

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146705.12 - phase: 0 /pseudo
(811 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
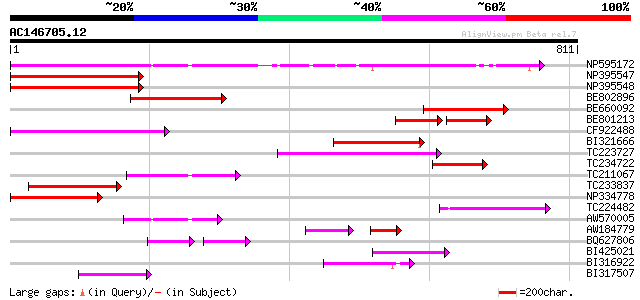
Score E
Sequences producing significant alignments: (bits) Value
NP595172 polyprotein [Glycine max] 363 e-100
NP395547 reverse transcriptase [Glycine max] 291 6e-79
NP395548 reverse transcriptase [Glycine max] 287 1e-77
BE802896 187 2e-47
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 186 4e-47
BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis ... 105 8e-43
CF922488 167 2e-41
BI321666 165 6e-41
TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 135 6e-32
TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (... 128 9e-30
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 124 2e-28
TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 109 6e-24
NP334778 reverse transcriptase [Glycine max] 107 2e-23
TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, part... 97 4e-20
AW570005 87 3e-17
AW184779 56 4e-15
BQ627806 46 1e-12
BI425021 65 2e-10
BI316922 62 1e-09
BI317507 59 9e-09
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 363 bits (933), Expect = e-100
Identities = 259/779 (33%), Positives = 393/779 (50%), Gaps = 15/779 (1%)
Frame = +1
Query: 1 NKATRKDHFPLPFIDQMLERLAKHSHFCYLDGYSGFFQIPIHPNDQEKTTFTCPFGTFAY 60
N T KD FP+P +D++L+ L +F LD SG+ QI + P D+EKT F G + +
Sbjct: 1969 NAITVKDSFPMPTVDELLDELHGAQYFSKLDLRSGYHQILVQPEDREKTAFRTHHGHYEW 2148
Query: 61 RRMPFGLCNAPATFQRCMMSIFSDFVEKIMEVFMDDFSVHGSNFDDCLTNLEKVLERCEQ 120
MPFGL NAPATFQ M IF + K + VF DD ++ +++ D L +LE VL+ +Q
Sbjct: 2149 LVMPFGLTNAPATFQCLMNKIFQFALRKFVLVFFDDILIYSASWKDHLKHLESVLQTLKQ 2328
Query: 121 VNLVLNWEKCHFMVREGIVLGHLVFDRGIEVDRAKIEIIKKMLPPTSVKEIRSFLGHAGF 180
L KC F E LGH V G+ ++ K++ + P +VK++R FLG G+
Sbjct: 2329 HQLFARLSKCSFGDTEVDYLGHKVSGLGVSMENTKVQAVLDWPTPNNVKQLRGFLGLTGY 2508
Query: 181 YRRFIKDFSSITKPLTSLLLKDADFTFDDSCLQAFCRLKEALITAPIIQPPDWNLPFEIM 240
YRRFIK +++I PLT LL KD+ F +++ AF +LK+A+ AP++ PD++ PF +
Sbjct: 2509 YRRFIKSYANIAGPLTDLLQKDS-FLWNNEAEAAFVKLKKAMTEAPVLSLPDFSQPFILE 2685
Query: 241 CDASDYAVGAVLGQRNDKKMHAIYYASKTLDGAQVNYATTEKELLAVVYAIDKFRQYLVG 300
DAS VGAVLGQ H I Y SK L + +ELLA+ A+ KFR YL+G
