

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
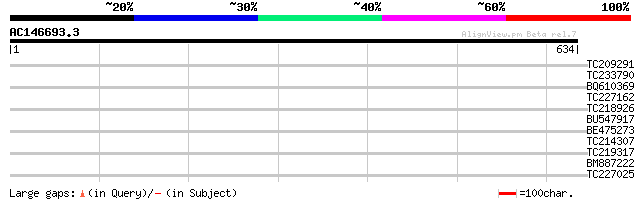
Query= AC146693.3 - phase: 0 /pseudo
(634 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC209291 similar to UP|O23522 (O23522) Triacylglycerol lipase li... 32 0.66
TC233790 31 1.9
BQ610369 similar to GP|9759119|dbj AMP-binding protein {Arabidop... 31 1.9
TC227162 homologue to UP|Q9SH64 (Q9SH64) F22C12.10, partial (43%) 30 2.5
TC218926 similar to UP|CAZ_DROME (Q27294) RNA-binding protein ca... 30 2.5
BU547917 30 4.3
BE475273 weakly similar to GP|20514261|gb| tubulin folding cofac... 29 5.6
TC214307 similar to UP|P93704 (P93704) HMG-1, partial (48%) 28 9.5
TC219317 similar to GB|AAP13385.1|30023704|BT006277 At3g56360 {A... 28 9.5
BM887222 28 9.5
TC227025 homologue to UP|Q9ZPA0 (Q9ZPA0) Inosine monophosphate d... 28 9.5
>TC209291 similar to UP|O23522 (O23522) Triacylglycerol lipase like protein,
partial (16%)
Length = 497
Score = 32.3 bits (72), Expect = 0.66
Identities = 21/69 (30%), Positives = 30/69 (43%)
Frame = +2
Query: 322 SERLSPNSLPRLILSPHTPKCLRPKSPKWPNKWPPLLKH*ESFPVNPKQTPKLMSTPFLM 381
S + +S PR+ S TP C R +S PN W FP++P + PK + +
Sbjct: 284 SSATANSSKPRIKPSIQTPPCQR-RSRHTPNTW--------HFPIDPTE*PKAFTPRHSL 436
Query: 382 GVVSWKGQL 390
G W L
Sbjct: 437 GYPKWVDDL 463
>TC233790
Length = 408
Score = 30.8 bits (68), Expect = 1.9
Identities = 28/92 (30%), Positives = 36/92 (38%)
Frame = +2
Query: 311 KSPRTEKPNRISERLSPNSLPRLILSPHTPKCLRPKSPKWPNKWPPLLKH*ESFPVNPKQ 370
+SPRT N + PN L L+ + P P PK+P KW +L K
Sbjct: 164 RSPRT---NPMESPRHPNQNQNLNLNTNNPTTKTPCKPKYP-KWNKVLL--------TKT 307
Query: 371 TPKLMSTPFLMGVVSWKGQLLKPKELRERVLG 402
T LMST L L P R ++ G
Sbjct: 308 TTTLMSTKLLTTTKQTNHALSAPCSSRMKIKG 403
>BQ610369 similar to GP|9759119|dbj AMP-binding protein {Arabidopsis
thaliana}, partial (24%)
Length = 441
Score = 30.8 bits (68), Expect = 1.9
Identities = 20/65 (30%), Positives = 31/65 (46%), Gaps = 2/65 (3%)
Frame = -2
Query: 311 KSPRTEKPNRIS--ERLSPNSLPRLILSPHTPKCLRPKSPKWPNKWPPLLKH*ESFPVNP 368
+SPR + P R+ + SP + P SP C P S + PP+L+ + +
Sbjct: 311 RSPRWDPPRRLPPLQPFSPLASPSAEPSPSPATCSTPPSTRMT---PPILRSQSNHTRDS 141
Query: 369 KQTPK 373
+QTPK
Sbjct: 140 RQTPK 126
>TC227162 homologue to UP|Q9SH64 (Q9SH64) F22C12.10, partial (43%)
Length = 1808
Score = 30.4 bits (67), Expect = 2.5
Identities = 15/41 (36%), Positives = 20/41 (48%), Gaps = 3/41 (7%)
Frame = +1
Query: 94 NHAVQSKRWGVSL*CVRAFQ---GNAETLSPPWLGKVAYSP 131
++ +QS WG ++ C R Q G L PW GK Y P
Sbjct: 652 HNVLQSTWWGKTVYCTRVHQRC*GEHPILQGPWRGKTLYIP 774
>TC218926 similar to UP|CAZ_DROME (Q27294) RNA-binding protein cabeza
(Sarcoma-associated RNA-binding fly homolog) (P19),
partial (5%)
Length = 906
Score = 30.4 bits (67), Expect = 2.5
Identities = 16/36 (44%), Positives = 19/36 (52%), Gaps = 1/36 (2%)
Frame = +1
Query: 321 ISERLSPNSLPRLILSPHTPKCL-RPKSPKWPNKWP 355
+S LSP L L SPH+P L P P WP+ P
Sbjct: 55 LSRFLSPEGLNSLAHSPHSPPPLPEPPPPSWPSPPP 162
>BU547917
Length = 571
Score = 29.6 bits (65), Expect = 4.3
