

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146631.5 - phase: 0 /pseudo
(359 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
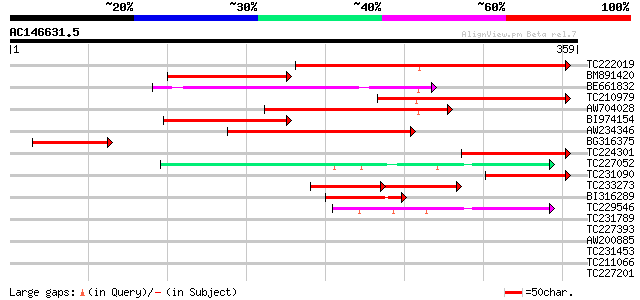
Score E
Sequences producing significant alignments: (bits) Value
TC222019 similar to UP|Q6K4N7 (Q6K4N7) Amino acid transporter-li... 285 2e-77
BM891420 similar to GP|20197120|gb| expressed protein {Arabidops... 132 2e-31
BE661832 130 7e-31
TC210979 123 1e-28
AW704028 114 9e-26
BI974154 104 5e-23
AW234346 100 8e-22
BG316375 77 2e-14
TC224301 weakly similar to UP|Q9MBG9 (Q9MBG9) Gb|AAC79623.2, par... 67 1e-11
TC227052 weakly similar to GB|AAM19768.1|20453045|AY094388 AT3g5... 67 1e-11
TC231090 62 5e-10
TC233273 60 2e-09
BI316289 52 3e-07
TC229546 weakly similar to GB|AAM19768.1|20453045|AY094388 AT3g5... 44 9e-05
TC231789 similar to GB|AAN31090.1|23506157|AY149936 At1g80510/T2... 37 0.011
TC227393 similar to UP|Q9FMC0 (Q9FMC0) Amino acid transporter-li... 35 0.040
AW200885 similar to GP|15529155|gb AT3g30390/T6J22_16 {Arabidops... 35 0.040
TC231453 weakly similar to UP|Q85V22 (Q85V22) Histidine amino ac... 34 0.12
TC211066 31 0.76
TC227201 weakly similar to GB|AAM19768.1|20453045|AY094388 AT3g5... 30 1.3
>TC222019 similar to UP|Q6K4N7 (Q6K4N7) Amino acid transporter-like, partial
(84%)
Length = 979
Score = 285 bits (730), Expect = 2e-77
Identities = 136/176 (77%), Positives = 155/176 (87%), Gaps = 2/176 (1%)
Frame = +1
Query: 182 VEFITLEGDNLTGLFPGTSLDIGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGG 241
+EFITLEGDNLTGLFPGTSLD+G LDS+HLFG+L AL+I+PTVWLKDLR+IS LS GG
Sbjct: 19 LEFITLEGDNLTGLFPGTSLDLGSFRLDSVHLFGILAALIIIPTVWLKDLRIISILSAGG 198
Query: 242 IAATILIIISVFSVGTT--VGFHHTGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMAD 299
+ AT+LI++ VF VGT VGFHHTG++VNWSGIP AIG++GFCFAGHSVFPNIYQSMAD
Sbjct: 199 VFATLLIVVCVFCVGTINGVGFHHTGQLVNWSGIPLAIGIHGFCFAGHSVFPNIYQSMAD 378
Query: 300 KKQYTKALITCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGAFASKVALWTT 355
K+Q+TKALI CFVL I IYG VA+MGFL FG +TLSQITLNMP AFASKVALWTT
Sbjct: 379 KRQFTKALIICFVLSITIYGGVAIMGFLMFGGETLSQITLNMPRDAFASKVALWTT 546
>BM891420 similar to GP|20197120|gb| expressed protein {Arabidopsis
thaliana}, partial (12%)
Length = 433
Score = 132 bits (333), Expect = 2e-31
Identities = 59/78 (75%), Positives = 71/78 (90%)
Frame = +2
Query: 101 NVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFESREGLTSYPDIGEAA 160
NVMAGVG+LSTPYT+K+AGWM +VLM++FA +CCYTATLMR+CFESREG+TSYPDIGEAA
Sbjct: 176 NVMAGVGILSTPYTLKEAGWMSMVLMVLFAVICCYTATLMRYCFESREGITSYPDIGEAA 355
Query: 161 FGRYGRIFVSIILYTELY 178