Sbjct: 2686 TDASGIGVGAVLGQNG----HPIAYFSKKLAPRMQKQSAYTRELLAITEALSKFRHYLLG 2853
Query: 301 SKIIVYTDHSAIKYLLNKKDAKPRLIRWILLLQEFDLEIKDKKGVENVVADHLSRLRETN 360
+K I+ TD ++K L+++ P W+ +D +I+ K G +N AD LSR
Sbjct: 2854 NKFIIRTDQRSLKSLMDQSLQTPEQQAWLHKFLGYDFKIEYKPGKDNQAADALSR----- 3018
Query: 361 KDELPLDDSFPDDQLFLLAQTDAPWYADFVNFLAAGVL--PPELNYQQKKKFFNDLKHYY 418
+F+LA ++ ++ F+ L A ++ P + K D HY
Sbjct: 3019 --------------MFMLAWSEP--HSIFLEELRARLISDPHLKQLMETYKQGADASHYT 3150
Query: 419 WDEPYLFRRGSDGIFRRCIPENE--VSSILTHCHSSSYGGHASTQKTSFKILHSGFWWPS 476
E L+ + R IP E V+ IL HSS GGHA +T + L + F+WP
Sbjct: 3151 VREGLLYWKD-----RVVIPAEEEIVNKILQEYHSSPIGGHAGITRTLAR-LKAQFYWPK 3312
Query: 477 LFKDVHLFISKCDKCQRTGSITKRNEMPLNNILEVEI-FDVW---GIDFMGPFPSSFGNQ 532
+ +DV +I KC CQ+ S N +P + + I VW +DF+ P+SFG
Sbjct: 3313 MQEDVKAYIQKCLICQQAKS---NNTLPAGLLQPLPIPQQVWEDVAMDFITGLPNSFGLS 3483
Query: 533 YILVAVDYVSKWVEAIASPTN-DAQVVIKMFKKVIFPRFGVPRVVISDGGSHFISRHFEK 591
I+V +D ++K+ I + +++VV + F I G+PR ++SD F S ++
Sbjct: 3484 VIMVVIDRLTKYAHFIPLKADYNSKVVAEAFMSHIVKLHGIPRSIVSDRDRVFTSTFWQH 3663
Query: 592 LLQKLGVRHKIATPYHPQTSGQVEVSNRQIKAILEKTVSTSRTDWSNKLDDALWAYRTAY 651
L + G +++ YHPQ+ GQ EV N+ ++ L W L A + Y TAY
Sbjct: 3664 LFKLQGTTLAMSSAYHPQSDGQSEVLNKCLEMYLRCFTYEHPKGWVKALPWAEFWYNTAY 3843
Query: 652 KTPIGMTPFKLVYGKSCHLPVELEHKAYWAIRNLNLDPNLAGDKRKLQLNELEELRMDAY 711
+GMTPF+ +YG+ P L +A ++D + + QL + + L
Sbjct: 3844 HMSLGMTPFRALYGRE---PPTLTRQA------CSIDDPA---EVREQLTDRDALLAKLK 3987
Query: 712 ENARIYKERTKTWHDKKIIKRHFKSGDLVLL------FNSRLKLFPGKLRSRWSGPFQV 764
N ++ K DKK + F+ GD VL+ +S + KL R+ GPF+V
Sbjct: 3988 INLTRAQQVMKRQADKKRLDVSFQIGDEVLVKLQPYRQHSAVLRKNQKLSMRYFGPFKV 4164
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 291 bits (746), Expect = 6e-79
Identities = 133/191 (69%), Positives = 159/191 (82%)
Frame = +1
Query: 1 NKATRKDHFPLPFIDQMLERLAKHSHFCYLDGYSGFFQIPIHPNDQEKTTFTCPFGTFAY 60
N+ATRKDH+PLPF+DQML+RLA+ S + +LDGYSG+ QI + P DQEKT FTCPF FAY
Sbjct: 190 NEATRKDHYPLPFMDQMLKRLARQSFYRFLDGYSGYNQIAVDPQDQEKTAFTCPFSVFAY 369
Query: 61 RRMPFGLCNAPATFQRCMMSIFSDFVEKIMEVFMDDFSVHGSNFDDCLTNLEKVLERCEQ 120
RRMPFGLCNA TFQRCMM+IF D VEK +EVFMDDFS G++F +CL NLEKVL+RCE+
Sbjct: 370 RRMPFGLCNASTTFQRCMMAIFDDMVEKCIEVFMDDFSFFGASFGNCLANLEKVLQRCEK 549
