

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146712.3 - phase: 0 /pseudo
(276 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
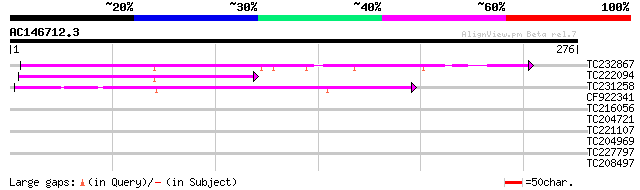
Score E
Sequences producing significant alignments: (bits) Value
TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial ... 79 2e-15
TC222094 75 4e-14
TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kin... 65 3e-11
CF922341 37 0.010
TC216056 similar to GB|AAP37836.1|30725628|BT008477 At2g39450 {A... 30 1.2
TC204721 similar to GB|AAL84954.1|19310435|AY078954 F17H15.1/F17... 28 3.5
TC221107 similar to UP|Q6NLR6 (Q6NLR6) At4g24230, partial (14%) 28 3.5
TC204969 similar to UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease ... 28 5.9
TC227797 similar to UP|Q8H1N8 (Q8H1N8) At1g20880/F9H16_14, parti... 28 5.9
TC208497 27 7.7
>TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial (3%)
Length = 1144
Score = 79.3 bits (194), Expect = 2e-15
Identities = 76/265 (28%), Positives = 122/265 (45%), Gaps = 15/265 (5%)
Frame = -3
Query: 6 LKEYATPSMEEPQAIIVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLS 65
L++Y++ ++ + I P V+ P+L+ L+Q N F G P EDP H+++++ +
Sbjct: 902 LEDYSSSTVPQFFTSIAQPEVQAQTITYPPSLIQLIQSNFFHGLPNEDPYAHLATYIEIC 723
Query: 66 GTIK---ENQEAV*LHLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPL- 121
TI+ +A+ L L FSL A W HS + S+ SWD + FL ++ P K
Sbjct: 722 NTIRLAGVPADAIRLSLLSFSLSGEAKRWLHSFKGNSLKSWDEVVEKFLKKYFPESKTAE 543
Query: 122 -NLEIKS-HGLIKKMGSLSMKRGR-VSRKSLDFVPTMAWKSGL*STPF-NGLSYTTKMSV 177
I S H + S +++R R + RK+ PT + + F + L +K V
Sbjct: 542 GKAAISSFHQFPDESLSEALERFRGLLRKT----PTHGFSEPIQLNIFIDELRPESKQLV 375
Query: 178 DAAAGGSLMNKNYTEAYALMKTW-------LRITTNGPTREPSPLLLPLRKRQVYTKSLN 230
DA+ GG + K EA L+++ LR + PT++ LL L +
Sbjct: 374 DASVGGKIKMKTPDEAMDLIESMAASDIAILRDRAHIPTKKS---LLELTSQD------- 225
Query: 231 ITTLLLRLKL*PKNSKNLMLMLSHL 255
TLL + KL K + L LS L
Sbjct: 224 --TLLAQNKLLSKQLETLTKTLSKL 156
>TC222094
Length = 984
Score = 74.7 bits (182), Expect = 4e-14
Identities = 38/120 (31%), Positives = 66/120 (54%), Gaps = 3/120 (2%)
Frame = -2
Query: 5 TLKEYATPSMEEPQAIIVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRL 64
TL++Y++P + + I P V+ NF +L+ L+Q N F G PTEDP H+++++ +
Sbjct: 845 TLEDYSSPIIPQYFTSIARPEVQTANFSYPYSLIQLIQGNLFYGLPTEDPYAHLATYIDI 666
Query: 65 SGTIK---ENQEAV*LHLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLKPL 121
T+K ++A+ L LF FSL A W S + ++ +W+ FL ++ P K +
Sbjct: 665 CNTVKIVGVPEDAIHLDLFCFSLAGEARTWLRSFKGNNLRTWNEXXEKFLKKYFPESKTI 486
>TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kinase/response
regulator, partial (3%)
Length = 1374
Score = 65.5 bits (158), Expect = 3e-11
Identities = 57/200 (28%), Positives = 89/200 (44%), Gaps = 4/200 (2%)
Frame = +2
Query: 3 ERTLKEYATPSMEEPQAIIVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFL 62
ERTL+E P YP E + +K L++L+ + F G EDP+ H+ F
Sbjct: 434 ERTLREMVAPDFTYESLCNQYPD-EDVPYVLKTGLIHLLPK--FHGLVGEDPHKHLKEFH 604
Query: 63 RLSGTIKE---NQEAV*LHLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARFSPHLK 119
+ T+K ++ + L FP SL A W + L SI +WD ++R FL +F P +
Sbjct: 605 IVCSTMKPPDVQEDHIFLKAFPHSLEGVAKDWLYYLAPRSIFNWDDLKRVFLEKFFPASR 784
Query: 120 PLNLEIKSHGLIKKMGSLSMKRGRVSRKSLDFVP-TMAWKSGL*STPFNGLSYTTKMSVD 178
+ G+ + + +KS P K L + LS + +D
Sbjct: 785 TTTIRKDISGIRQLSRESLYEYWERFKKSCASCPHHQISKQLLL*YFYEELSNMKRSMID 964
Query: 179 AAAGGSLMNKNYTEAYALMK 198
