

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146584.2 - phase: 0 /pseudo
(1452 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
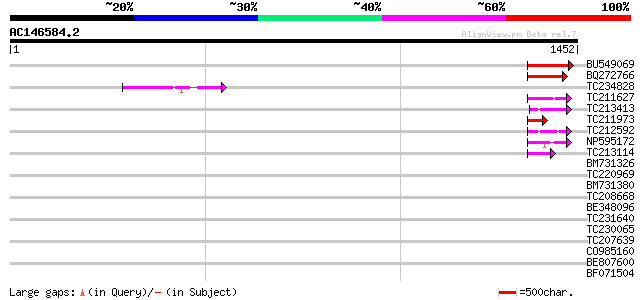
Score E
Sequences producing significant alignments: (bits) Value
BU549069 160 3e-39
BQ272766 weakly similar to GP|28558781|gb| pol protein {Cucumis ... 130 3e-30
TC234828 68 3e-11
TC211627 65 3e-10
TC213413 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 57 8e-08
TC211973 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 54 7e-07
TC212592 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, parti... 53 9e-07
NP595172 polyprotein [Glycine max] 52 1e-06
TC213114 weakly similar to UP|Q8W150 (Q8W150) Polyprotein, parti... 51 3e-06
BM731326 weakly similar to GP|21740635|em OSJNBb0043H09.2 {Oryza... 42 0.002
TC220969 similar to UP|Q94IB3 (Q94IB3) WRKY DNA-binding protein,... 39 0.017
BM731380 34 0.41
TC208668 homologue to UP|Q9NGW8 (Q9NGW8) Developmental protein D... 33 0.70
BE348096 33 0.70
TC231640 homologue to UP|Q918P0 (Q918P0) Latency associated anti... 33 0.92
TC230065 similar to UP|O14329 (O14329) SPBC16E9.14c protein, par... 33 1.2
TC207639 similar to UP|O48592 (O48592) GT2 protein, partial (24%) 33 1.2
CO985160 33 1.2
BE807600 similar to GP|24461867|gb CTV.22 {Poncirus trifoliata},... 32 1.6
BF071504 32 2.7
>BU549069
Length = 615
Score = 160 bits (406), Expect = 3e-39
Identities = 74/117 (63%), Positives = 94/117 (80%)
Frame = -1
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQV 1385
+++ P FIG +QIL+R VAY++ LPP LSNLHNVFHVSQLR Y+ DPSHV++ DDVQV
Sbjct: 570 RKLTPHFIGHFQILKRAXPVAYQIALPPSLSNLHNVFHVSQLRMYIHDPSHVVKLDDVQV 391
Query: 1386 RDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESYPELF 1442
++NLT ETLP+RI+DR+ K LR KE PLV+V+W G +GE TWELES+M +YP LF
Sbjct: 390 KENLTYETLPLRIEDRRTKHLRRKENPLVKVIWGGTSGEDATWELESQMRVAYPSLF 220
>BQ272766 weakly similar to GP|28558781|gb| pol protein {Cucumis melo}, partial
(9%)
Length = 410
Score = 130 bits (328), Expect = 3e-30
Identities = 60/101 (59%), Positives = 80/101 (78%)
Frame = -3
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ P FIGP+QIL++V VAY++ LPP L++LHNVFHVSQ+ KY+ DPSH++ D VQV+
Sbjct: 303 KLTPHFIGPFQILKKVDFVAYQIALPPSLTSLHNVFHVSQIHKYINDPSHMVNLDVVQVK 124
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLT 1427
+NL ETLP+RI D + K LRGKEI LV+V+W GA+GE T
Sbjct: 123 ENLAHETLPLRIKDMRTKHLRGKEILLVKVIWGGASGEEAT 1
>TC234828
