 + phase: 0 /pseudo
(758 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
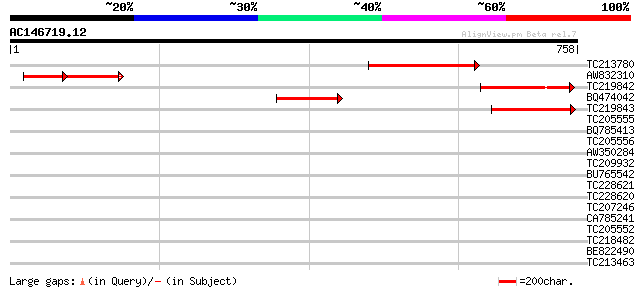
Score E
Sequences producing significant alignments: (bits) Value
TC213780 178 7e-45
AW832310 weakly similar to GP|20804590|db P0421H07.9 {Oryza sati... 100 7e-41
TC219842 similar to UP|Q94BY0 (Q94BY0) AT3g49400/F2K15_260 (Frag... 136 4e-32
BQ474042 weakly similar to GP|14517468|gb| AT3g49400/F2K15_260 {... 132 4e-31
TC219843 similar to UP|Q94BY0 (Q94BY0) AT3g49400/F2K15_260 (Frag... 124 1e-28
TC205555 36 0.073
BQ785413 33 0.47
TC205556 33 0.61
AW350284 similar to GP|22946224|gb| CG31868-PA {Drosophila melan... 32 1.4
TC209932 32 1.4
BU765542 31 1.8
TC228621 similar to UP|Q8RXD8 (Q8RXD8) WD repeat protein-like, p... 31 1.8
TC228620 weakly similar to UP|Q8RXD8 (Q8RXD8) WD repeat protein-... 31 1.8
TC207246 similar to UP|Q6YWF4 (Q6YWF4) Rhomboid family protein-l... 31 2.3
CA785241 31 2.3
TC205552 similar to UP|PMM_ARATH (O80840) Probable phosphomannom... 30 4.0
TC218482 30 5.2
BE822490 SP|P43565|RI1 Serine/threonine-protein kinase RIM15 (EC... 29 6.8
TC213463 similar to UP|Q8H3D7 (Q8H3D7) Leaf senescence related p... 29 8.9
>TC213780
Length = 447
Score = 178 bits (452), Expect = 7e-45
Identities = 102/148 (68%), Positives = 110/148 (73%)
Frame = +1
Query: 480 QEIHLNHVLA*LCPLEILSLQLFIALILIN*IECMKEGF*GQPLNTSGLVGYKLMFC*NL 539
QE HL HVL CPLEILSL LFI L+L N*IECMKEGF*GQ LNT GLV YK MF *NL
Sbjct: 4 QERHLIHVLVLQCPLEILSLLLFIVLMLRN*IECMKEGF*GQLLNTFGLVDYKWMFS*NL 183
Query: 540 PFHATLKNFPLFQRKN*LIGGPISYGL*TIINVSISLWFFGI*LQHYWLSRKTSQNM*NI 599
PFH TLK P+ +KN*LIG PISYGL*T INV ISLWF GI*LQHY SR T NM NI
Sbjct: 184 PFHVTLKEIPVMWKKN*LIGEPISYGL*TNINVMISLWFSGI*LQHYQHSRITI*NMQNI 363
Query: 600 *WSNGSHYHFWDLI*IFLLRKCYHVSFQ 627
+GSH+ F++ I FLL+K YH S Q
Sbjct: 364 YLLSGSHHLFYN*IWTFLLKKXYHSSLQ 447
>AW832310 weakly similar to GP|20804590|db P0421H07.9 {Oryza sativa (japonica
cultivar-group)}, partial (10%)
Length = 428
Score = 99.8 bits (247), Expect(2) = 7e-41
Identities = 54/82 (65%), Positives = 62/82 (74%)
Frame = +3
Query: 71 ACLQLCIGMINQLFDQYHGLHLAWLPTQGA*LLFAPLMDTLKFIVRPFVIFVLNGLRLWT 130
+C +LCIGMI+ L DQYHGL LAWL QGA* LFA L D +FIV FVI VLNGLR WT
Sbjct: 180 SCRRLCIGMISLLPDQYHGLRLAWLQIQGA**LFATLKDM*RFIVHLFVITVLNGLRWWT 359
Query: 131 *LKGCMNISDVLSFKAQEGVIH 152