Query: 121 VNLVLNWEKCHFMVREGIVLGHLVFDRGIEVDRAKIEIIKKMLPPTSVKEIRSFLGHAGF 180
NLVLNWEKCHFMV+EGIVLGH + RGIEV + K+++I K+ PP +VK I SFLGH GF
Sbjct: 550 SNLVLNWEKCHFMVQEGIVLGHKISKRGIEVVKEKLDVIDKLPPPVNVKGIHSFLGHVGF 729
Query: 181 YRRFIKDFSSI 191
YRRFIKDF+ +
Sbjct: 730 YRRFIKDFTKV 762
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 287 bits (735), Expect = 1e-77
Identities = 131/191 (68%), Positives = 158/191 (82%)
Frame = +1
Query: 1 NKATRKDHFPLPFIDQMLERLAKHSHFCYLDGYSGFFQIPIHPNDQEKTTFTCPFGTFAY 60
N+ATRKDHFPLPF+DQMLERLA H+++C+LD Y G+ QI + P DQEK FTCPFG FAY
Sbjct: 190 NEATRKDHFPLPFMDQMLERLAGHAYYCFLDAYFGYNQIVVDPKDQEKMAFTCPFGVFAY 369
Query: 61 RRMPFGLCNAPATFQRCMMSIFSDFVEKIMEVFMDDFSVHGSNFDDCLTNLEKVLERCEQ 120
RR+PFGLCNAP TFQ CM++IF+D VEK +EVFMDDFSV + + CL LE VL+RC +
Sbjct: 370 RRIPFGLCNAPTTFQMCMLAIFADIVEKSIEVFMDDFSVFVPSLESCLKKLEMVLQRCVE 549
Query: 121 VNLVLNWEKCHFMVREGIVLGHLVFDRGIEVDRAKIEIIKKMLPPTSVKEIRSFLGHAGF 180
NLVLNWEKCHFMVREGIVLGH + RGIEVD+ KI++I+K+ PP++VK IRSFLG A F
Sbjct: 550 TNLVLNWEKCHFMVREGIVLGHKISTRGIEVDQTKIDVIEKLPPPSNVKGIRSFLGQARF 729
Query: 181 YRRFIKDFSSI 191
YRRFIKDF+ +
Sbjct: 730 YRRFIKDFTKV 762
>BE802896
Length = 416
Score = 187 bits (474), Expect = 2e-47
Identities = 87/138 (63%), Positives = 108/138 (78%)
Frame = -2
Query: 173 SFLGHAGFYRRFIKDFSSITKPLTSLLLKDADFTFDDSCLQAFCRLKEALITAPIIQPPD 232
SFLGHAGFYRRFI+DF + PL++LL K+ +F F+D C AF LK ALIT PIIQ PD
Sbjct: 415 SFLGHAGFYRRFIRDFRKVALPLSNLLQKEVEFDFNDKCK*AFDCLKRALITTPIIQAPD 236
Query: 233 WNLPFEIMCDASDYAVGAVLGQRNDKKMHAIYYASKTLDGAQVNYATTEKELLAVVYAID 292
W PFE+MCDAS+YA+G VL Q+ DK IYY+S+TLD AQ NY TTEKELLA+V+A++
Sbjct: 235 WTAPFELMCDASNYALGVVLAQKIDKLPRVIYYSSRTLDAAQANYTTTEKELLAIVFALE 56
Query: 293 KFRQYLVGSKIIVYTDHS 310
KF YL+G++IIVY DH+
Sbjct: 55 KFHSYLLGTRIIVYIDHA 2
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 186 bits (472), Expect = 4e-47
Identities = 80/122 (65%), Positives = 106/122 (86%)
Frame = -3
Query: 592 LLQKLGVRHKIATPYHPQTSGQVEVSNRQIKAILEKTVSTSRTDWSNKLDDALWAYRTAY 651
LL+K GV H+++TPYHPQT+GQ E+SNR+IK ILEK V SR DWS +LDDALWA+RTAY
Sbjct: 367 LLKKYGVVHRVSTPYHPQTNGQAEISNREIKRILEKIVQPSRKDWSTRLDDALWAHRTAY 188
Query: 652 KTPIGMTPFKLVYGKSCHLPVELEHKAYWAIRNLNLDPNLAGDKRKLQLNELEELRMDAY 711