FG+YGRI VS+ L +Y
Sbjct: 356 FGKYGRIIVSVSLSIFIY 409
>BE661832
Length = 635
Score = 130 bits (328), Expect = 7e-31
Identities = 70/182 (38%), Positives = 103/182 (56%), Gaps = 2/182 (1%)
Frame = +1
Query: 91 SYTQTVFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFESREGL 150
S+ T FNGIN + S PYT+ GW+ L L+ A+ YT L++ + +
Sbjct: 121 SFFHTCFNGINAV------SVPYTLASGGWLSLALLFAIAAATFYTGVLIKR*MDKDSNI 282
Query: 151 TSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDIGGLHLDS 210
+YPD+GE AFG+ GR+ +S ++YTEL+ V F+ LEGDNL+ LFP + L +
Sbjct: 283 RTYPDMGELAFGKTGRLIISGLIYTELFLVSVGFLILEGDNLSNLFPTVEIHTADLAIGG 462
Query: 211 MHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGT--TVGFHHTGRVV 268
LF +L ALV L +LR++SY+S + A+ +II+S+ T VGFH G +
Sbjct: 463 KKLFVILVALV------LDNLRILSYVSASRVFASAIIILSISWTATFDGVGFHQXGTLG 624
Query: 269 NW 270
NW
Sbjct: 625 NW 630
>TC210979
Length = 928
Score = 123 bits (308), Expect = 1e-28
Identities = 57/126 (45%), Positives = 85/126 (67%), Gaps = 4/126 (3%)
Frame = +2
Query: 234 ISYLSVGGIAATILIIISVFSVG----TTVGFHHTGRVVNWSGIPFAIGVYGFCFAGHSV 289
+SY+S G+ ATIL+++ +F VG + T + N + P AIG+YG+C+AGH+V
Sbjct: 2 LSYISACGVIATILVVLCLFWVGLLDNADIHTQGTTKTFNLATFPVAIGLYGYCYAGHAV 181
Query: 290 FPNIYQSMADKKQYTKALITCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGAFASK 349
F NIY +MA++ Q+ L+ CF +C +Y +VA+MG+ +FG TLSQ TLNMP A+K
Sbjct: 182 FSNIYTAMANRNQFPGVLLVCFAICTSMYCAVAIMGYTAFGKATLSQYTLNMPQHLVATK 361
Query: 350 VALWTT 355
+A+WTT
Sbjct: 362 IAVWTT 379
>AW704028
Length = 373
Score = 114 bits (284), Expect = 9e-26
Identities = 56/121 (46%), Positives = 78/121 (64%), Gaps = 2/121 (1%)
Frame = +3
Query: 162 GRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDIGGLHLDSMHLFGVLTALV 221
G+ GR+ +S +YTELY + F+ LEGDNL L P + I G + LF +L AL+
Sbjct: 6 GKIGRLIISGYMYTELYLVSIGFLILEGDNLNNLCPIEEVQIAGFVIGGKQLFVILVALI 185
Query: 222 ILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGT--TVGFHHTGRVVNWSGIPFAIGV 279
ILPTVWL +L ++SY+S G+ A+ +II+S+ GT VGFH G +VNW GIP A+ +
Sbjct: 186 ILPTVWLDNLSMLSYVSASGVFASAVIILSISWTGTFDGVGFHQKGTLVNWRGIPTAVSL 365
Query: 280 Y 280
Y
Sbjct: 366 Y 368
>BI974154
Length = 420
Score = 104 bits (260), Expect = 5e-23
Identities = 47/81 (58%), Positives = 60/81 (74%)
Frame = +2
Query: 98 NGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFESREGLTSYPDIG 157
NGINV+ GVG+LSTPY K GW+GL +++IFA + YT L+R C +S L +YPDIG
Sbjct: 5 NGINVLCGVGILSTPYAAKVGGWLGLSILVIFAIISFYTGLLLRSCLDSEPELETYPDIG 184
Query: 158 EAAFGRYGRIFVSIILYTELY 178
+AAFG GRI +SI+LY ELY
Sbjct: 185 QAAFGTTGRIAISIVLYVELY 247
>AW234346