*LK CM I +VLSF+ Q G++H
Sbjct: 360 *LKVCMTIFNVLSFEVQ-GLLH 422
Score = 87.0 bits (214), Expect(2) = 7e-41
Identities = 42/60 (70%), Positives = 49/60 (81%), Gaps = 1/60 (1%)
Frame = +2
Query: 19 FPNALAWSHDNLIAAASGHFVTILRPDLPNG-PRGLIKVIPNEPLILGFIDRKACLQLCI 77
FPNA+AWS DNLIA ASGH VTILRPDLP G PRG+IKV P+EPL +GF++RK L C+
Sbjct: 2 FPNAIAWSDDNLIAVASGHLVTILRPDLPIGAPRGVIKVSPSEPLPIGFVERKDLLSGCL 181
>TC219842 similar to UP|Q94BY0 (Q94BY0) AT3g49400/F2K15_260 (Fragment),
partial (3%)
Length = 773
Score = 136 bits (342), Expect = 4e-32
Identities = 83/126 (65%), Positives = 88/126 (68%)
Frame = +3
Query: 630 RTFLLDCCTCSI*FVDV*CWHSWMLIS*P**IANFKTWMGYVLSLKRK*LSG*KFF*AVK 689
+TFL T SI* VDV*C SWMLI * +FK+W VL K K LSG FF* VK
Sbjct: 9 QTFLXXYYTXSI*SVDV*CXXSWMLIK*LEYTKSFKSWKR*VLPWKNKLLSGQNFF*TVK 188
Query: 690 ESFVKGTWVLVSLLSKLACSIKKQLHPCLAVGILLD*HRWNNGLL*IRKTYMTS*NPLHQ 749
ES VKG W LV LLSKLAC I+KQ+ L VGILLD*HRWNNGLL*IR YMTS* LHQ
Sbjct: 189 ESSVKGLWALVFLLSKLACPIQKQISN-LVVGILLD*HRWNNGLL*IRNIYMTS*KSLHQ 365
Query: 750 KLLMRK 755
KL MRK
Sbjct: 366 KLSMRK 383
>BQ474042 weakly similar to GP|14517468|gb| AT3g49400/F2K15_260 {Arabidopsis
thaliana}, partial (5%)
Length = 434
Score = 132 bits (333), Expect = 4e-31
Identities = 68/88 (77%), Positives = 76/88 (86%)
Frame = +1
Query: 357 V*RFGWVTMTNCSNHQKLIQIPFYY*RRL*LQMLSQFQYFQ*LCMFSTPPRCF*P*AKFL 416
V*RFGW+TMTNC NHQK I++P +Y*RRL*L MLSQF YFQ* MF+T PRCF*P AKFL
Sbjct: 4 V*RFGWLTMTNC*NHQKWIKLPSHY*RRL*LSMLSQFLYFQ*PHMFNTLPRCF*PRAKFL 183
Query: 417 VPLRYGYVTYPLGNLISLAHMMHIIMLL 444
V L+YGYVTY LGNLISLAH+MH+I LL
Sbjct: 184 VHLKYGYVTYLLGNLISLAHIMHMIWLL 267
>TC219843 similar to UP|Q94BY0 (Q94BY0) AT3g49400/F2K15_260 (Fragment),
partial (4%)
Length = 758
Score = 124 bits (312), Expect = 1e-28
Identities = 73/112 (65%), Positives = 77/112 (68%)
Frame = +2
Query: 645 DV*CWHSWMLIS*P**IANFKTWMGYVLSLKRK*LSG*KFF*AVKESFVKGTWVLVSLLS 704
D *C SWMLI FK+W YV K K LSG KF AVKES VKG W V LLS
Sbjct: 2 DE*C*QSWMLIKXLEYTKTFKSWKRYVXPWKNKLLSGQKFXXAVKESSVKGLWXXVFLLS 181
Query: 705 KLACSIKKQLHPCLAVGILLD*HRWNNGLL*IRKTYMTS*NPLHQKLLMRKG 756
+LAC I+KQ+ P L VGILLD*HRWNNGLL*IR YMTS* PLHQKL M KG
Sbjct: 182 ELACPIQKQI-PNLVVGILLD*HRWNNGLL*IRDIYMTS*KPLHQKLSMSKG 334
>TC205555
Length = 552
Score = 35.8 bits (81), Expect = 0.073
Identities = 15/27 (55%), Positives = 17/27 (62%)
Frame = -1