K PIGM+P+++V+GK+CHLPVE+EHKAYWA++ N + AG++RKLQL+EL+E+R++AY
Sbjct: 187 KAPIGMSPYRVVFGKACHLPVEIEHKAYWAVKTCNFSMDQAGEERKLQLSELDEIRLEAY 8
Query: 712 EN 713
EN
Sbjct: 7 EN 2
>BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis thaliana},
partial (3%)
Length = 416
Score = 105 bits (262), Expect(2) = 8e-43
Identities = 45/64 (70%), Positives = 57/64 (88%)
Frame = +2
Query: 626 EKTVSTSRTDWSNKLDDALWAYRTAYKTPIGMTPFKLVYGKSCHLPVELEHKAYWAIRNL 685
EK V++SR DWS+KL+DALWA +TA KTPIG+TPF++VY K+CHLPVEL+HKAYWA++ L
Sbjct: 224 EKNVASSRKDWSSKLEDALWACKTAKKTPIGLTPFQMVYRKACHLPVELKHKAYWAMKCL 403
Query: 686 NLDP 689
N DP
Sbjct: 404 NFDP 415
Score = 87.8 bits (216), Expect(2) = 8e-43
Identities = 40/66 (60%), Positives = 50/66 (75%)
Frame = +3
Query: 553 NDAQVVIKMFKKVIFPRFGVPRVVISDGGSHFISRHFEKLLQKLGVRHKIATPYHPQTSG 612
NDA++VIK KK IF RFG+PR++ISDGGSHF K+L+ VRHK+ T YHPQT+G
Sbjct: 3 NDAKIVIKFLKKNIFSRFGMPRILISDGGSHFYYSQLNKVLKHDSVRHKVETSYHPQTNG 182
Query: 613 QVEVSN 618
Q +VSN
Sbjct: 183 QAKVSN 200
>CF922488
Length = 741
Score = 167 bits (423), Expect = 2e-41
Identities = 91/228 (39%), Positives = 133/228 (57%)
Frame = +3
Query: 1 NKATRKDHFPLPFIDQMLERLAKHSHFCYLDGYSGFFQIPIHPNDQEKTTFTCPFGTFAY 60
N A+ KD FPLP I+ +++ S F ++DG+SG+ QI I P D EKTTF +GTF Y
Sbjct: 54 N*ASPKDKFPLPHINVLVDNTTSFSQFSFMDGFSGYNQIKIAPEDMEKTTFITLWGTFCY 233
Query: 61 RRMPFGLCNAPATFQRCMMSIFSDFVEKIMEVFMDDFSVHGSNFDDCLTNLEKVLERCEQ 120
+ M FGL N AT+QR M+++F D + K +EV+MDD V ++ L NL K+ R +
Sbjct: 234 KAMSFGLKNVGATYQRAMVALF*DMMHKEIEVYMDDMIVKSRTEEEHLVNLRKLFRRLRK 413
Query: 121 VNLVLNWEKCHFMVREGIVLGHLVFDRGIEVDRAKIEIIKKMLPPTSVKEIRSFLGHAGF 180
L LN KC F V+ +L + RGIEVD K+++I +M P + K+++ FLG +
Sbjct: 414 YRLRLNPAKCMFEVKSRKLLDFIDS*RGIEVDSNKVKVILEMAKPHTEKQVQGFLGRLNY 593
Query: 181 YRRFIKDFSSITKPLTSLLLKDADFTFDDSCLQAFCRLKEALITAPII 228
RFI + +PL LL K+ +D C AF R+K+ LI ++
Sbjct: 594 IVRFIS*LIATCEPLFILLCKNQFVKWDHDC*VAFERIKQCLINPHVL 737
>BI321666
Length = 430
Score = 165 bits (418), Expect = 6e-41
Identities = 78/134 (58%), Positives = 100/134 (74%), Gaps = 4/134 (2%)
Frame = +2
Query: 464 SFKILHSGFWWPSLFKDVHLFISKCDKCQRTGSITKRNEMPLNNILEVEIFDVWGIDFMG 523
S +L S F+ PS+FKD ++ C+KCQRT S++KRNE+PL+ ILEVEIFD WGIDF+G
Sbjct: 5 STNVLQSRFYLPSIFKDAYVHAQSCNKCQRTRSVSKRNELPLHTILEVEIFDYWGIDFVG 184
