

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146588.5 - phase: 0 /pseudo
(637 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
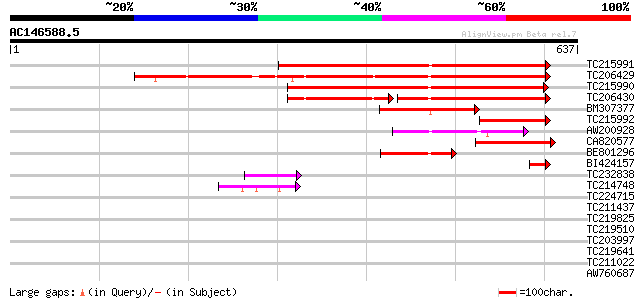
Score E
Sequences producing significant alignments: (bits) Value
TC215991 495 e-140
TC206429 similar to UP|Q9LFQ3 (Q9LFQ3) Transcriptional regulator... 495 e-140
TC215990 479 e-135
TC206430 similar to UP|Q9LFQ3 (Q9LFQ3) Transcriptional regulator... 233 2e-61
BM307377 168 6e-42
TC215992 137 2e-32
AW200928 116 3e-26
CA820577 weakly similar to PIR|T51447|T51 transcription regulato... 115 8e-26
BE801296 80 2e-15
BI424157 similar to PIR|T51447|T51 transcription regulator-like ... 44 2e-04
TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {A... 43 4e-04
TC214748 homologue to UP|Q39882 (Q39882) Nodulin-26, partial (98%) 42 8e-04
TC224715 homologue to UP|Q39882 (Q39882) Nodulin-26, partial (98%) 41 0.002
TC211437 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%) 40 0.003
TC219825 weakly similar to UP|MID2_YEAST (P36027) Mating process... 39 0.007
TC219510 UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetylase HD2-P3... 39 0.007
TC203997 similar to UP|Q9FFJ6 (Q9FFJ6) Phosphate/phosphoenolpyru... 38 0.016
TC219641 similar to UP|Q6VBJ3 (Q6VBJ3) Epa4p, partial (6%) 38 0.016
TC211022 similar to UP|O81600 (O81600) H-protein promoter bindin... 36 0.046
AW760687 36 0.046
>TC215991
Length = 1356
Score = 495 bits (1275), Expect = e-140
Identities = 248/307 (80%), Positives = 275/307 (88%), Gaps = 2/307 (0%)
Frame = +1
Query: 303 CSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSERFLLTAKPLVKYVSPVFH 362
CSREEHE DG+HDNK ESEGEAEG+ADA+DVEGDG SLPYSERFLLT KPL K+V P+ H
Sbjct: 1 CSREEHE-DGEHDNKAESEGEAEGIADAHDVEGDGMSLPYSERFLLTVKPLAKHVPPMLH 177
Query: 363 GKEENVQIFYGNDSFYVLFRLHQTLYERIRSAKINSSSAEKKWRASNDTSSTDRYARFMN 422
K+ N ++FYGNDS YVL RLHQTLYERI+SAKINSSSA++KW+AS+DTSSTD+Y RFMN
Sbjct: 178 EKDRNSRVFYGNDSIYVLLRLHQTLYERIQSAKINSSSADRKWKASSDTSSTDQYDRFMN 357
Query: 423 SLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQ 482
+LYSLLDGSSDN+KFEDDCRAIIG QSYVLFTLDKLIYKLVKQLQAVA+ DEMD KLLQ
Sbjct: 358 ALYSLLDGSSDNTKFEDDCRAIIGIQSYVLFTLDKLIYKLVKQLQAVAA--DEMDTKLLQ 531
Query: 483 LYAYEQSRKSGSFFDVVYHENARVLLHDENIYRIECS--PTRLSIQLMDYGHDKPEVTAV 540
LYAYE+SRK G F D+VYHENARVLLHDENIYRIE S P +LSIQLMD GHDKPEVTAV
Sbjct: 532 LYAYEKSRKPGKFVDMVYHENARVLLHDENIYRIEYSPGPMKLSIQLMDSGHDKPEVTAV 711
Query: 541 SIDPNFSTYLHNDFLSVVPDSKEKSGIFLKRNKLKCARSDELSSQVMDGIQVINGLECKI 600