Length = 857
Score = 67.8 bits (164), Expect = 3e-11
Identities = 82/280 (29%), Positives = 117/280 (41%), Gaps = 14/280 (5%)
Frame = +1
Query: 289 LIATERVTLVHSVLSLRGRRLQVESLL*LVLRQRTRIV*SEVLAILIILL*LLLLILVLR 348
LI V L + L + ++ + LL*+V ++ +* V L+I *+ +I V R
Sbjct: 64 LIPQGAVMLATTTLVVEDQKYPLGYLL*VVQKRLPPTI*YGVSV*LLINY*MYFMIQVPR 243
Query: 349 IVLLLSIVFLLWVLICLI*MERWLSKLQLRVR*LLLSYV*NVLCLYLVVISKWI*FVYL* 408
I+ L V+ W + W L+ * L *+VL L V + *F YL*
Sbjct: 244 ILSYLMHVWRGWGYVRRSCHMIWWCLLRRASL*PPLGCA*SVL*LLRVDLLWRT*FAYL* 423
Query: 409 AVWM*FWV*TG*STTTFLLIVLASQCISL--------------PSKRKVVRSFYLLSS*S 454
+ M* W G T + +C L P+K++ + S
Sbjct: 424 LI*M*SWEWIG-YLLTISF*TVRRRC*CLVEM*FQVNH*KRMQPTKKRKMFELTWCCSLC 600
Query: 455 NWNATVS*CFR*WLLYLLRIKQ*LIGYRWCVNFLKFFQMRFLMCLQREKLSFLLILFLER 514
W T+ L+ YRWC NF KFF+M F+ C REK + LL L R
Sbjct: 601 MWKRTLK----------------LVVYRWCQNFQKFFRMMFVNCHLREKWNSLLTWCLGR 732
Query: 515 SRSRWHLIVCLLPSYLS*RNSWRTCLRRSLLDQVFHLGER 554
+ R LI CL + *R+ +RT +L Q H GER
Sbjct: 733 IQCRLRLIECLRWN*QR*RHKYRTF*ANNLFVQAHHRGER 852
>TC211627
Length = 1034
Score = 64.7 bits (156), Expect = 3e-10
Identities = 40/113 (35%), Positives = 60/113 (52%), Gaps = 1/113 (0%)
Frame = +3
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLR-KYVPDPSHVIQSDDVQV 1385
++ +F GPYQI RVG VAYR+ LPP S +H +FHVS L+ + P P ++
Sbjct: 492 KLSKRFYGPYQIQARVGQVAYRLQLPP-TSKIHPIFHVSLLKVHHGPIPPELLALPPFST 668
Query: 1386 RDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESY 1438
++ V+ P++ D K+ IP V V WT E TWE +++ + Y
Sbjct: 669 TNHPLVQ--PLQFLDWKMDESTTPPIPQVLVQWTNLAPEDTTWESWTQLKDIY 821
>TC213413 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (5%)
Length = 761
Score = 56.6 bits (135), Expect = 8e-08
Identities = 33/108 (30%), Positives = 53/108 (48%)
Frame = +1
Query: 1331 KFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVRDNLT 1390
++ GP+++ ER+G V YR+ L H S +H VFHVS L+ +V DP + + V
Sbjct: 85 RYYGPFEVQERLGKVVYRLKLTAH-SRIHPVFHVSLLKAFVGDP-ETTHAGPLPVMRTEE 258
Query: 1391 VETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESY 1438
P+ + D K+ +V V W A+ + +WE + E Y
Sbjct: 259 ATNTPLTVIDSKLVPADNGPRRMVLVQWPSASRQDASWEDWQVLRERY 402
>TC211973 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (4%)
Length = 730
Score = 53.5 bits (127), Expect = 7e-07
Identities = 22/52 (42%), Positives = 37/52 (70%)
Frame = +1
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHV 1377
+++ P+F P+Q+ +VGT+AY++ LP H+ +H VFHVS L+K P+ H+