Query: 524 PFPSSFGNQYILVAVDYVSKWVEAIASPTNDAQVVIKMFKKVIFPRFGVPRVVISDGGSH 583
PFP SF N+YILV VDYVSKWVEA+A +DA++VIK KK IF R GVP V+I +GGSH
Sbjct: 185 PFPPSFSNEYILVVVDYVSKWVEAVACQKSDAKIVIKFLKKQIFSRLGVPWVLIDNGGSH 364
Query: 584 F----ISRHFEKLL 593
++R F ++
Sbjct: 365 LCNA*LARFFNTIM 406
>TC223727 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (9%)
Length = 843
Score = 135 bits (341), Expect = 6e-32
Identities = 71/236 (30%), Positives = 123/236 (52%), Gaps = 2/236 (0%)
Frame = +1
Query: 384 PWYADFVNFLAAGVLPPELNYQQKKKFFNDLKHYYWDEPYLFRRGSDGIFRRCIPENEVS 443
PWY D ++ + PE+ K+ ++ L++R D RC+ E +
Sbjct: 136 PWYFDIKRYVISKEYLPEIADNDKRTLRRLAAGFFMSGSILYKRNHDMKPLRCVDAREAN 315
Query: 444 SILTHCHSSSYGGHASTQKTSFKILHSGFWWPSLFKDVHLFISKCDKCQRTGSITKRNEM 503
++ H S+G HA+ + KIL +G++W ++ D + + KC KCQ
Sbjct: 316 HMIEEVHEGSFGTHANGHAMARKILRAGYYWLTMESDCCVHVRKCHKCQAFADNVNAPPH 495
Query: 504 PLNNILEVEIFDVWGIDFMGPF--PSSFGNQYILVAVDYVSKWVEAIASPTNDAQVVIKM 561
PLN + F +WGID +G +S G+++ILVA+DY +KWVEA + VV++
Sbjct: 496 PLNVMSSPWPFSMWGIDVIGAIEPKASNGHRFILVAIDYFTKWVEAASYTDVMRGVVVRF 675
Query: 562 FKKVIFPRFGVPRVVISDGGSHFISRHFEKLLQKLGVRHKIATPYHPQTSGQVEVS 617
KK I R+G+PR +I+D G++ ++ ++ ++ ++H TPY P+ + VEV+
Sbjct: 676 IKKEIICRYGLPRKIITDNGTNLNNKMMGEICEEFKIQHHNPTPYRPKMN*AVEVA 843
>TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (32%)
Length = 482
Score = 128 bits (322), Expect = 9e-30
Identities = 55/78 (70%), Positives = 68/78 (86%)
Frame = -3
Query: 606 YHPQTSGQVEVSNRQIKAILEKTVSTSRTDWSNKLDDALWAYRTAYKTPIGMTPFKLVYG 665
YHPQT+GQ EVSN++IK +LE V +SR DW+ KLDDA WAYR A+KTPIG++PF+LVYG
Sbjct: 234 YHPQTNGQAEVSNKEIKRVLENIVVSSRKDWALKLDDAFWAYRIAFKTPIGLSPFQLVYG 55
Query: 666 KSCHLPVELEHKAYWAIR 683
K+CHL VELEHKAYWA++
Sbjct: 54 KACHLSVELEHKAYWALK 1
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 124 bits (311), Expect = 2e-28
Identities = 66/163 (40%), Positives = 97/163 (59%)
Frame = +1
Query: 167 SVKEIRSFLGHAGFYRRFIKDFSSITKPLTSLLLKDADFTFDDSCLQAFCRLKEALITAP 226
SV +IRSF G A FYRRF+ +FS++ PL L+ K+ FT+ + QAF LKE L AP
Sbjct: 109 SVGDIRSFHGLASFYRRFVPNFSTVASPLNELVKKNMAFTWGEKQEQAFALLKEKLTKAP 288
Query: 227 IIQPPDWNLPFEIMCDASDYAVGAVLGQRNDKKMHAIYYASKTLDGAQVNYATTEKELLA 286