S+DPNFSTYLHNDFLSVVPD K+KSGIFLKRNK + A +DE SSQ M+G+Q+INGLECKI
Sbjct: 712 SMDPNFSTYLHNDFLSVVPDKKQKSGIFLKRNKRRYASNDEFSSQAMEGLQIINGLECKI 891
Query: 601 ACGSSKV 607
AC SSKV
Sbjct: 892 ACSSSKV 912
>TC206429 similar to UP|Q9LFQ3 (Q9LFQ3) Transcriptional regulatory-like
protein, partial (23%)
Length = 1771
Score = 495 bits (1274), Expect = e-140
Identities = 279/483 (57%), Positives = 346/483 (70%), Gaps = 16/483 (3%)
Frame = +1
Query: 141 EQSNGRTNINNASGLAATLSRP----GYVYREGGLDLPSSEGADSTRPDTSTNGAIIEDT 196
+ S RTN++ + G A T SRP V + G++ PS EG D P NG + E +
Sbjct: 4 DNSLNRTNLDVSPGRALTPSRPTDVDDSVSKSQGVNAPSVEGCDMATPVPVANGVLSESS 183
Query: 197 KAHRCHKESVGHFKSEREEGELSPNGDFEEDNFAVYANAGLEAVHKGKDCNTSQHYQNRR 256
K + H ES G K E+EEGELSPNGD EEDN Y ++ ++++ K K + YQ+R
Sbjct: 184 KV-KTHDESFGPCKIEKEEGELSPNGDSEEDNIVAYGDSNVQSMAKSKHNIERRKYQSRN 360
Query: 257 -EEQICGVAGGENDDESDGSPHRSSDDSENASENG-DVSGTESADGEECSREEHEEDGD- 313
E++ C AGG+ND ++D +DSEN SE G DVSG+ESA G+EC RE+HEE+ D
Sbjct: 361 GEDESCPEAGGDNDADAD------DEDSENVSEAGEDVSGSESA-GDECFREDHEEEEDI 519
Query: 314 -HDN---KVESEGEAEGMADANDVEGDGASLPYSERFLLTAKPLVKYVSPVFHGKE-ENV 368
HD+ K ESEGEAEG+ DA GDG SLP SERFL + KPL K+VS V +E ++
Sbjct: 520 EHDDVDGKAESEGEAEGICDAQ-AGGDGTSLPLSERFLSSVKPLTKHVSAVSFVEEMKDS 696
Query: 369 QIFYGNDSFYVLFRLHQTLYERIRSAKINSSSAEKKWRASNDTSSTDRYARFMNSLYSLL 428
++FYGND FYVLFRLHQ LYERI SAK +S SAE KW+A D SS D Y+RF+N+LY+LL
Sbjct: 697 RVFYGNDDFYVLFRLHQALYERILSAKTHSMSAEMKWKAK-DASSPDPYSRFINALYNLL 873
Query: 429 DGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYEQ 488
DGS++N+KFED+CRAIIG QSYVLFTLDKLIYKLV+QLQ VA TDE+DNKLLQLY YE+
Sbjct: 874 DGSAENAKFEDECRAIIGNQSYVLFTLDKLIYKLVRQLQTVA--TDEVDNKLLQLYEYEK 1047
Query: 489 SRKSGSFFDVVYHENARVLLHDENIYRIECS--PTRLSIQLMDYGHDKPEVTAVSIDPNF 546
SRK G D VYH NA V+LH+ENIYR++CS P+RLSIQLMD ++KPE+ AVSIDPNF
Sbjct: 1048SRKPGKLNDSVYHANAHVILHEENIYRLQCSSTPSRLSIQLMDNMNEKPELFAVSIDPNF 1227
Query: 547 STYLHNDFLSVVPDSKEKSGIFLKRNKLKCARSDELSS--QVMDGIQVINGLECKIACGS 604
S YLHNDFLSV P+ KE GI L RNK + + DELS+ M+G++VINGLECKIAC S
Sbjct: 1228SFYLHNDFLSVFPNKKEPHGIILHRNKRQYGKLDELSAICSAMEGVKVINGLECKIACSS 1407
Query: 605 SKV 607
SK+
Sbjct: 1408SKI 1416
>TC215990
Length = 884
Score = 479 bits (1233), Expect = e-135
Identities = 239/296 (80%), Positives = 264/296 (88%), Gaps = 3/296 (1%)
Frame = +2
Query: 313 DHDNKVESEGEAEGMADANDVEGDGASLPYSERFLLTAKPLVKYVSPVFHGKEENVQIFY 372
+HDNK ESEGEAEGM DANDVEGDGASLPYSERFL+T KPL K+V PV H K+ V++FY
Sbjct: 2 EHDNKAESEGEAEGMTDANDVEGDGASLPYSERFLVTVKPLAKHVPPVLHDKQRTVRVFY 181