Length = 371
Score = 100 bits (250), Expect = 8e-22
Identities = 49/119 (41%), Positives = 76/119 (63%)
Frame = +2
Query: 139 LMRHCFESREGLTSYPDIGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPG 198
L++ C + + ++PDIG+ AFG GRI VSI + +EL+ F+ LEGDNL L P
Sbjct: 11 LVKRCMDMDPDIKNFPDIGQRAFGDKGRIIVSIAMNSELFLVVTGFLILEGDNLNKLVPN 190
Query: 199 TSLDIGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGT 257
L++ GL + +F ++ ALVILPTV L+DL ++SY+S G A+ + ++S++ GT
Sbjct: 191 MQLELAGLTIGGTTIFTMIAALVILPTVLLEDLSMLSYVSASGALASSIFLLSIYWNGT 367
>BG316375
Length = 427
Score = 76.6 bits (187), Expect = 2e-14
Identities = 37/51 (72%), Positives = 43/51 (83%)
Frame = +1
Query: 15 RETTDSYTIATAPNFVSILRGPSSIYSSFSNRSKSDLDIDGKTPCLSGLDG 65
+ETTDSYT+A PNF SILR PS IYSSF +RSK++LDIDGKTP LSG +G
Sbjct: 226 KETTDSYTLAATPNFESILRVPSIIYSSFESRSKNNLDIDGKTPFLSGHEG 378
Score = 37.4 bits (85), Expect = 0.011
Identities = 14/17 (82%), Positives = 16/17 (93%)
Frame = +1
Query: 272 GIPFAIGVYGFCFAGHS 288
GIP AIG++GFCFAGHS
Sbjct: 376 GIPLAIGIHGFCFAGHS 426
>TC224301 weakly similar to UP|Q9MBG9 (Q9MBG9) Gb|AAC79623.2, partial (29%)
Length = 667
Score = 67.4 bits (163), Expect = 1e-11
Identities = 29/69 (42%), Positives = 43/69 (62%)
Frame = +2
Query: 287 HSVFPNIYQSMADKKQYTKALITCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGAF 346
H + Y SM DK Q+++ CF +C L Y + V+ +L FG + SQ+TLN+P G F
Sbjct: 2 HPIXXTXYNSMRDKSQFSRXXSICFSVCXLGYAAAGVLXYLMFGQEVESQVTLNLPTGKF 181
Query: 347 ASKVALWTT 355
+S VA++TT
Sbjct: 182 SSHVAIFTT 208
>TC227052 weakly similar to GB|AAM19768.1|20453045|AY094388
AT3g56200/F18O21_160 {Arabidopsis thaliana;} , partial
(68%)
Length = 1627
Score = 67.0 bits (162), Expect = 1e-11
Identities = 64/278 (23%), Positives = 110/278 (39%), Gaps = 28/278 (10%)
Frame = +3
Query: 96 VFNGINVMAGVGLLSTPYTVKQAGWMGLVLMLIFASVCCYTATLMRHCFESREGLTSYPD 155
VFN + G G++S P +K G + M++ +V + F T+Y
Sbjct: 159 VFNVATSIVGAGIMSIPAIMKVLGVVPAFAMILVVAVLAELSVDFLMRFTHSGETTTYAG 338
Query: 156 IGEAAFGRYGRIFVSIILYTELYSYCVEFITLEGDNLTGLFPGTSLDIG------GLHLD 209
+ AFG G + + + + ++ + GD L+G G + +G G+H
Sbjct: 339 VMREAFGSGGALAAQVCVIITNVGGLILYLIIIGDVLSGKQNGGEVHLGILQQWFGIHWW 518
Query: 210 SMHLFGVLTALV--ILPTVWLKDLRVISYLSVGGIAATILIIISVFSVGTTVGFHHTGRV 267
+ F +L LV +LP V K + + Y S + +++V VG G T V
Sbjct: 519 NSREFALLFTLVFVMLPLVLYKRVESLKYSSA------VSTLLAVAFVGICCGLAITALV 680
Query: 268 VN--------------------WSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKAL 307
++ +P + + F F H I +A Q T A+
Sbjct: 681 QGKTQTPRLFPRLDYQTSFFDLFTAVPVVVTAFTFHFNVHP----IGFELAKASQMTTAV 848