++ PD++ FE+ CDAS V AVL Q H I Y S+ L A +NY T +KEL A
Sbjct: 289 VLALPDFSKTFELECDASGVGVRAVLLQGG----HPIAYFSEKLHSATLNYPTYDKELYA 456
Query: 287 VVYAIDKFRQYLVGSKIIVYTDHSAIKYLLNKKDAKPRLIRWI 329
++ A + +LV + ++++DH ++KY+ K R +W+
Sbjct: 457 LIRAPQTWEHFLVCKEFVIHSDHQSLKYIRGKSKLNKRHAKWV 585
>TC233837 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 402
Score = 109 bits (272), Expect = 6e-24
Identities = 56/133 (42%), Positives = 83/133 (62%)
Frame = +2
Query: 27 FCYLDGYSGFFQIPIHPNDQEKTTFTCPFGTFAYRRMPFGLCNAPATFQRCMMSIFSDFV 86
F ++DG+SG+ QI + D EKTTF +GTF+YR M FGL N AT+QR M+++F D +
Sbjct: 2 FSFMDGFSGYNQI*MAREDVEKTTFVTLWGTFSYRVMAFGLKNTGATYQRAMVALFHDMM 181
Query: 87 EKIMEVFMDDFSVHGSNFDDCLTNLEKVLERCEQVNLVLNWEKCHFMVREGIVLGHLVFD 146
K +EV++DD + L NL K+ R ++ L LN KC F V+ G +LG +V
Sbjct: 182 HKEIEVYVDDMIAKSRTETEHLVNLCKLFGRLQKYQLKLNPTKCTFGVKSGKLLGFIVSQ 361
Query: 147 RGIEVDRAKIEII 159
+GIE+D K++ +
Sbjct: 362 KGIEIDPEKVKAL 400
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 107 bits (268), Expect = 2e-23
Identities = 52/132 (39%), Positives = 81/132 (60%)
Frame = +3
Query: 1 NKATRKDHFPLPFIDQMLERLAKHSHFCYLDGYSGFFQIPIHPNDQEKTTFTCPFGTFAY 60
N+A+ KD+FPLP ID ++ +A + F ++DG+SG+ QI + P D EKTTF +GTF Y
Sbjct: 30 NRASPKDNFPLPHIDILMANMASFALFSFMDGFSGYNQIKMAPEDMEKTTFITLWGTFCY 209
Query: 61 RRMPFGLCNAPATFQRCMMSIFSDFVEKIMEVFMDDFSVHGSNFDDCLTNLEKVLERCEQ 120
+ M FGL N AT+ R M+++F D + K +E ++D+ ++ L NL+ + + +
Sbjct: 210 KVMSFGLKNFGATYHRAMVALFQDMMHKEIEAYVDEMIAKSRMEEEHLVNLQNLFGQLRK 389
Query: 121 VNLVLNWEKCHF 132
L LN KC F
Sbjct: 390 YRLRLNPRKCVF 425
>TC224482 similar to UP|Q6WAY3 (Q6WAY3) Gag/pol polyprotein, partial (6%)
Length = 669
Score = 96.7 bits (239), Expect = 4e-20
Identities = 59/160 (36%), Positives = 85/160 (52%), Gaps = 1/160 (0%)
Frame = +1
Query: 615 EVSNRQIKAILEKTVSTSRTDWSNKLDDALWAYRTAYKTPIGMTPFKLVYGKSCHLPVEL 674
E +N+ IK I++K ++ S DW L AL YRT+ +T G TPF LVYG LP E+
Sbjct: 1 EAANKNIKKIIQK-MTVSYKDWHEMLPFALHGYRTSVRTSTGATPFSLVYGMEAVLPFEV 177
Query: 675 EHKAYWAIRNLNLDPNLAGDKRKLQLNELEELRMDAYENARIYKERTKTWHDKKIIKRHF 734
E + + L + R QLN +E R+ A + R+Y++R K+ DKK+ R F
Sbjct: 178 EVPSLRILAESGLKESEWAQTRYDQLNLIEGKRLTAMSHGRLYQQRMKSAFDKKVCLRKF 357
Query: 735 KSGDLVL-LFNSRLKLFPGKLRSRWSGPFQVRTVYPYGAI 773
