

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146632.13 - phase: 0
(126 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
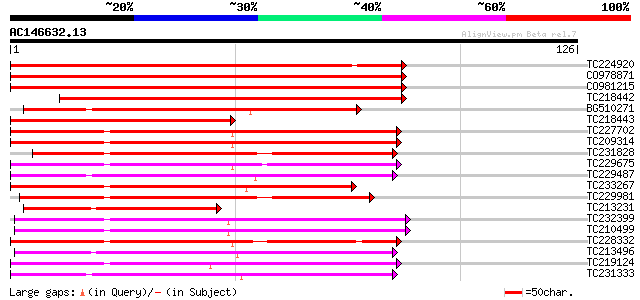
Sequences producing significant alignments: (bits) Value
TC224920 similar to UP|Q9LRT1 (Q9LRT1) Receptor protein kinase, ... 163 2e-41
CO978871 115 4e-27
CO981215 110 2e-25
TC218442 similar to UP|Q70AH8 (Q70AH8) Receptor-like kinase with... 89 7e-19
BG510271 81 1e-16
TC218443 similar to GB|BAB85647.1|19032343|AB076907 inflorescenc... 70 3e-13
TC227702 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial... 67 2e-12
TC209314 similar to UP|Q70CF1 (Q70CF1) Receptor-like protein kin... 67 3e-12
TC231828 similar to UP|Q6YW24 (Q6YW24) SERK1 protein-like, parti... 63 3e-11
TC229675 similar to UP|Q70CF1 (Q70CF1) Receptor-like protein kin... 62 5e-11
TC229487 similar to UP|Q6ZIG4 (Q6ZIG4) Receptor protein kinase P... 60 2e-10
TC233267 homologue to UP|Q7Y229 (Q7Y229) At1g56720, partial (33%) 60 3e-10
TC229981 similar to UP|Q9FGC3 (Q9FGC3) Serine/threonine protein ... 60 3e-10
TC213231 similar to UP|Q9FEU2 (Q9FEU2) Receptor protein kinase, ... 60 3e-10
TC232399 similar to UP|Q39086 (Q39086) Receptor kinase , partial... 60 3e-10
TC210499 weakly similar to UP|Q9LDS6 (Q9LDS6) Serine/threonine k... 60 3e-10
TC228332 similar to UP|Q94KD9 (Q94KD9) At1g01540/F22L4_6, partia... 59 4e-10
TC213496 weakly similar to UP|Q8K2Y2 (Q8K2Y2) Receptor-interacti... 59 8e-10
TC219124 similar to UP|Q9LH71 (Q9LH71) Receptor-like serine/thre... 58 1e-09
TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, parti... 58 1e-09
>TC224920 similar to UP|Q9LRT1 (Q9LRT1) Receptor protein kinase, partial
(26%)
Length = 1039
Score = 163 bits (413), Expect = 2e-41
Identities = 81/88 (92%), Positives = 83/88 (94%)
Frame = +1
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHV 60
MSNRFQSALGYVAPELACQ LRVNEKCDVYGFGVMILE+VTG+RPVEYGEDNVLILNDHV
Sbjct: 457 MSNRFQSALGYVAPELACQSLRVNEKCDVYGFGVMILELVTGRRPVEYGEDNVLILNDHV 636
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVLP 88
RVLLE GN LECVD S MSEYPEDEVLP
Sbjct: 637 RVLLEQGNVLECVDQS-MSEYPEDEVLP 717
>CO978871
Length = 862
Score = 115 bits (289), Expect = 4e-27
Identities = 51/88 (57%), Positives = 69/88 (77%)
Frame = -3
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHV 60
+S++ QSALGY+APE AC+ +++ EKCDVYGFGV++LEIVTGKRPVEY ED+V++L D V
Sbjct: 635 LSSKIQSALGYMAPEFACKTVKITEKCDVYGFGVLVLEIVTGKRPVEYMEDDVVVLCDMV 456
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVLP 88
R LE G EC+D L ++P +E +P
Sbjct: 455 RGALEEGRVEECIDERLQGKFPAEEAIP 372
>CO981215
Length = 584
Score = 110 bits (274), Expect = 2e-25
Identities = 48/88 (54%), Positives = 68/88 (76%)
Frame = -3
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHV 60
+S++ QSALGY+APE AC+ +++ +KCDVYGFG+++LEIVTGKRPVEY ED+V++L D V
Sbjct: 435 LSSKIQSALGYMAPEFACRTVKITKKCDVYGFGILVLEIVTGKRPVEYMEDDVVVLCDMV 256
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVLP 88
R LE G +CVD L+ + +E +P
Sbjct: 255 RGALEEGKVEQCVDGRLLGNFAAEEAIP 172
>TC218442 similar to UP|Q70AH8 (Q70AH8) Receptor-like kinase with LRR