Query: 373 GNDSFYVLFRLHQTLYERIRSAKINSSSAEKKWRASNDTSSTDRYARFMNSLYSLLDGSS 432
GNDSFYVLFRLHQTLYERI+SAK+NSSSAEKKWRASNDT S+D+Y RFM++LY+LLDGSS
Sbjct: 182 GNDSFYVLFRLHQTLYERIQSAKVNSSSAEKKWRASNDTGSSDQYGRFMDALYNLLDGSS 361
Query: 433 DNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYEQSRKS 492
D++KFED+CRAIIGTQSYVLFTLDKLIYKLVKQLQ VA T+EMDNKLLQLY YE SRK
Sbjct: 362 DSTKFEDECRAIIGTQSYVLFTLDKLIYKLVKQLQVVA--TEEMDNKLLQLYTYENSRKP 535
Query: 493 GSFFDVVYHENARVLLHDENIYRIECS--PTRL-SIQLMDYGHDKPEVTAVSIDPNFSTY 549
G F D+VYHENARVLLHDENIYRIECS PT+L SIQLMDYG+DKPE+TAVS+DPNFS Y
Sbjct: 536 GRFVDLVYHENARVLLHDENIYRIECSPAPTQLSSIQLMDYGYDKPEMTAVSMDPNFSAY 715
Query: 550 LHNDFLSVVPDSKEKSGIFLKRNKLKCARSDELSSQVMDGIQVINGLECKIACGSS 605
LHNDFLSVVPD KEKSGI+LKRNK K A SDE SSQ +DG+ INGLECKIAC SS
Sbjct: 716 LHNDFLSVVPDKKEKSGIYLKRNKRKYAISDEYSSQTLDGLXXINGLECKIACSSS 883
>TC206430 similar to UP|Q9LFQ3 (Q9LFQ3) Transcriptional regulatory-like
protein, partial (14%)
Length = 1282
Score = 233 bits (594), Expect = 2e-61
Identities = 120/176 (68%), Positives = 142/176 (80%), Gaps = 4/176 (2%)
Frame = +3
Query: 436 KFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYEQSRKSGSF 495
KFED+CRAIIG QSYVLFTLDKLIYKLV+QLQ VA TDE+DNKLLQLY YE+SRKSG
Sbjct: 324 KFEDECRAIIGNQSYVLFTLDKLIYKLVRQLQTVA--TDEVDNKLLQLYEYEKSRKSGKL 497
Query: 496 FDVVYHENARVLLHDENIYRIECS--PTRLSIQLMDYGHDKPEVTAVSIDPNFSTYLHND 553
D VYH NA V+LH++NIYR++CS P+RL IQLMD ++KPE+ AVSIDPNFS YLH+D
Sbjct: 498 NDSVYHANAHVILHEDNIYRLQCSSTPSRLFIQLMDNMNEKPELFAVSIDPNFSFYLHSD 677
Query: 554 FLSVVPDSKEKSGIFLKRNKLKCARSDELSS--QVMDGIQVINGLECKIACGSSKV 607
FLSV P+ KE GI L RNK + DELS+ M+G++V+NGLECKIAC SSK+
Sbjct: 678 FLSVFPNKKEPHGIILHRNKRQYGNLDELSAICSAMEGVKVVNGLECKIACSSSKI 845
Score = 117 bits (292), Expect = 2e-26
Identities = 66/120 (55%), Positives = 80/120 (66%), Gaps = 1/120 (0%)
Frame = +1
Query: 313 DHDNKVESEGEAEGMADANDVEGDGASLPYSERFLLTAKPLVKYVSPVFHGKE-ENVQIF 371
+ D K ESEGEAEG+ DA GDG SLP SERFL + KPL K+VS V +E ++ ++F
Sbjct: 10 EFDGKAESEGEAEGICDAQ-AGGDGTSLPLSERFLSSVKPLTKHVSAVSFVEEMKDSRVF 186
Query: 372 YGNDSFYVLFRLHQTLYERIRSAKINSSSAEKKWRASNDTSSTDRYARFMNSLYSLLDGS 431
YGND FYV FRLHQ LYER+ SAK +S SAE KW+A D SS D Y+ + LL S
Sbjct: 187 YGNDDFYVFFRLHQALYERLLSAKTHSMSAEMKWKA-KDASSPDPYSSLKMNAGQLLGTS 363
>BM307377
Length = 426
Score = 168 bits (426), Expect = 6e-42
Identities = 92/141 (65%), Positives = 102/141 (72%), Gaps = 28/141 (19%)
Frame = +3
Query: 416 RYARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAVAS---- 471
+Y RFMN+LYSLLDGSSDN+KFEDDCRAIIG QSYVLFTLDKLIYKLVKQ++ ++S