Query: 308 ITCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGA 345
+LC +IY ++ + G++ FGD T S I +N A
Sbjct: 849 RLALLLCAVIYLAIGLFGYMLFGDSTQSDILINFDQNA 962
>TC231090
Length = 559
Score = 61.6 bits (148), Expect = 5e-10
Identities = 26/54 (48%), Positives = 36/54 (66%)
Frame = +2
Query: 302 QYTKALITCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGAFASKVALWTT 355
Q+ L+ CF +C L+Y AV+G+ FG+ LSQ TLNMP A+K+A+WTT
Sbjct: 2 QFPGVLLACFGICTLLYAGAAVLGYTMFGEAILSQFTLNMPKELVATKIAVWTT 163
>TC233273
Length = 601
Score = 60.1 bits (144), Expect = 2e-09
Identities = 25/48 (52%), Positives = 37/48 (77%)
Frame = +1
Query: 191 NLTGLFPGTSLDIGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLS 238
+L+ LFP L++GG+ L+S LF V+T L +LPTVWL+DL ++SY+S
Sbjct: 4 SLSSLFPSAHLNLGGIELNSHTLFAVITTLAVLPTVWLRDLSILSYIS 147
Score = 47.0 bits (110), Expect = 1e-05
Identities = 19/52 (36%), Positives = 34/52 (64%), Gaps = 1/52 (1%)
Frame = +1
Query: 236 YLSVGGIAATILIIISVFSVGTT-VGFHHTGRVVNWSGIPFAIGVYGFCFAG 286
++ GG+ A+IL++ + VG VGFH G +N + +P +G+YG+C++G
Sbjct: 445 FVIAGGVVASILVVXCLLWVGIEDVGFHSKGTTLNLATLPVDVGLYGYCYSG 600
>BI316289
Length = 412
Score = 52.4 bits (124), Expect = 3e-07
Identities = 24/51 (47%), Positives = 37/51 (72%)
Frame = +3
Query: 201 LDIGGLHLDSMHLFGVLTALVILPTVWLKDLRVISYLSVGGIAATILIIIS 251
L++GG+ L+S LF V+T L +LPTVWL+DL ++SY+S G + ++IS
Sbjct: 3 LNLGGIELNSRTLFAVITTLAVLPTVWLRDLSILSYIS-GKSLFLVQVVIS 152
>TC229546 weakly similar to GB|AAM19768.1|20453045|AY094388
AT3g56200/F18O21_160 {Arabidopsis thaliana;} , partial
(83%)
Length = 1336
Score = 44.3 bits (103), Expect = 9e-05
Identities = 39/157 (24%), Positives = 65/157 (40%), Gaps = 16/157 (10%)
Frame = +3
Query: 205 GLHLDSMHLFGVLTAL--VILPTVWLKDLRVISYLSVGG--IAATILIIISVFSVGTTVG 260
G+H S F +L L ++LP V + + + + S +A + I +V ++ V
Sbjct: 366 GIHWWSSREFALLVVLFLILLPLVLYRRVESLKFSSAISTLLAVAFVTICTVLAIVAIVE 545
Query: 261 FH------------HTGRVVNWSGIPFAIGVYGFCFAGHSVFPNIYQSMADKKQYTKALI 308
HT ++ +P + Y F F H I +A + A+
Sbjct: 546 GRTQSPRLIPCLDQHTSFFDLFTAVPVVVTAYTFHFNVHP----IGFELAKPSEMATAVR 713
Query: 309 TCFVLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGA 345
+LC +IY S+ + G+L FGD T S I +N A
Sbjct: 714 IALLLCCVIYFSIGLSGYLLFGDSTQSDILVNFDQNA 824
>TC231789 similar to GB|AAN31090.1|23506157|AY149936 At1g80510/T21F11_16
{Arabidopsis thaliana;} , partial (42%)
Length = 1044
Score = 37.4 bits (85), Expect = 0.011
Identities = 15/29 (51%), Positives = 20/29 (68%)
Frame = +2
Query: 312 VLCILIYGSVAVMGFLSFGDDTLSQITLN 340
+LCIL+Y S A+ G+L FG DT S + N
Sbjct: 110 ILCILVYSSTAISGYLLFGKDTESDVLTN 196