repeats (Fragment), partial (65%)
Length = 611
Score = 88.6 bits (218), Expect = 7e-19
Identities = 41/77 (53%), Positives = 54/77 (69%)
Frame = +3
Query: 12 VAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVRVLLEHGNALE 71
+A E A + +++ EKCDVYGFG LEIVTGKRPVEY ED+V++L D VR LE G E
Sbjct: 3 MAXEFAXKTVKITEKCDVYGFGXXXLEIVTGKRPVEYMEDDVVVLCDMVRGALEEGRVEE 182
Query: 72 CVDPSLMSEYPEDEVLP 88
C+D L ++P +E +P
Sbjct: 183CIDERLQGKFPAEEAIP 233
>BG510271
Length = 420
Score = 80.9 bits (198), Expect = 1e-16
Identities = 43/76 (56%), Positives = 59/76 (77%), Gaps = 1/76 (1%)
Frame = +1
Query: 4 RFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDN-VLILNDHVRV 62
+F +A+GYVAPELA Q LR +EKCDVY FGV++LE+VTG+RPVE N V++L ++V
Sbjct: 193 KFHNAVGYVAPELA-QGLRQSEKCDVYSFGVILLELVTGRRPVESPTTNEVVVLCEYVTG 369
Query: 63 LLEHGNALECVDPSLM 78
LLE G+A +C D +L+
Sbjct: 370 LLETGSASDCFDRNLL 417
>TC218443 similar to GB|BAB85647.1|19032343|AB076907 inflorescence and root
apices receptor-like kinase {Arabidopsis thaliana;} ,
partial (39%)
Length = 1216
Score = 70.1 bits (170), Expect = 3e-13
Identities = 31/50 (62%), Positives = 41/50 (82%)
Frame = +2
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGE 50
+S++ QSALGY+APE AC+ +++ EKCDV V++LEIVTGKRPVEY E
Sbjct: 974 LSSKIQSALGYMAPEFACKTVKITEKCDVLWVRVLVLEIVTGKRPVEYME 1123
>TC227702 similar to UP|Q9LPG0 (Q9LPG0) T3F20.24 protein, partial (21%)
Length = 1142
Score = 67.4 bits (163), Expect = 2e-12
Identities = 38/88 (43%), Positives = 53/88 (60%), Gaps = 1/88 (1%)
Frame = +1
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY-GEDNVLILNDH 59
+S R +GY+APE A + + +K DVY FG++ LEIV+GK Y ++ + L D
Sbjct: 244 ISTRIAGTIGYMAPEYAMRGY-LTDKADVYSFGIVALEIVSGKSNTNYRPKEEFVYLLDW 420
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
VL E GN LE VDPSL S+Y +E +
Sbjct: 421 AYVLQEQGNLLELVDPSLGSKYSSEEAM 504
>TC209314 similar to UP|Q70CF1 (Q70CF1) Receptor-like protein kinase
(Fragment), partial (77%)
Length = 782
Score = 66.6 bits (161), Expect = 3e-12
Identities = 38/88 (43%), Positives = 54/88 (61%), Gaps = 1/88 (1%)
Frame = +2
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY-GEDNVLILNDH 59
+S R +GY+APE A + + +K DVY FGV+ LEIV+GK +Y ++ + L D
Sbjct: 92 ISTRIAGTIGYMAPEYAMRGY-LTDKADVYSFGVVALEIVSGKSNTKYRPKEEFVYLLDW 268
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
VL E GN LE VDP+L S+Y +E +
Sbjct: 269 AYVLQEQGNLLELVDPNLGSKYSPEEAM 352
>TC231828 similar to UP|Q6YW24 (Q6YW24) SERK1 protein-like, partial (42%)
Length = 1530
Score = 63.2 bits (152), Expect = 3e-11
Identities = 32/81 (39%), Positives = 51/81 (62%)
Frame = +2
Query: 6 QSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVRVLLE 65
+ GY+APE + V+EK DV+ FG+++LEIVTG+RPV+ + N+L+ + L+E
Sbjct: 806 EGTFGYLAPEYFMHGI-VDEKTDVFAFGILLLEIVTGRRPVDSSKQNLLL---WAKPLME 973
Query: 66 HGNALECVDPSLMSEYPEDEV 86
GN E DP L +Y +++
Sbjct: 974 SGNIAELADPRLEGKYDGEQL 1036
>TC229675 similar to UP|Q70CF1 (Q70CF1) Receptor-like protein kinase
(Fragment), partial (54%)
Length = 800
Score = 62.4 bits (150), Expect = 5e-11
Identities = 39/89 (43%), Positives = 53/89 (58%), Gaps = 2/89 (2%)