Identities = 13/28 (46%), Positives = 16/28 (56%)
Frame = +2
Query: 325 LSPNSLPRLILSPHTPKCLRPKSPKWPN 352
L P+S+PR S H + P S KWPN
Sbjct: 359 LKPSSMPRFH*SLHESRSQLPNSHKWPN 442
>BE475273 weakly similar to GP|20514261|gb| tubulin folding cofactor C
{Arabidopsis thaliana}, partial (22%)
Length = 401
Score = 29.3 bits (64), Expect = 5.6
Identities = 21/67 (31%), Positives = 25/67 (36%)
Frame = +2
Query: 312 SPRTEKPNRISERLSPNSLPRLILSPHTPKCLRPKSPKWPNKWPPLLKH*ESFPVNPKQT 371
SP + PN P S I SP+ + SPK P +PP S P T
Sbjct: 140 SPNSPNPNPFPPT-PPTSNSTSIKSPNQSPI*KNSSPKTPTPYPPTTSEPPSKPSPT*ST 316
Query: 372 PKLMSTP 378
P S P
Sbjct: 317 PSTTSLP 337
>TC214307 similar to UP|P93704 (P93704) HMG-1, partial (48%)
Length = 553
Score = 28.5 bits (62), Expect = 9.5
Identities = 18/56 (32%), Positives = 29/56 (51%), Gaps = 1/56 (1%)
Frame = +1
Query: 357 LLKH*ESFPVNPKQTPKLMSTPFLMGVVSWKGQLLKPKELRERVL-GFLVRKMRNR 411
+LK S P PK+ K M F + + +W+G++ + L + L +RKMR R
Sbjct: 31 MLKRPPSLP-RPKRRNKSMRKLFWLTIRNWRGRIQRKTSLTSQSLKSMTMRKMRRR 195
>TC219317 similar to GB|AAP13385.1|30023704|BT006277 At3g56360 {Arabidopsis
thaliana;} , partial (33%)
Length = 829
Score = 28.5 bits (62), Expect = 9.5
Identities = 20/68 (29%), Positives = 28/68 (40%), Gaps = 9/68 (13%)
Frame = -2
Query: 320 RISERLSPNSLP---------RLILSPHTPKCLRPKSPKWPNKWPPLLKH*ESFPVNPKQ 370
+I+ R +P++LP L H P CLRP P+ P S P+ P
Sbjct: 672 KITNRAAPDALPLNPPPPTPSSSSLRRHPPPCLRPTPPRI*LPPAPASHRIPSAPLPPAS 493
Query: 371 TPKLMSTP 378
P S+P
Sbjct: 492 PPPPSSSP 469
>BM887222
Length = 422
Score = 28.5 bits (62), Expect = 9.5
Identities = 9/20 (45%), Positives = 13/20 (65%)
Frame = +1
Query: 66 FHHFMGRYEASIPCPIFPPI 85
FHHF+G+ CP+ PP+
Sbjct: 259 FHHFLGKISEKRQCPLPPPL 318
>TC227025 homologue to UP|Q9ZPA0 (Q9ZPA0) Inosine monophosphate
dehydrogenase, complete
Length = 2097
Score = 28.5 bits (62), Expect = 9.5
Identities = 20/73 (27%), Positives = 26/73 (35%), Gaps = 16/73 (21%)
Frame = +2
Query: 312 SPRTEKPNRISERLSPN---SLPRLILSPHTPKCLRP-------------KSPKWPNKWP 355
S R + ++ SP SLP SP + P +SP WP WP
Sbjct: 197 SRRASPTHTMTSSFSPTTSTSLPTSWTSPRASRAAFPLPCRLLPLRWTPCRSPPWPPPWP 376
Query: 356 PLLKH*ESFPVNP 368
P S P +P
Sbjct: 377 PSAASPSSTPTSP 415
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.364 0.164 0.631
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 33,477,016
Number of Sequences: 63676
Number of extensions: 528180
Number of successful extensions: 7995
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 3228
Number of HSP's successfully gapped in prelim test: 398
Number of HSP's that attempted gapping in prelim test: 4632
Number of HSP's gapped (non-prelim): 3907
length of query: 634
length of database: 12,639,632
effective HSP length: 103
effective length of query: 531
effective length of database: 6,081,004
effective search space: 3229013124
effective search space used: 3229013124
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.5 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146693.3