Sbjct: 3 QYDRFMNALYSLLDGSSDNTKFEDDCRAIIGIQSYVLFTLDKLIYKLVKQVKYISSSFCL 182
Query: 472 ----------------------GTDEMDNKLLQLYAYEQSRKSGSFFDVVYHENARVLLH 509
DEMD KLLQLYAYE+SRK G F D+VYHENARVLLH
Sbjct: 183 LHTIISFLRLKYLFILF*LQAVAADEMDTKLLQLYAYEKSRKPGKFVDMVYHENARVLLH 362
Query: 510 DENIYRIECS--PTRLSIQLM 528
DENIYRIE S P +LSIQLM
Sbjct: 363 DENIYRIEYSPGPMKLSIQLM 425
>TC215992
Length = 1071
Score = 137 bits (344), Expect = 2e-32
Identities = 65/79 (82%), Positives = 71/79 (89%)
Frame = +3
Query: 529 DYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKRNKLKCARSDELSSQVMD 588
D GHDKPEVTAVS+DPNFSTYLHNDFLSVVPD KEKSGIFLKRNK + A +DE SSQ M+
Sbjct: 3 DSGHDKPEVTAVSMDPNFSTYLHNDFLSVVPDKKEKSGIFLKRNKRRYAGNDEFSSQAME 182
Query: 589 GIQVINGLECKIACGSSKV 607
G+Q+INGLECKIAC SSKV
Sbjct: 183 GLQIINGLECKIACSSSKV 239
>AW200928
Length = 457
Score = 116 bits (290), Expect = 3e-26
Identities = 73/158 (46%), Positives = 91/158 (57%), Gaps = 6/158 (3%)
Frame = +2
Query: 431 SSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEMDNKLLQLYAYEQSR 490
SSDN++FEDD II Y +FT+ K IY+LVK Q VA+ DE D KL QLYAY++SR
Sbjct: 2 SSDNTEFEDDRPPIIAIPPYDIFTIYKPIYQLVKHAQPVAA--DETDYKLSQLYAYDKSR 175
Query: 491 KSGSFFDVVYHENARVLLHDENIYRIECSPTRLSIQLMDYGHDKP------EVTAVSIDP 544
F D+ YH N R L R+ T + Y D P VS+DP
Sbjct: 176 TPAEFADIPYHRNDRALSMTRTYNRLHIHLTNETA----YPTDGPLHMINLR*LHVSLDP 343
Query: 545 NFSTYLHNDFLSVVPDSKEKSGIFLKRNKLKCARSDEL 582
FSTY+H DFLSV+ D+K+ GI LKR K + AR+DEL
Sbjct: 344 *FSTYIHYDFLSVLSDTKQTPGILLKRTKRRHARNDEL 457
>CA820577 weakly similar to PIR|T51447|T51 transcription regulator-like
protein - Arabidopsis thaliana, partial (6%)
Length = 480
Score = 115 bits (287), Expect = 8e-26
Identities = 59/92 (64%), Positives = 70/92 (75%), Gaps = 2/92 (2%)
Frame = +3
Query: 524 SIQLMDYGHDKPEVTAVSIDPNFSTYLHNDFLSVVPDSKEKSGIFLKRNKLKCARSDELS 583
SIQLMD ++KPE+ AVSIDPNFS YLHNDFLSV P+ KE GI L RNK + + DELS
Sbjct: 3 SIQLMDNMNEKPELFAVSIDPNFSFYLHNDFLSVFPNKKEPHGIILHRNKRQYGKLDELS 182
Query: 584 S--QVMDGIQVINGLECKIACGSSKVCNHLLL 613
+ M+G++VINGLECKIAC SSKVC +L
Sbjct: 183 AIXSAMEGVKVINGLECKIACSSSKVCCFFIL 278
>BE801296
Length = 256
Score = 80.5 bits (197), Expect = 2e-15
Identities = 44/86 (51%), Positives = 58/86 (67%)
Frame = +2
Query: 417 YARFMNSLYSLLDGSSDNSKFEDDCRAIIGTQSYVLFTLDKLIYKLVKQLQAVASGTDEM 476
Y M++LY+ L G D++K+ED C A++ T SY LFTL L+YKL KQL AVA T+++
Sbjct: 5 YGMLMHALYNTLYGPHDSTKWEDHCYAMMRTYSYDLFTLYHLMYKLEKQLPAVA--TEDI 178
Query: 477 DNKLLQLYAYEQSRKSGSFFDVVYHE 502
D+KLL LYAYE R+ D YHE