Frame = +1
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY--GEDNVLILND 58
+S R +GY+APE A + + +K DVY FGV+ LE V+GK + ED V +L D
Sbjct: 4 ISTRVAGTIGYMAPEYAMRGY-LTDKADVYSFGVVALETVSGKSNTNFRPNEDFVYLL-D 177
Query: 59 HVRVLLEHGNALECVDPSLMSEYPEDEVL 87
VL E G+ LE VDP+L SEY +E +
Sbjct: 178WAYVLQERGSLLELVDPNLGSEYLTEEAM 264
>TC229487 similar to UP|Q6ZIG4 (Q6ZIG4) Receptor protein kinase PERK1-like
protein, partial (53%)
Length = 1004
Score = 60.5 bits (145), Expect = 2e-10
Identities = 33/87 (37%), Positives = 49/87 (55%), Gaps = 1/87 (1%)
Frame = +1
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNV-LILNDH 59
++ R + LGY+APE A + + NE CDVY FG+++LE+ +GK+P+E V +ND
Sbjct: 235 VTTRVKGTLGYLAPEYA-MLGKANESCDVYSFGILLLELASGKKPLEKLSSAVKRSINDW 411
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEV 86
L E DP L Y E+E+
Sbjct: 412 ALPLACEKKFSELADPKLEGNYAEEEL 492
>TC233267 homologue to UP|Q7Y229 (Q7Y229) At1g56720, partial (33%)
Length = 637
Score = 60.1 bits (144), Expect = 3e-10
Identities = 33/78 (42%), Positives = 51/78 (65%), Gaps = 1/78 (1%)
Frame = +3
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGED-NVLILNDH 59
++ R GYVAPE A L +NEK DVY FGV++LE +TG+ PV+YG N + L D
Sbjct: 114 VTTRVMGTFGYVAPEYANTGL-LNEKSDVYSFGVLLLEGITGRDPVDYGRPANEVNLVDW 290
Query: 60 VRVLLEHGNALECVDPSL 77
+++++ + + E VDP++
Sbjct: 291 LKMMVGNRRSEEVVDPNI 344
>TC229981 similar to UP|Q9FGC3 (Q9FGC3) Serine/threonine protein kinase-like,
partial (42%)
Length = 801
Score = 59.7 bits (143), Expect = 3e-10
Identities = 28/79 (35%), Positives = 53/79 (66%)
Frame = +3
Query: 3 NRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGEDNVLILNDHVRV 62
++F+ GY+APE + V+EK DV+ FGV++LE+V+G+R ++Y + ++++ +
Sbjct: 216 SKFEGTFGYLAPEYLLHGI-VDEKTDVFAFGVLLLELVSGRRALDYSQQSLVL---WAKP 383
Query: 63 LLEHGNALECVDPSLMSEY 81
LL+ + +E VDPSL ++
Sbjct: 384 LLKKNDIMELVDPSLAGDF 440
>TC213231 similar to UP|Q9FEU2 (Q9FEU2) Receptor protein kinase, partial (9%)
Length = 830
Score = 59.7 bits (143), Expect = 3e-10
Identities = 26/44 (59%), Positives = 36/44 (81%)
Frame = +2
Query: 4 RFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVE 47
+F + GY+APE AC +L++ EK DVY FGV++LEI+TGKRPV+
Sbjct: 83 QFAGSYGYIAPEYAC-MLKITEKSDVYSFGVVLLEIITGKRPVD 211
>TC232399 similar to UP|Q39086 (Q39086) Receptor kinase , partial (23%)
Length = 1167
Score = 59.7 bits (143), Expect = 3e-10
Identities = 33/89 (37%), Positives = 49/89 (54%), Gaps = 1/89 (1%)
Frame = +2
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVE-YGEDNVLILNDHV 60
+NR GY++PE A Q L +EK DV+ FGV+++EIV+G+R Y +DN L L
Sbjct: 542 TNRVVGTYGYMSPEYAMQGL-FSEKSDVFSFGVLVIEIVSGRRNSRFYDDDNALSLLGFA 718
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVLPC 89
+ GN L +DP + ++L C
Sbjct: 719 WIQWREGNILSVIDPEIYDVTHHKDILRC 805
>TC210499 weakly similar to UP|Q9LDS6 (Q9LDS6) Serine/threonine kinase-like
protein , partial (27%)
Length = 717
Score = 59.7 bits (143), Expect = 3e-10
Identities = 34/89 (38%), Positives = 51/89 (57%), Gaps = 1/89 (1%)
Frame = +3
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVE-YGEDNVLILNDHV 60