Query: 165 TTTKKFPIRWGELHGSNEVIS*TMSVY 191
TTT +P RWG LHGS I T+S Y
Sbjct: 306 TTTTPYPTRWGRLHGSTSAIMMTLSPY 226
>BQ785413
Length = 376
Score = 33.1 bits (74), Expect = 0.47
Identities = 13/20 (65%), Positives = 14/20 (70%)
Frame = -1
Query: 165 TTTKKFPIRWGELHGSNEVI 184
TTT +P RWG LHGS VI
Sbjct: 103 TTTTPYPTRWGRLHGSTSVI 44
>TC205556
Length = 648
Score = 32.7 bits (73), Expect = 0.61
Identities = 13/23 (56%), Positives = 14/23 (60%)
Frame = -3
Query: 162 IKATTTKKFPIRWGELHGSNEVI 184
I TTT +P RWG LHGS I
Sbjct: 397 ITTTTTTPYPTRWGRLHGSTSAI 329
>AW350284 similar to GP|22946224|gb| CG31868-PA {Drosophila melanogaster},
partial (1%)
Length = 767
Score = 31.6 bits (70), Expect = 1.4
Identities = 12/20 (60%), Positives = 13/20 (65%)
Frame = +1
Query: 165 TTTKKFPIRWGELHGSNEVI 184
TTT +P RWG LHGS I
Sbjct: 61 TTTTPYPTRWGRLHGSTSAI 120
>TC209932
Length = 829
Score = 31.6 bits (70), Expect = 1.4
Identities = 12/20 (60%), Positives = 13/20 (65%)
Frame = -3
Query: 165 TTTKKFPIRWGELHGSNEVI 184
TTT +P RWG LHGS I
Sbjct: 599 TTTTPYPTRWGRLHGSTSAI 540
>BU765542
Length = 427
Score = 31.2 bits (69), Expect = 1.8
Identities = 18/42 (42%), Positives = 22/42 (51%), Gaps = 1/42 (2%)
Frame = -1
Query: 154 IFPNRYTCIKATTTKKFPIRWGELHGSNEVIS*TMS-VYRSM 194
I P TTTK F RWG+LH SN T++ VYR +
Sbjct: 190 IMPETNILETTTTTKPFFTRWGQLHRSNNT*CSTVNHVYRQI 65
>TC228621 similar to UP|Q8RXD8 (Q8RXD8) WD repeat protein-like, partial (70%)
Length = 1014
Score = 31.2 bits (69), Expect = 1.8
Identities = 14/25 (56%), Positives = 17/25 (68%)
Frame = -1
Query: 160 TCIKATTTKKFPIRWGELHGSNEVI 184
T ATTTK +P R G LHGS++ I
Sbjct: 1014 TLCSATTTKFYPTR*GRLHGSHDAI 940
>TC228620 weakly similar to UP|Q8RXD8 (Q8RXD8) WD repeat protein-like,
partial (29%)
Length = 622
Score = 31.2 bits (69), Expect = 1.8
Identities = 14/25 (56%), Positives = 17/25 (68%)
Frame = -3
Query: 160 TCIKATTTKKFPIRWGELHGSNEVI 184
T ATTTK +P R G LHGS++ I
Sbjct: 485 TLCSATTTKFYPTR*GRLHGSHDAI 411
>TC207246 similar to UP|Q6YWF4 (Q6YWF4) Rhomboid family protein-like, partial
(15%)
Length = 1053
Score = 30.8 bits (68), Expect = 2.3
Identities = 20/58 (34%), Positives = 34/58 (58%)
Frame = +2
Query: 270 HLNFIPILIEMPLSVFLLLVGSLVKFLYGDFTNQIALLLRTEKSQLL*SLLVSFTHTI 327
+++ IP+L+E+ L + L++GS +KF Q+ L + QLL* LL S+ T+
Sbjct: 350 YIHQIPLLVELDLYLQ*LVLGSCIKF-KTSMLLQVMLQRTCSRRQLL*QLLFSY*ATL 520