Sbjct: 179 DDKLLHLYAYENPRQPVR*VDSYYHE 256
>BI424157 similar to PIR|T51447|T51 transcription regulator-like protein -
Arabidopsis thaliana, partial (2%)
Length = 421
Score = 43.9 bits (102), Expect = 2e-04
Identities = 18/23 (78%), Positives = 21/23 (91%)
Frame = +1
Query: 585 QVMDGIQVINGLECKIACGSSKV 607
Q +DG+Q+INGLECKIAC SSKV
Sbjct: 1 QTLDGLQIINGLECKIACSSSKV 69
>TC232838 similar to GB|AAP21377.1|30102918|BT006569 At1g47970 {Arabidopsis
thaliana;} , partial (46%)
Length = 902
Score = 43.1 bits (100), Expect = 4e-04
Identities = 26/66 (39%), Positives = 36/66 (54%), Gaps = 3/66 (4%)
Frame = +2
Query: 265 GGENDDESDGSPHRSSDDS-ENASENGDVS-GTESADGEECSREEHEEDGDHDNKVESE- 321
G +NDDE D +P DD E+ E GDV G E D + S ++ ++D D D + E E
Sbjct: 374 GDDNDDEEDDAPDGGDDDDDEDDEEEGDVQRGGEPDDDDNDSDDDSDDDEDEDEEDEEEQ 553
Query: 322 GEAEGM 327
GE E +
Sbjct: 554 GEEEDL 571
Score = 35.4 bits (80), Expect = 0.078
Identities = 25/81 (30%), Positives = 35/81 (42%), Gaps = 4/81 (4%)
Frame = +2
Query: 265 GGENDDESDGSPHRSSDDSENASENGDVSGTESADGEEC----SREEHEEDGDHDNKVES 320
GGE DD+ + S S DD + E+ + G E G E EE+ D + E
Sbjct: 467 GGEPDDDDNDSDDDSDDDEDEDEEDEEEQGEEEDLGTEYLIRPLETAEEEEASSDFEPEE 646
Query: 321 EGEAEGMADANDVEGDGASLP 341
GE E D +D + + A P
Sbjct: 647 NGEEE-EEDVDDEDCEKAEAP 706
Score = 30.8 bits (68), Expect = 1.9
Identities = 19/77 (24%), Positives = 33/77 (42%), Gaps = 4/77 (5%)
Frame = +2
Query: 267 ENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAE- 325
E DDE S DD+E+ + + + D + ++ + GD D+ + E E +
Sbjct: 281 EEDDEEYESVDEVVDDAEDDEDEEEDDDDDEGDDNDDEEDDAPDGGDDDDDEDDEEEGDV 460
Query: 326 ---GMADANDVEGDGAS 339
G D +D + D S
Sbjct: 461 QRGGEPDDDDNDSDDDS 511
>TC214748 homologue to UP|Q39882 (Q39882) Nodulin-26, partial (98%)
Length = 1254
Score = 42.0 bits (97), Expect = 8e-04
Identities = 28/102 (27%), Positives = 49/102 (47%), Gaps = 10/102 (9%)
Frame = -2
Query: 235 AGLEAVHKGKDCNTSQHYQNRREEQI----CGVAGGENDDESDGS---PHRSSDDSENAS 287
+G++ VH+ + + QH + C GG ++ E +G DD ++A+
Sbjct: 611 SGVDCVHQTEGHHDLQHQSVANSDSSGKPECWNRGGSDEGEEEGGHDGAEELGDDVDDAA 432
Query: 288 ENGDVSGTESADGE---ECSREEHEEDGDHDNKVESEGEAEG 326
E GDV+ E A+G+ + + + DGD D + ES G+ G
Sbjct: 431 E*GDVAAHEGAEGDGGVDVAAGDVGADGDGDEEGESMGDGSG 306
>TC224715 homologue to UP|Q39882 (Q39882) Nodulin-26, partial (98%)
Length = 1270
Score = 40.8 bits (94), Expect = 0.002
Identities = 32/118 (27%), Positives = 54/118 (45%), Gaps = 10/118 (8%)
Frame = -2
Query: 237 LEAVHKGKDCNTSQHY----QNRREEQICGVAGGENDDESDGSPHRSS---DDSENASEN 289