>TC227393 similar to UP|Q9FMC0 (Q9FMC0) Amino acid transporter-like protein,
partial (47%)
Length = 1065
Score = 35.4 bits (80), Expect = 0.040
Identities = 18/64 (28%), Positives = 29/64 (45%)
Frame = +3
Query: 277 IGVYGFCFAGHSVFPNIYQSMADKKQYTKALITCFVLCILIYGSVAVMGFLSFGDDTLSQ 336
+ V+ + H +I + D Q + T VLC +Y ++ GFL FG+ TL
Sbjct: 69 VPVFVTAYICHYNVHSIDNELEDSSQMQGVVQTALVLCSSVYVMISFFGFLLFGEGTLDD 248
Query: 337 ITLN 340
+ N
Sbjct: 249 VLAN 260
>AW200885 similar to GP|15529155|gb AT3g30390/T6J22_16 {Arabidopsis
thaliana}, partial (24%)
Length = 404
Score = 35.4 bits (80), Expect = 0.040
Identities = 18/58 (31%), Positives = 30/58 (51%), Gaps = 6/58 (10%)
Frame = +2
Query: 299 DKKQYTKALITCFVLCILIYGSVAVMGFLSFGDDTLSQITLN------MPAGAFASKV 350
D+ Q + T +LC +Y + ++ GF FGD+TL I N +P G+F + +
Sbjct: 5 DRSQMKAIVRTSLLLCSSVYIATSLFGFFLFGDNTLDDILANFDGDLGVPYGSFLTDI 178
>TC231453 weakly similar to UP|Q85V22 (Q85V22) Histidine amino acid
transporter, partial (51%)
Length = 811
Score = 33.9 bits (76), Expect = 0.12
Identities = 24/81 (29%), Positives = 37/81 (45%), Gaps = 8/81 (9%)
Frame = +1
Query: 276 AIGVYGFCFAGHSVFPNIYQSM------ADKKQYTKALITCFVLCILIYGSVAVMGFLSF 329
A+G F FAGH+V I ++ K K I +V+ + Y VA++G+ +F
Sbjct: 121 ALGQISFAFAGHAVALEIQATIPSTPEKPSKIPMWKGAIGAYVINAICYFPVALVGYWAF 300
Query: 330 GDDTLSQITLNM--PAGAFAS 348
G D + + PA AS
Sbjct: 301 GRDVEDNVLMEFERPAWLIAS 363
>TC211066
Length = 805
Score = 31.2 bits (69), Expect = 0.76
Identities = 13/22 (59%), Positives = 16/22 (72%)
Frame = +3
Query: 334 LSQITLNMPAGAFASKVALWTT 355
LSQ TLNMP A+ +A+WTT
Sbjct: 51 LSQFTLNMPKELVATNIAVWTT 116
>TC227201 weakly similar to GB|AAM19768.1|20453045|AY094388
AT3g56200/F18O21_160 {Arabidopsis thaliana;} , partial
(35%)
Length = 925
Score = 30.4 bits (67), Expect = 1.3
Identities = 10/37 (27%), Positives = 20/37 (54%)
Frame = +3
Query: 312 VLCILIYGSVAVMGFLSFGDDTLSQITLNMPAGAFAS 348
++C+ IY ++ G+L FGD + + +N + S
Sbjct: 162 IICVAIYFAIGFFGYLLFGDSIMPDVLVNFDQNSHTS 272
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.140 0.433
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,281,779
Number of Sequences: 63676
Number of extensions: 241603
Number of successful extensions: 1728
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 1706
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1721
length of query: 359
length of database: 12,639,632
effective HSP length: 98
effective length of query: 261
effective length of database: 6,399,384
effective search space: 1670239224
effective search space used: 1670239224
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146631.5