Sbjct: 208 EKLSPRFYAPFQVFNKVGTIAYKLDLPSHI-KIHPVFHVSLLKKASPNLYHL 360
>TC212592 weakly similar to UP|Q84ZV5 (Q84ZV5) Polyprotein, partial (4%)
Length = 664
Score = 53.1 bits (126), Expect = 9e-07
Identities = 35/112 (31%), Positives = 59/112 (52%)
Frame = +2
Query: 1327 EVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYVPDPSHVIQSDDVQVR 1386
++ +F GP+Q++ERVG VAYR+ LP + LH VFH S L+ + +P +
Sbjct: 74 KLSKRFFGPFQVVERVGKVAYRLQLPVD-AKLHPVFHCSLLKPFQGNPPDTAAPLPPTLF 250
Query: 1387 DNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLESY 1438
D+ +V V + R T+ +I V V W G + + TWE +++ + +
Sbjct: 251 DHQSVIAPLVILATR---TVNDDDIE-VLVQWQGLSPDDATWEKWTELCKEF 394
>NP595172 polyprotein [Glycine max]
Length = 4659
Score = 52.4 bits (124), Expect = 1e-06
Identities = 34/122 (27%), Positives = 58/122 (46%), Gaps = 8/122 (6%)
Frame = +1
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLR--------KYVPDPSHV 1377
+++ ++ GP+++L ++G VAY++ L P + +H VFHVSQL+ Y+P P V
Sbjct: 4126 QKLSMRYFGPFKVLAKIGDVAYKLEL-PSAARIHPVFHVSQLKPFNGTAQDPYLPLPLTV 4302
Query: 1378 IQSDDVQVRDNLTVETLPVRIDDRKVKTLRGKEIPLVRVVWTGATGESLTWELESKMLES 1437
+ V PV+I ++ +I + V W + TWE + S
Sbjct: 4303 TEMGPVM---------QPVKILASRIIIRGHNQIEQILVQWENGLQDEATWEDIEDIKAS 4455
Query: 1438 YP 1439
YP
Sbjct: 4456 YP 4461
Score = 34.3 bits (77), Expect = 0.41
Identities = 54/208 (25%), Positives = 90/208 (42%), Gaps = 4/208 (1%)
Frame = +2
Query: 536 WRTCLRRSLLDQVFHLGERRFC**RRKMVVCGYVLIIGS*TR*LSRIGIHFRELMI*WIS 595
++ C ++ +QV +C RRKM V II * + L RI ++ W++
Sbjct: 1850 YKKCWFKASFNQVTVHFPYPYCLLRRKMEVGDSAQIIEL*MQSL*RIAFQCQQ----WMN 2017
Query: 596 *WVHVFSARL----I*GQVITRLK*KMKICRRQLSERVMVTMNIKLCLLVLPMHLECLWS 651
++ + I*GQ + ++I R+ L E +M MN C LVL MH +
Sbjct: 2018 C*MNYMELNIFQNWI*GQDTIKFWYSLRIERKLLLEHIMAIMNG**CHLVLLMHQPHSNA 2197
Query: 652 I*IASSMHFWIGLWLCLSMIF*FTPRLKKNMLSI*RLFCKC*KRRNFMLNCLSVNSG*KK 711
* L+LC M *+ L + + S L + * +++L+C +V+ +K
Sbjct: 2198 **TKYFSLLSENLFLCFLMTS*YIVLLGRIISSTWNLCYRP*SNISYLLDCPNVHLEIQK 2377
Query: 712 *VF*AMLFLVMVLQWIRLKLKRYHNGRL 739
+ A +L W + K+Y G L
Sbjct: 2378 LITWATRYLG*EFPWRIQRYKQY*IGLL 2461
>TC213114 weakly similar to UP|Q8W150 (Q8W150) Polyprotein, partial (7%)
Length = 810
Score = 51.2 bits (121), Expect = 3e-06
Identities = 29/81 (35%), Positives = 48/81 (58%), Gaps = 10/81 (12%)
Frame = +3
Query: 1326 KEVDPKFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRKYV-----PDPSHVIQS 1380
