

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146649.7 - phase: 0 /pseudo
(859 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
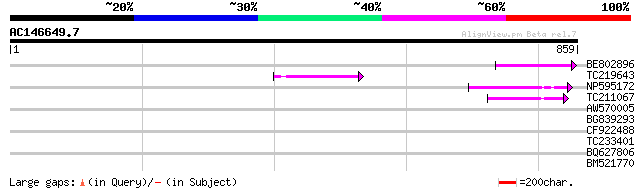
Score E
Sequences producing significant alignments: (bits) Value
BE802896 59 7e-09
TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotei... 59 1e-08
NP595172 polyprotein [Glycine max] 55 2e-07
TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%) 53 5e-07
AW570005 40 0.004
BG839293 40 0.006
CF922488 33 0.70
TC233401 homologue to UP|Q8CBJ2 (Q8CBJ2) Mus musculus 16 days ne... 32 1.6
BQ627806 30 3.5
BM521770 30 4.5
>BE802896
Length = 416
Score = 59.3 bits (142), Expect = 7e-09
Identities = 36/123 (29%), Positives = 59/123 (47%)
Frame = -2
Query: 736 RFVAKSAQHALPFFNLLKKETKFEWTNECEKALQHLKNSLSAPPVLSRPDEGETLYLYLS 795
RF+ + ALP NLL+KE +F++ ++C+ A LK +L P++ PD L
Sbjct: 385 RFIRDFRKVALPLSNLLQKEVEFDFNDKCK*AFDCLKRALITTPIIQAPDWTAPFELMCD 206
Query: 796 VSSEAVSGTLNIETQEGQKPTYFTSKALLGPETRYQKIEKVALALVTAARTH*KYFMAHT 855
S+ A+ L + + + Y++S+ L + Y EK LA+V A Y +
Sbjct: 205 ASNYALGVVLAQKIDKLPRVIYYSSRTLDAAQANYTTTEKELLAIVFALEKFHSYLLGTR 26
Query: 856 IAV 858
I V
Sbjct: 25 IIV 17
>TC219643 weakly similar to UP|Q6WAY7 (Q6WAY7) Gag/pol polyprotein (Fragment),
partial (8%)
Length = 1320
Score = 58.5 bits (140), Expect = 1e-08
Identities = 42/139 (30%), Positives = 65/139 (46%), Gaps = 2/139 (1%)
Frame = +3
Query: 400 GVQPDRSRSTLHKVIVERAEEGKQGPTQRGNTS-PFGDDPTKC-FKLGKGIPEHARAHLI 457
G+ P+ R H E+ + GP Q G K K+G GI R LI
Sbjct: 915 GLPPELERMVAH-------EDQEMGPHQEETELVDLGSGSGKREVKIGTGITAPIREELI 1073
Query: 458 ACLWLNTDLFARSITDMPDIDPSVACHQLTVSPSASVVAQRRKKQFPEKAEGTEKFVKDL 517
L D+FA S DMP + + H+L ++P S V Q+ ++ PE + ++ VK
Sbjct: 1074 ILLRDYQDIFAWSYQDMPGLSSDIVQHRLPLNPECSPVKQKLRRMKPETSLKIKEEVKK* 1253
Query: 518 AEEKFIAEAKYTTWLSNIL 536
+ F+A A+Y W++NI+
Sbjct: 1254 FDAGFLAVARYPKWVANIV 1310
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 54.7 bits (130), Expect = 2e-07
Identities = 46/161 (28%), Positives = 71/161 (43%), Gaps = 3/161 (1%)
Frame = +1
Query: 695 FLSNKIWD*GQSRQMPSIYGTPYSPKKKCIQTLNGML--TSLFR-FVAKSAQHALPFFNL 751
+L +K+ G S + + P ++ L G L T +R F+ A A P +L
Sbjct: 2383 YLGHKVSGLGVSMENTKVQAVLDWPTPNNVKQLRGFLGLTGYYRRFIKSYANIAGPLTDL 2562
Query: 752 LKKETKFEWTNECEKALQHLKNSLSAPPVLSRPDEGETLYLYLSVSSEAVSGTLNIETQE 811
L+K++ F W NE E A LK +++ PVLS PD + L S V L Q
Sbjct: 2563 LQKDS-FLWNNEAEAAFVKLKKAMTEAPVLSLPDFSQPFILETDASGIGVGAVLG---QN 2730
Query: 812 GQKPTYFTSKALLGPETRYQKIEKVALALVTAARTH*KYFM 852
G YF+ K L P + Q L +T A + ++++
Sbjct: 2731 GHPIAYFSKK--LAPRMQKQSAYTRELLAITEALSKFRHYL 2847
>TC211067 similar to UP|Q9LQH2 (Q9LQH2) F15O4.13, partial (9%)
Length = 589
Score = 53.1 bits (126), Expect = 5e-07
Identities = 37/123 (30%), Positives = 55/123 (44%)
Frame = +1
Query: 724 IQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEKALQHLKNSLSAPPVLSR 783
I++ +G+ + RFV + A P L+KK F W + E+A LK L+ PVL+
Sbjct: 121 IRSFHGLASFYRRFVPNFSTVASPLNELVKKNMAFTWGEKQEQAFALLKEKLTKAPVLAL 300
Query: 784 PDEGETLYLYLSVSSEAVSGTLNIETQEGQKPTYFTSKALLGPETRYQKIEKVALALVTA 843
PD +T L S V L +G P + S+ L Y +K AL+ A