>CA785241
Length = 435
Score = 30.8 bits (68), Expect = 2.3
Identities = 14/26 (53%), Positives = 18/26 (68%), Gaps = 3/26 (11%)
Frame = -2
Query: 162 IKATT---TKKFPIRWGELHGSNEVI 184
IK TT +K FP +WG+LH SN+ I
Sbjct: 209 IKITTRRQSKPFPTKWGQLHESNDTI 132
>TC205552 similar to UP|PMM_ARATH (O80840) Probable phosphomannomutase (PMM)
, partial (25%)
Length = 1033
Score = 30.0 bits (66), Expect = 4.0
Identities = 11/16 (68%), Positives = 12/16 (74%)
Frame = -3
Query: 165 TTTKKFPIRWGELHGS 180
TTT +P RWG LHGS
Sbjct: 686 TTTTPYPTRWGRLHGS 639
>TC218482
Length = 1214
Score = 29.6 bits (65), Expect = 5.2
Identities = 17/48 (35%), Positives = 24/48 (49%)
Frame = +1
Query: 628 GCRTFLLDCCTCSI*FVDV*CWHSWMLIS*P**IANFKTWMGYVLSLK 675
G F+L C C + F+ W I+ P + N K W+ Y+LSLK
Sbjct: 169 GLSIFVLVCQKCIVFFLKSSEWIQMPKIANPNCVDNLKMWILYLLSLK 312
>BE822490 SP|P43565|RI1 Serine/threonine-protein kinase RIM15 (EC 2.7.1.-).
[Baker's yeast] {Saccharomyces cerevisiae}, partial (0%)
Length = 712
Score = 29.3 bits (64), Expect = 6.8
Identities = 12/25 (48%), Positives = 13/25 (52%)
Frame = +3
Query: 160 TCIKATTTKKFPIRWGELHGSNEVI 184
T TTT +P WG LHGS I
Sbjct: 600 TTTTTTTTTPYPTXWGRLHGSTXXI 674
>TC213463 similar to UP|Q8H3D7 (Q8H3D7) Leaf senescence related protein-like,
partial (10%)
Length = 674
Score = 28.9 bits (63), Expect = 8.9
Identities = 13/21 (61%), Positives = 15/21 (70%)
Frame = -2
Query: 165 TTTKKFPIRWGELHGSNEVIS 185
TTTK +PIR HGS +VIS
Sbjct: 535 TTTKSYPIR*DRFHGSYKVIS 473
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.360 0.162 0.604
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 40,925,220
Number of Sequences: 63676
Number of extensions: 678089
Number of successful extensions: 6827
Number of sequences better than 10.0: 38
Number of HSP's better than 10.0 without gapping: 3581
Number of HSP's successfully gapped in prelim test: 277
Number of HSP's that attempted gapping in prelim test: 3166
Number of HSP's gapped (non-prelim): 4039
length of query: 758
length of database: 12,639,632
effective HSP length: 104
effective length of query: 654
effective length of database: 6,017,328
effective search space: 3935332512
effective search space used: 3935332512
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 37 (21.8 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146719.12