AA+GG+L + TEA L+K
Sbjct: 965 AASGGALGDMTPTEARNLIK 1024
>CF922341
Length = 675
Score = 37.0 bits (84), Expect = 0.010
Identities = 25/108 (23%), Positives = 52/108 (48%)
Frame = +1
Query: 7 KEYATPSMEEPQAIIVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLRLSG 66
++YA ++EE + + P + I P ++ +++ G T P H+ + + G
Sbjct: 349 EDYAFANLEE---LFLVPNI------ITPPKFKVLDFDKYKG--TTCPKNHLKMYCQKMG 495
Query: 67 TIKENQEAV*LHLFPFSLRDRASAWFHSLEVGSITSWDGMRRAFLARF 114
+++E + +H F SL A W+ +LE + SW + AF+ ++
Sbjct: 496 AYAKDEELL-IHSFQESLTGVAVTWYTNLEPSRVHSWKDLMVAFVRQY 636
>TC216056 similar to GB|AAP37836.1|30725628|BT008477 At2g39450 {Arabidopsis
thaliana;} , partial (88%)
Length = 1617
Score = 30.0 bits (66), Expect = 1.2
Identities = 15/51 (29%), Positives = 26/51 (50%)
Frame = +2
Query: 13 SMEEPQAIIVYPTVEGNNFEIKPALLNLVQQNQFSGSPTEDPNLHISSFLR 63
S+ P + I P + N ++ + + Q+Q SGSP PN++ F+R
Sbjct: 1298 SLSSPVSCICEPFLVVRNLNLRASH*EVGYQSQISGSPDNIPNVYHDRFIR 1450
>TC204721 similar to GB|AAL84954.1|19310435|AY078954 F17H15.1/F17H15.1
{Arabidopsis thaliana;} , partial (30%)
Length = 1762
Score = 28.5 bits (62), Expect = 3.5
Identities = 16/48 (33%), Positives = 20/48 (41%)
Frame = +2
Query: 3 ERTLKEYATPSMEEPQAIIVYPTVEGNNFEIKPALLNLVQQNQFSGSP 50
ERTLK TP E +VY + G N PA+ Q + P
Sbjct: 410 ERTLKIDGTPEQIESAKQLVYQVISGENRVRNPAMSGGYPQQGYQSRP 553
>TC221107 similar to UP|Q6NLR6 (Q6NLR6) At4g24230, partial (14%)
Length = 498
Score = 28.5 bits (62), Expect = 3.5
Identities = 15/37 (40%), Positives = 19/37 (50%)
Frame = +3
Query: 192 EAYALMKTWLRITTNGPTREPSPLLLPLRKRQVYTKS 228
E Y L K + T GP REP P+ L L R + +S
Sbjct: 363 ELYGLHK----VATEGPCREPQPMALKLAARAKWVES 461
>TC204969 similar to UP|Q9ZTT3 (Q9ZTT3) Subtilisin-like protease C1, partial
(12%)
Length = 830
Score = 27.7 bits (60), Expect = 5.9
Identities = 10/25 (40%), Positives = 20/25 (80%)
Frame = +3
Query: 251 MLSHLLLHPLLVKSVVYPVISVLIV 275
++SHLLL+P L +S++ P++ L++
Sbjct: 303 LISHLLLYPQLAQSILVPLLVELLL 377
>TC227797 similar to UP|Q8H1N8 (Q8H1N8) At1g20880/F9H16_14, partial (59%)
Length = 1737
Score = 27.7 bits (60), Expect = 5.9
Identities = 23/92 (25%), Positives = 36/92 (39%)
Frame = +3
Query: 47 SGSPTEDPNLHISSFLRLSGTIKENQEAV*LHLFPFSLRDRASAWFHSLEVGSITSWDGM 106
S SP P ++ LSG + E V +HL W H ++ G + DG
Sbjct: 444 SSSPIRTPEDPRATVS*LSGIPRPQGERVLIHLLLLMAEGPIVIWLHLVDHGPLCLMDG- 620
Query: 107 RRAFLARFSPHLKPLNLEIKSHGLIKKMGSLS 138
L PL LE+ + ++ +G L+
Sbjct: 621 ---------SDLPPLMLEVCNQLVVLMLGVLA 689
>TC208497
Length = 1555
Score = 27.3 bits (59), Expect = 7.7
Identities = 21/51 (41%), Positives = 31/51 (60%), Gaps = 4/51 (7%)
Frame = +2
Query: 214 PLLLPLR--KRQVYTKSLNITTLLLR-LKL*PK-NSKNLMLMLSHLLLHPL 260
PL+ PL KRQ+ +S ++ TLL+R L * K N K +M+ L ++L L
Sbjct: 344 PLVCPLTL*KRQMRMRSNHLVTLLIRFLNT*MKGNLKKMMIWLLRIMLSSL 496
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.138 0.406
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,843,915
Number of Sequences: 63676
Number of extensions: 162037
Number of successful extensions: 1100
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 1094
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1098
length of query: 276
length of database: 12,639,632
effective HSP length: 96
effective length of query: 180
effective length of database: 6,526,736
effective search space: 1174812480
effective search space used: 1174812480
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146712.3