GDLVL + +K GK + GPF V+ + GA+
Sbjct: 358 HEGDLVLKKMSHAVKDHRGKWAPNYEGPFVVKRAFSGGAL 477
>AW570005
Length = 413
Score = 87.0 bits (214), Expect = 3e-17
Identities = 51/141 (36%), Positives = 75/141 (53%)
Frame = -2
Query: 164 PPTSVKEIRSFLGHAGFYRRFIKDFSSITKPLTSLLLKDADFTFDDSCLQAFCRLKEALI 223
PP + + +R FL GFYRRFIK ++++ PL+ LL KD+ F + AF LK +
Sbjct: 409 PPRTARSLRGFLRLTGFYRRFIKGYAAMAAPLSHLLTKDS-FVWSPEADVAFQALKNVVT 233
Query: 224 TAPIIQPPDWNLPFEIMCDASDYAVGAVLGQRNDKKMHAIYYASKTLDGAQVNYATTEKE 283
++ PD+ PF + DAS +GAVL Q H I + SK V +T E
Sbjct: 232 NTLVLALPDFTKPFTVETDASGSDMGAVLSQEG----HPIAFFSKEFCPKLVRSSTYVHE 65
Query: 284 LLAVVYAIDKFRQYLVGSKII 304
L A+ + K+RQYL+G ++
Sbjct: 64 LAAITNVVKKWRQYLLGHHLV 2
>AW184779
Length = 432
Score = 55.8 bits (133), Expect(2) = 4e-15
Identities = 23/69 (33%), Positives = 37/69 (53%)
Frame = +1
Query: 424 LFRRGSDGIFRRCIPENEVSSILTHCHSSSYGGHASTQKTSFKILHSGFWWPSLFKDVHL 483
L++R D + RC+ E +L H S+G HA+ + KIL G++W ++ D +
Sbjct: 16 LYKRNHDMVLLRCVDAREAEQMLVEVHEGSFGTHANIHAMAQKILRVGYYWLTMESDCCI 195
Query: 484 FISKCDKCQ 492
+ KC KCQ
Sbjct: 196 HVWKCHKCQ 222
Score = 44.3 bits (103), Expect(2) = 4e-15
Identities = 21/47 (44%), Positives = 32/47 (67%), Gaps = 2/47 (4%)
Frame = +3
Query: 516 VWGIDFMGPFP--SSFGNQYILVAVDYVSKWVEAIASPTNDAQVVIK 560
+WGID +G +S G+ +ILVA+DY +KWVEA++ + VVI+
Sbjct: 291 MWGIDVIGAIEPKASNGHHFILVAIDYFTKWVEAVSYASVTRSVVIR 431
>BQ627806
Length = 435
Score = 46.2 bits (108), Expect(2) = 1e-12
Identities = 22/67 (32%), Positives = 39/67 (57%)
Frame = +1
Query: 278 ATTEKELLAVVYAIDKFRQYLVGSKIIVYTDHSAIKYLLNKKDAKPRLIRWILLLQEFDL 337
+T +EL A+ A+ K+RQYL+G ++ TDH ++K L+++ P ++ L FD
Sbjct: 229 STYVRELAAITVAVKKWRQYLLGHHFVILTDHRSLKELMSQAVQTPEQQIYLARLMGFDY 408
Query: 338 EIKDKKG 344
I+ + G
Sbjct: 409 TIQYRAG 429
Score = 45.4 bits (106), Expect(2) = 1e-12
Identities = 26/67 (38%), Positives = 39/67 (57%)
Frame = +3
Query: 198 LLLKDADFTFDDSCLQAFCRLKEALITAPIIQPPDWNLPFEIMCDASDYAVGAVLGQRND 257
LL+KD F +++ +AF +LK AL AP++ PD+N F + DAS +GA+L Q +
Sbjct: 3 LLVKD-QFHWNEEADRAFSQLKLALCQAPVLGLPDFNSSFVVETDASGIGMGAILSQNHH 179
Query: 258 KKMHAIY 264
IY
Sbjct: 180 PLAFFIY 200
>BI425021
Length = 426
Score = 64.7 bits (156), Expect = 2e-10
