

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146631.2 + phase: 0 /pseudo
(953 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
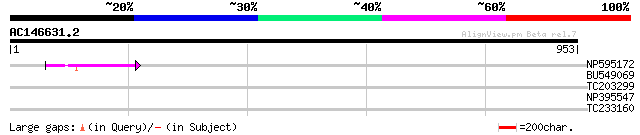
NP595172 polyprotein [Glycine max] 56 9e-08
BU549069 37 0.032
TC203299 weakly similar to PDB|1CBG.0|1311386|1CBG Cyanogenic Be... 31 3.0
NP395547 reverse transcriptase [Glycine max] 30 6.6
TC233160 similar to UP|Q8S976 (Q8S976) Auxin response factor 10 ... 29 8.6
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 55.8 bits (133), Expect = 9e-08
Identities = 54/164 (32%), Positives = 82/164 (49%), Gaps = 4/164 (2%)
Frame = +2
Query: 61 YCWLRRRMVA*GCV*ITVS*IK*R*RIGIHFRGLTI*WIS*WVHVCLVKL----I*GQVT 116
YC LRR+M I *++ *RI + W++ ++ + + I*GQ T
Sbjct: 1910 YCLLRRKMEVGDSAQIIEL*MQSL*RIAFQCQQ----WMNC*MNYMELNIFQNWI*GQDT 2077
Query: 117 IRLR*RMRICRRRPLGHVMVTMNIR*CLSVILMRLLCLWSI*TEFSMLIWINLWLCLLMT 176
I+ +RI R+ L H+M MN *C V+LM + *T++ L+ NL+LC LMT
Sbjct: 2078 IKFWYSLRIERKLLLEHIMAIMNG**CHLVLLMHQPHSNA**TKYFSLLSENLFLCFLMT 2257
Query: 177 S*FILRLKKSMQNI*GLCCKC*KKRSCMPSCLNVSLVK*SKFSW 220
S*+I+ L + + + LC + * S + C NV L +W
Sbjct: 2258 S*YIVLLGRIISSTWNLCYRP*SNISYLLDCPNVHLEIQKLITW 2389
Score = 38.5 bits (88), Expect = 0.014
Identities = 45/188 (23%), Positives = 74/188 (38%)
Frame = +3
Query: 564 CCSFCLCMFDLSKV*G*ASEACSVVDTVRCARAEVGDHFHGFCDEFTEHSERA*CHLGCC 623
C S M DLS * A T + +G HGF FT+ CH G
Sbjct: 3324 CQSLYTEMLDLSTS*IKQYTASRTTPTFTYSSTSLGRCSHGFHHRFTKFLWII-CHNGSH 3500
Query: 624 RQVDEVGTLYSDQC*LSGSSVGGDLYS*YCEVAWSSVEYCVGKRPEVHFKILEEFARCIG 683
++ + +S + *L +Y + ++ W + Y V R + L F +
Sbjct: 3501 **THQICSFHSIKS*L*Q*GGSRGIYESHSQITWDTKIYSV*SR*GFYQYFLATFIQVTR 3680
Query: 684 FKVEVEFGLSSANRWSVGENDSIVRGFAKRMCT*ARWSLG*SPTVDRVHVQQQLPFEYWY 743
+ + LSS RW++ + + + + * LG T+ RV V + +E +
Sbjct: 3681 YYIGHVISLSSTVRWTI*GAE*MFGDVS*MLHV*TS*RLGKGITMGRVLV*HCISYELRH 3860
Query: 744 GTVQSFVW 751
++QS VW
Sbjct: 3861 DSIQSLVW 3884
>BU549069
Length = 615
Score = 37.4 bits (85), Expect = 0.032
Identities = 38/134 (28%), Positives = 67/134 (49%)
Frame = -3
Query: 817 ESYSYDRCRTCIEV*EVDS*VY*SISDI*ESWNSGI*SGFATTSFEFA*CVSCVATSEVC 876
ES+S D + E+ + + +Y S + *+S + GI + +F + C+SCV+T V
Sbjct: 613 ESHSKDWGWSSTEIPKTHTSLYRSFPNS*KSXSCGIPNCITPITF*SSQCLSCVSTPYVY 434
Query: 877 SGSFACYSNR*CAG*RQPYS*DLTVENQRP*GEEVERQGYTSSEGCLGWSNR*KFDLGAG 936
S +C * R+ *++T+E++ + +++G + G LG R + +G
Sbjct: 433 P*SISCGQIG*RTSKRELDI*NITIEDRG*KNKAPKKEGESIG*GDLGRYIRRRCHVGIR 254
Query: 937 E*DAGVLSRVVCLR 950
E DA LS V +R
Sbjct: 253 ESDASSLSIFV*VR 212
>TC203299 weakly similar to PDB|1CBG.0|1311386|1CBG Cyanogenic
Beta-Glucosidase Mol_id: 1; Molecule: Cyanogenic
Beta-Glucosidase; Chain: Null; Ec: 3.2.1.21. {Trifolium
repens;} , partial (80%)
Length = 1781
Score = 30.8 bits (68), Expect = 3.0
Identities = 15/36 (41%), Positives = 18/36 (49%), Gaps = 1/36 (2%)
Frame = +3
Query: 194 CCKC*-KKRSCMPSCLNVSLVK*SKFSWPHYFW*WY 228
CCKC KKRS P C N+ FS+ W W+
Sbjct: 81 CCKCNSKKRSAKPPCFNIQQ---KPFSFHFSLWNWF 179
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 29.6 bits (65), Expect = 6.6
Identities = 20/50 (40%), Positives = 30/50 (60%)
Frame = +2
Query: 114 QVTIRLR*RMRICRRRPLGHVMVTMNIR*CLSVILMRLLCLWSI*TEFSM 163
QVTIRL+ +RI +++ L + V + I C SV +M LL +* +F M
Sbjct: 290 QVTIRLQWILRIKKKQLLHVLSVFLLIAACRSVYVMPLLLFRDV*WQFLM 439
>TC233160 similar to UP|Q8S976 (Q8S976) Auxin response factor 10 (Fragment),
partial (26%)
Length = 658
Score = 29.3 bits (64), Expect = 8.6
Identities = 20/54 (37%), Positives = 27/54 (49%), Gaps = 7/54 (12%)
Frame = +1
Query: 309 FTEAGRTFCCLL*CV*VGLGW---CFDARW*SGSLW----F*AVEDS*KELSYT 355
F E+ FC L CV G W C+D+ G+LW F VE ++LSY+
Sbjct: 139 FCESQEAFCWGLNCVFEGRKWGFVCWDSTSEEGNLWWT*DFLRVESCRRKLSYS 300
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.373 0.168 0.680
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 48,039,602
Number of Sequences: 63676
Number of extensions: 758399
Number of successful extensions: 9388
Number of sequences better than 10.0: 10
Number of HSP's better than 10.0 without gapping: 3529
Number of HSP's successfully gapped in prelim test: 299
Number of HSP's that attempted gapping in prelim test: 5690
Number of HSP's gapped (non-prelim): 4168
length of query: 953
length of database: 12,639,632
effective HSP length: 106
effective length of query: 847
effective length of database: 5,889,976
effective search space: 4988809672
effective search space used: 4988809672
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (22.0 bits)
S2: 64 (29.3 bits)
Medicago: description of AC146631.2