++R GY++PE A + + K DVY FGV++LEIV+G++ Y D++L L H
Sbjct: 210 TSRIVGTYGYMSPEYAMEGT-FSTKSDVYSFGVLLLEIVSGRKNTSFYDVDHLLNLIGHA 386
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEVLPC 89
L G +L+ +DPSL + DEV C
Sbjct: 387 WELWNQGESLQLLDPSLNDSFDPDEVKRC 473
>TC228332 similar to UP|Q94KD9 (Q94KD9) At1g01540/F22L4_6, partial (42%)
Length = 977
Score = 59.3 bits (142), Expect = 4e-10
Identities = 33/91 (36%), Positives = 56/91 (61%), Gaps = 4/91 (4%)
Frame = +1
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEY----GEDNVLIL 56
++ R GYVAPE AC + + EK DVY FG++I+E++TG+ PV+Y GE N++
Sbjct: 229 VTTRVMGTFGYVAPEYACTGM-LTEKSDVYSFGILIMELITGRSPVDYSKPQGEVNLI-- 399
Query: 57 NDHVRVLLEHGNALECVDPSLMSEYPEDEVL 87
+ ++ ++ + + E VDP + +E P + L
Sbjct: 400 -EWLKSMVGNRKSEEVVDPKI-AEKPSSKAL 486
>TC213496 weakly similar to UP|Q8K2Y2 (Q8K2Y2) Receptor-interacting
serine-threonine kinase 3, partial (7%)
Length = 760
Score = 58.5 bits (140), Expect = 8e-10
Identities = 34/86 (39%), Positives = 47/86 (54%), Gaps = 1/86 (1%)
Frame = +1
Query: 2 SNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYG-EDNVLILNDHV 60
+ R LGY+APELA + DVY FGV++LE+ G+RP+E + ++L D V
Sbjct: 268 TTRVVGTLGYLAPELAT-VAAPTSATDVYSFGVVLLEVACGRRPIETSVAEEEVVLIDWV 444
Query: 61 RVLLEHGNALECVDPSLMSEYPEDEV 86
R L G A E D + EY E +V
Sbjct: 445 RELYAKGCAREAADLRIRGEYDEGDV 522
>TC219124 similar to UP|Q9LH71 (Q9LH71) Receptor-like serine/threonine
kinase, partial (15%)
Length = 871
Score = 57.8 bits (138), Expect = 1e-09
Identities = 35/88 (39%), Positives = 49/88 (54%), Gaps = 1/88 (1%)
Frame = +3
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGK-RPVEYGEDNVLILNDH 59
+S R GY+APE + +K DVY FGV+ LEIV+GK + + L L D
Sbjct: 132 ISTRVAGTYGYMAPEYEMHGY-LTDKADVYSFGVVALEIVSGKSNTIHRSKQEALHLLDW 308
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEVL 87
+L E GN +E VD L S++ E+EV+
Sbjct: 309 AHLLKEKGNLMELVDRRLGSDFNENEVM 392
>TC231333 similar to UP|Q8L741 (Q8L741) AT3g13690/MMM17_12, partial (38%)
Length = 1009
Score = 57.8 bits (138), Expect = 1e-09
Identities = 33/87 (37%), Positives = 50/87 (56%), Gaps = 1/87 (1%)
Frame = +3
Query: 1 MSNRFQSALGYVAPELACQILRVNEKCDVYGFGVMILEIVTGKRPVEYGE-DNVLILNDH 59
+ R GY+APE A Q ++ EK DVY FG+++LE+VTG++ V+ L++
Sbjct: 513 VETRVIGTFGYLAPEYA-QSGQITEKADVYSFGIVLLELVTGRKAVDINRPKGQQCLSEW 689
Query: 60 VRVLLEHGNALECVDPSLMSEYPEDEV 86
R LLE + +DPSL + Y + EV
Sbjct: 690 ARPLLEKQATYKLIDPSLRNCYVDQEV 770
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.141 0.423
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,591,537
Number of Sequences: 63676
Number of extensions: 36550
Number of successful extensions: 716
Number of sequences better than 10.0: 502
Number of HSP's better than 10.0 without gapping: 651
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 652
length of query: 126
length of database: 12,639,632
effective HSP length: 86
effective length of query: 40
effective length of database: 7,163,496
effective search space: 286539840
effective search space used: 286539840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 53 (25.0 bits)
Medicago: description of AC146632.13