Identities = 38/112 (33%), Positives = 58/112 (50%), Gaps = 1/112 (0%)
Frame = -1
Query: 519 IDFMGPFPSSFGNQYILVAVDYVSKWVEAIASPTND-AQVVIKMFKKVIFPRFGVPRVVI 577
+DF+ P G+ I V V+ SK + PT+ A +V +F ++ G PR ++
Sbjct: 381 MDFIVGLPPYHGHTTIFVVVNRFSKGIHLGTLPTSHTAHMVASLFLNIVIKLHGFPRSIV 202
Query: 578 SDGGSHFISRHFEKLLQKLGVRHKIATPYHPQTSGQVEVSNRQIKAILEKTV 629
SD FIS ++ L + G ++++ YHPQT GQ EV NR I+ L V
Sbjct: 201 SDRDPLFISHFWQDLFRLSGTVLRMSSAYHPQTDGQTEVLNRVIEQYLRAFV 46
>BI316922
Length = 405
Score = 62.0 bits (149), Expect = 1e-09
Identities = 42/135 (31%), Positives = 71/135 (52%), Gaps = 5/135 (3%)
Frame = +3
Query: 450 HSSSYGGHASTQKTSFKILHSGFWWPSLFKDVHLFISKCDKCQRTGSITKRNEMPLNNIL 509
H G H+ +Q + ++L G++ ++ K ++ KC++C + G+I+ L+NI+
Sbjct: 9 HRGICGMHSKSQLMTTRVLRVGYY**TMRKYCTEYVKKCEEC*KFGNISHLLVEELHNIV 188
Query: 510 EVEIFDVWGIDFMGPFPSSFGN-QYILVAVDYVSKWVE----AIASPTNDAQVVIKMFKK 564
F + G+D + PFP S +Y+LV +D +KW+E AI S N V K +
Sbjct: 189 APWPFAI*GVDILRPFPLSKRQVKYLLVGIDQFTKWIETEHIAIISIAN----VRKFV*R 356
Query: 565 VIFPRFGVPRVVISD 579
I FG+P +ISD
Sbjct: 357 NIVC*FGIPNTLISD 401
>BI317507
Length = 359
Score = 58.9 bits (141), Expect = 9e-09
Identities = 32/105 (30%), Positives = 58/105 (54%), Gaps = 1/105 (0%)
Frame = -1
Query: 99 VHGSNFDDCLTNLEKVLERCEQVNLVLNWEKCHFMVREGIVLGHLVFDRGIEVDRAKIEI 158
++ ++ + +L VL+ ++ LV N +KC+F LGH++ + +D K++
Sbjct: 344 IYSPDWKSHIMHLTAVLDVLKKERLVANRKKCYFSQTTIEYLGHVISKDCVAMDSNKVKS 165
Query: 159 IKKMLPPTSVKEIRSFLGHAGFYRRFIKDFSSIT-KPLTSLLLKD 202
+ + P +VK + SFL G+YR+FIKD+ + +PLT L D
Sbjct: 164 VIEWPVPKNVKRVCSFLRLTGYYRKFIKDYGKLAPRPLTDLTKND 30
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.140 0.431
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,026,514
Number of Sequences: 63676
Number of extensions: 555938
Number of successful extensions: 2499
Number of sequences better than 10.0: 62
Number of HSP's better than 10.0 without gapping: 2463
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2482
length of query: 811
length of database: 12,639,632
effective HSP length: 105
effective length of query: 706
effective length of database: 5,953,652
effective search space: 4203278312
effective search space used: 4203278312
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146705.12