++ VH+ + + QH N E C +GG ++ + +G + DD ++A+E
Sbjct: 978 VDRVHQTEGHHDLQHQGVANSNSSGESECWNSGGGDEGQEEGGDDGAEELGDDVDDAAEE 799
Query: 290 GDVSGTESADGE---ECSREEHEEDGDHDNKVESEGEAEGMADANDVEGDGASLPYSE 344
GDV+ E A+G+ + + + DGD D + E G+ G D G G S E
Sbjct: 798 GDVAADEGAEGDGGVDVAAGDVGADGDGDEEGECMGDGGG-----DEAGRGGSAVVGE 640
>TC211437 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%)
Length = 857
Score = 40.0 bits (92), Expect = 0.003
Identities = 24/76 (31%), Positives = 34/76 (44%), Gaps = 6/76 (7%)
Frame = +1
Query: 267 ENDDESDGSPHRSSDDSENASENGD------VSGTESADGEECSREEHEEDGDHDNKVES 320
E +DE DG +D + E+GD G E D E+ + E +DGD D+ +
Sbjct: 364 ETEDEEDGD--EDDEDGPDEDEDGDDEEFSGEEGDEGGDPEDEANGEGSDDGDEDDDGDD 537
Query: 321 EGEAEGMADANDVEGD 336
G+ E D D E D
Sbjct: 538 NGDEEDDDDGEDEEED 585
Score = 38.5 bits (88), Expect = 0.009
Identities = 22/74 (29%), Positives = 31/74 (41%), Gaps = 14/74 (18%)
Frame = +1
Query: 266 GENDDESDGSPHRSSDDSENASENGDVSGT--------------ESADGEECSREEHEED 311
G+ DDE DD E + E GD G E DG++ EE ++D
Sbjct: 385 GDEDDEDGPDEDEDGDDEEFSGEEGDEGGDPEDEANGEGSDDGDEDDDGDDNGDEEDDDD 564
Query: 312 GDHDNKVESEGEAE 325
G+ + + E E E E
Sbjct: 565 GEDEEEDEEEDEEE 606
Score = 38.1 bits (87), Expect = 0.012
Identities = 16/50 (32%), Positives = 27/50 (54%)
Frame = +1
Query: 262 GVAGGENDDESDGSPHRSSDDSENASENGDVSGTESADGEECSREEHEED 311
G GG+ +DE++G D+ ++ +NGD + + EE EE EE+
Sbjct: 457 GDEGGDPEDEANGEGSDDGDEDDDGDDNGDEEDDDDGEDEEEDEEEDEEE 606
>TC219825 weakly similar to UP|MID2_YEAST (P36027) Mating process protein
MID2 (Serine-rich protein SMS1) (Protein kinase A
interference protein), partial (19%)
Length = 862
Score = 38.9 bits (89), Expect = 0.007
Identities = 28/84 (33%), Positives = 36/84 (42%), Gaps = 8/84 (9%)
Frame = -2
Query: 262 GVAGGENDDESD---GSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKV 318
G GGE + D G S DD ++ E V G + D E+ E+ DH+
Sbjct: 378 GDDGGEVPEGGDVDVGD*KESEDDEDSEEEEEGVVGVDGEDHEDQGEEDEGVVEDHEEGD 199
Query: 319 ESEGEAEGMADAND-----VEGDG 337
EGE E + DA D EGDG
Sbjct: 198 P*EGEEEALEDAGDGGEGGGEGDG 127
>TC219510 UP|Q8LJS2 (Q8LJS2) Nucleolar histone deacetylase HD2-P39, complete
Length = 1196
Score = 38.9 bits (89), Expect = 0.007
Identities = 23/86 (26%), Positives = 39/86 (44%), Gaps = 7/86 (8%)
Frame = +3
Query: 274 GSPHRSSDDSENASENGDVSGTESADGEECSREEHEEDGDHDNKVESEGEAE-------G 326
G+P +++D ++ ++ D + GE+ S HE D D +++ ES+G+ E G
Sbjct: 501 GAPAKAADPKKDEDDDSDDESDDDLAGEDESGSSHEMDDDSNSEEESDGDDEETPAKKGG 680
Query: 327 MADANDVEGDGASLPYSERFLLTAKP 352