+++ P+F GPYQI +++G VA+ + LPP +H VFH S L+K V P P ++ S
Sbjct: 114 EKLSPRFYGPYQIKKQIGLVAFELDLPP-ARKIHPVFHASLLKKAVAATANPQPLPLMLS 290
Query: 1381 DDV-----QVRDNLTVETLPV 1396
+D+ Q++ L++ L V
Sbjct: 291 EDLSSEFFQLKSKLSITILMV 353
>BM731326 weakly similar to GP|21740635|em OSJNBb0043H09.2 {Oryza sativa
(japonica cultivar-group)}, partial (3%)
Length = 424
Score = 42.4 bits (98), Expect = 0.002
Identities = 18/39 (46%), Positives = 29/39 (74%)
Frame = +1
Query: 1331 KFIGPYQILERVGTVAYRVGLPPHLSNLHNVFHVSQLRK 1369
+F GPY ++ +VG VA+++ LP + +H VFHVSQL++
Sbjct: 172 RFYGPYLVMRQVGAVAFQLQLPSE-ARIHPVFHVSQLKR 285
>TC220969 similar to UP|Q94IB3 (Q94IB3) WRKY DNA-binding protein, partial
(8%)
Length = 677
Score = 38.9 bits (89), Expect = 0.017
Identities = 22/60 (36%), Positives = 40/60 (66%)
Frame = -3
Query: 320 RQRTRIV*SEVLAILIILL*LLLLILVLRIVLLLSIVFLLWVLICLI*MERWLSKLQLRV 379
+ T+I+ S +L ++++LL +LLL+L++ ++LL+ + L+ +L C M R KL LRV
Sbjct: 414 QHETQIICSALLLLMLLLLLMLLLLLLMLLLLLMLLSLLMLLLRCQERMNRHDRKLCLRV 235
>BM731380
Length = 433
Score = 34.3 bits (77), Expect = 0.41
Identities = 18/37 (48%), Positives = 29/37 (77%)
Frame = +2
Query: 330 VLAILIILL*LLLLILVLRIVLLLSIVFLLWVLICLI 366
VL +L+++L LLL++L+L VLLL +VFLL + + L+
Sbjct: 287 VLVLLLLVLLLLLVLLLLLQVLLLLLVFLLLLQVLLL 397
Score = 31.6 bits (70), Expect = 2.7
Identities = 28/62 (45%), Positives = 37/62 (59%)
Frame = +2
Query: 337 LL*LLLLILVLRIVLLLSIVFLLWVLICLI*MERWLSKLQLRVR*LLLSYV*NVLCLYLV 396
LL LLLL+LVL + L L +V LL VL+ L+ + L L L + LLL VL L LV
Sbjct: 236 LLLLLLLLLVLVLGLYLVLVLLLLVLLLLLVLLLLLQVLLLLLVFLLLL---QVLLLLLV 406
Query: 397 VI 398
++
Sbjct: 407 LL 412
Score = 31.2 bits (69), Expect = 3.5
Identities = 23/57 (40%), Positives = 36/57 (62%)
Frame = +2
Query: 331 LAILIILL*LLLLILVLRIVLLLSIVFLLWVLICLI*MERWLSKLQLRVR*LLLSYV 387
L +L++LL +L+L L L +VLLL ++ LL VL+ L+ + L L ++ LLL V
Sbjct: 236 LLLLLLLLLVLVLGLYLVLVLLLLVLLLLLVLLLLLQVLLLLLVFLLLLQVLLLLLV 406
Score = 31.2 bits (69), Expect = 3.5
Identities = 22/55 (40%), Positives = 36/55 (65%)
Frame = +2
Query: 330 VLAILIILL*LLLLILVLRIVLLLSIVFLLWVLICLI*MERWLSKLQLRVR*LLL 384
VL + ++L+ LLLL+L+L +VLLL + LL +L+ L+ ++ L L L + LL
Sbjct: 269 VLGLYLVLV-LLLLVLLLLLVLLLLLQVLLLLLVFLLLLQVLLLLLVLLLAQFLL 430
Score = 30.0 bits (66), Expect = 7.7
Identities = 24/53 (45%), Positives = 34/53 (63%)
Frame = +2
Query: 314 LL*LVLRQRTRIV*SEVLAILIILL*LLLLILVLRIVLLLSIVFLLWVLICLI 366
LL LVL +V +L +L++LL LLLL+ VL +LLL + LL VL+ L+