Sbjct: 301 PDFSKTFELECDASGVGVRAVL----LQGGHPIAYFSEKLHSATLNYPTYDKELYALIRA 468
Query: 844 ART 846
+T
Sbjct: 469 PQT 477
>AW570005
Length = 413
Score = 40.0 bits (92), Expect = 0.004
Identities = 38/139 (27%), Positives = 59/139 (42%), Gaps = 3/139 (2%)
Frame = -2
Query: 719 PKKKCIQTLNGML--TSLFR-FVAKSAQHALPFFNLLKKETKFEWTNECEKALQHLKNSL 775
P + ++L G L T +R F+ A A P +LL K++ F W+ E + A Q LKN +
Sbjct: 412 PPPRTARSLRGFLRLTGFYRRFIKGYAAMAAPLSHLLTKDS-FVWSPEADVAFQALKNVV 236
Query: 776 SAPPVLSRPDEGETLYLYLSVSSEAVSGTLNIETQEGQKPTYFTSKALLGPETRYQKIEK 835
+ VL+ PD + + S + L +QEG P F SK R
Sbjct: 235 TNTLVLALPDFTKPFTVETDASGSDMGAVL---SQEGH-PIAFFSKEFCPKLVRSSTYVH 68
Query: 836 VALALVTAARTH*KYFMAH 854
A+ + +Y + H
Sbjct: 67 ELAAITNVVKKWRQYLLGH 11
>BG839293
Length = 781
Score = 39.7 bits (91), Expect = 0.006
Identities = 22/55 (40%), Positives = 28/55 (50%)
Frame = +1
Query: 443 KLGKGIPEHARAHLIACLWLNTDLFARSITDMPDIDPSVACHQLTVSPSASVVAQ 497
K+G GI R LI L D+FA S DMP + + HQL ++P S V Q
Sbjct: 565 KIGTGITAPIREELIILLKDYQDIFAWSYQDMPGLSSDIVQHQLPLNPECSPVKQ 729
>CF922488
Length = 741
Score = 32.7 bits (73), Expect = 0.70
Identities = 19/61 (31%), Positives = 29/61 (47%)
Frame = +3
Query: 721 KKCIQTLNGMLTSLFRFVAKSAQHALPFFNLLKKETKFEWTNECEKALQHLKNSLSAPPV 780
+K +Q G L + RF++ P F LL K +W ++C A + +K L P V
Sbjct: 555 EKQVQGFLGRLNYIVRFIS*LIATCEPLFILLCKNQFVKWDHDC*VAFERIKQCLINPHV 734
Query: 781 L 781
L
Sbjct: 735 L 737
>TC233401 homologue to UP|Q8CBJ2 (Q8CBJ2) Mus musculus 16 days neonate
cerebellum cDNA, RIKEN full-length enriched library,
clone:9630015E22 product:odd Oz/ten-m homolog 1
(Drosophila), full insert sequence. (Fragment), partial
(11%)
Length = 403
Score = 31.6 bits (70), Expect = 1.6
Identities = 18/39 (46%), Positives = 22/39 (56%)
Frame = +2
Query: 201 VNHLVHSRHKSGGRPPLAFTTMNYPEEHLTRPFLFLFKL 239
+ H VHSR +S P +FTT +P L RPFL F L
Sbjct: 140 IRHEVHSR*QSNTFP-FSFTTRLFPSTLLIRPFLHHFSL 253
>BQ627806
Length = 435
Score = 30.4 bits (67), Expect = 3.5
Identities = 17/56 (30%), Positives = 26/56 (46%)
Frame = +3
Query: 751 LLKKETKFEWTNECEKALQHLKNSLSAPPVLSRPDEGETLYLYLSVSSEAVSGTLN 806
LL K+ +F W E ++A LK +L PVL PD + + S + L+
Sbjct: 3 LLVKD-QFHWNEEADRAFSQLKLALCQAPVLGLPDFNSSFVVETDASGIGMGAILS 167
>BM521770
Length = 428
Score = 30.0 bits (66), Expect = 4.5
Identities = 15/42 (35%), Positives = 23/42 (54%), Gaps = 2/42 (4%)
Frame = +2
Query: 444 LGKGIPE--HARAHLIACLWLNTDLFARSITDMPDIDPSVAC 483
LG IP+ + HL+ LWL+ DL ++ + +ID V C
Sbjct: 41 LGVAIPQAQEVQCHLLLLLWLDLDLPIAEVSGVEEID*GVHC 166
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.335 0.146 0.469
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 38,310,870
Number of Sequences: 63676
Number of extensions: 546084
Number of successful extensions: 4034
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 4004
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4034
length of query: 859
length of database: 12,639,632
effective HSP length: 105
effective length of query: 754
effective length of database: 5,953,652
effective search space: 4489053608
effective search space used: 4489053608
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146649.7