D A P S + TA P
Sbjct: 681 SGQEXDPMESAAKTPISAKKAKTATP 758
>TC203997 similar to UP|Q9FFJ6 (Q9FFJ6) Phosphate/phosphoenolpyruvate
translocator protein-like, partial (97%)
Length = 1271
Score = 37.7 bits (86), Expect = 0.016
Identities = 25/78 (32%), Positives = 35/78 (44%), Gaps = 5/78 (6%)
Frame = -3
Query: 264 AGGENDDESDGSPH---RSSDDSENASENGD--VSGTESADGEECSREEHEEDGDHDNKV 318
AG + D+ G P D SEN E G G DGE R+ E+DG+ +++
Sbjct: 498 AGDDGDEGEIGEPGLALEGHDVSENGGEEGSGGADGLVKRDGEVAERDVAEDDGEAEDEA 319
Query: 319 ESEGEAEGMADANDVEGD 336
E E A A+ + GD
Sbjct: 318 EGGDLEELEAGADGLHGD 265
>TC219641 similar to UP|Q6VBJ3 (Q6VBJ3) Epa4p, partial (6%)
Length = 942
Score = 37.7 bits (86), Expect = 0.016
Identities = 21/71 (29%), Positives = 34/71 (47%), Gaps = 4/71 (5%)
Frame = +1
Query: 267 ENDDESDGSPHRSSDDSENASENG----DVSGTESADGEECSREEHEEDGDHDNKVESEG 322
E D++ G +D ++ NG D G + DG + ++ +++GD D + E EG
Sbjct: 445 EGDEDFSGEGEEEADPEDDPEANGAGGSDSDGDDDDDGGD---DDDDDNGDDDGEDEEEG 615
Query: 323 EAEGMADANDV 333
E E D DV
Sbjct: 616 EDEDEEDEEDV 648
>TC211022 similar to UP|O81600 (O81600) H-protein promoter binding factor-2a,
partial (19%)
Length = 578
Score = 36.2 bits (82), Expect = 0.046
Identities = 20/48 (41%), Positives = 28/48 (57%), Gaps = 4/48 (8%)
Frame = +1
Query: 296 ESADGEECSREEHEEDGDHDNKVES----EGEAEGMADANDVEGDGAS 339
ESA+ E EEHE++GD D +VE+ E EA+ DA + + G S
Sbjct: 73 ESAEDREQEMEEHEDEGDKDTRVENVTKEELEADPPLDAEETKISGTS 216
>AW760687
Length = 484
Score = 36.2 bits (82), Expect = 0.046
Identities = 26/70 (37%), Positives = 35/70 (49%), Gaps = 3/70 (4%)
Frame = -1
Query: 260 ICGVAGGENDDESDGSPHRSSDDSENASENGDVS---GTESADGEECSREEHEEDGDHDN 316
I G E SDG+ DD ++A+E GDV+ G E A G + + DGD D
Sbjct: 382 IDGSVLAEQQAASDGA-EELGDDVDDAAE*GDVAAHEGAEGAGGVIVAAADVGADGDGDE 206
Query: 317 KVESEGEAEG 326
+ ES G+A G
Sbjct: 205 EGESMGDASG 176
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.129 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 25,036,288
Number of Sequences: 63676
Number of extensions: 323132
Number of successful extensions: 1575
Number of sequences better than 10.0: 102
Number of HSP's better than 10.0 without gapping: 1472
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1526
length of query: 637
length of database: 12,639,632
effective HSP length: 103
effective length of query: 534
effective length of database: 6,081,004
effective search space: 3247256136
effective search space used: 3247256136
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146588.5