Sbjct: 251 LLLLVLVLGLYLVLVLLLLVLLLLLVLLLLLQVL--LLLLVFLLLLQVLLLLL 403
>TC208668 homologue to UP|Q9NGW8 (Q9NGW8) Developmental protein DG1037
(Fragment), partial (4%)
Length = 1021
Score = 33.5 bits (75), Expect = 0.70
Identities = 17/29 (58%), Positives = 24/29 (82%)
Frame = +2
Query: 336 ILL*LLLLILVLRIVLLLSIVFLLWVLIC 364
ILL LLLL+L+L ++LLL ++ LL VL+C
Sbjct: 209 ILLLLLLLLLLLLLLLLLLLLLLLQVLVC 295
>BE348096
Length = 445
Score = 33.5 bits (75), Expect = 0.70
Identities = 17/39 (43%), Positives = 23/39 (58%)
Frame = -1
Query: 829 RR*SGSLCFKTVESS*EELSYA*SRVSGRSLCIEDMETL 867
RR S LC + S ++LSYA*SRV +C+ +E L
Sbjct: 187 RRPSSDLCSSIGQDSLKDLSYA*SRVGSHGVCLHILEAL 71
>TC231640 homologue to UP|Q918P0 (Q918P0) Latency associated antigen, partial
(7%)
Length = 466
Score = 33.1 bits (74), Expect = 0.92
Identities = 18/37 (48%), Positives = 29/37 (77%)
Frame = -1
Query: 330 VLAILIILL*LLLLILVLRIVLLLSIVFLLWVLICLI 366
+L + ++LL LLLL+L+L ++LLL ++FLL +L LI
Sbjct: 121 LLLLFLLLLLLLLLLLLLLLLLLLLLLFLLLLLGYLI 11
Score = 33.1 bits (74), Expect = 0.92
Identities = 16/36 (44%), Positives = 29/36 (80%)
Frame = -1
Query: 330 VLAILIILL*LLLLILVLRIVLLLSIVFLLWVLICL 365
+L +L++ L LLLL+L+L ++LLL ++ LL++L+ L
Sbjct: 130 LLLLLLLFLLLLLLLLLLLLLLLLLLLLLLFLLLLL 23
Score = 31.6 bits (70), Expect = 2.7
Identities = 16/36 (44%), Positives = 27/36 (74%)
Frame = -1
Query: 331 LAILIILL*LLLLILVLRIVLLLSIVFLLWVLICLI 366
L +L++L LLLL+L+L ++LLL ++ LL L+ L+
Sbjct: 130 LLLLLLLFLLLLLLLLLLLLLLLLLLLLLLFLLLLL 23
>TC230065 similar to UP|O14329 (O14329) SPBC16E9.14c protein, partial (6%)
Length = 610
Score = 32.7 bits (73), Expect = 1.2
Identities = 16/33 (48%), Positives = 22/33 (66%)
Frame = +1
Query: 341 LLLILVLRIVLLLSIVFLLWVLICLI*MERWLS 373
L L L L++VLLLS++FL W L L+ + LS
Sbjct: 202 LTLTLTLQLVLLLSLLFLTWTLFSLLTLSHLLS 300
>TC207639 similar to UP|O48592 (O48592) GT2 protein, partial (24%)
Length = 1689
Score = 32.7 bits (73), Expect = 1.2
Identities = 16/34 (47%), Positives = 28/34 (82%)
Frame = +2
Query: 333 ILIILL*LLLLILVLRIVLLLSIVFLLWVLICLI 366
++ +LL LLLL+L+L ++LLLS++ LL +L+ L+
Sbjct: 368 MVTVLLLLLLLLLLLLLLLLLSLLLLLLLLLLLL 469
Score = 32.0 bits (71), Expect = 2.0
Identities = 16/30 (53%), Positives = 25/30 (83%)
Frame = +2
Query: 330 VLAILIILL*LLLLILVLRIVLLLSIVFLL 359
VL +L++LL LLLL+L+L ++LLL ++ LL
Sbjct: 377 VLLLLLLLLLLLLLLLLLSLLLLLLLLLLL 466
Score = 30.0 bits (66), Expect = 7.7
Identities = 16/33 (48%), Positives = 25/33 (75%)
Frame = +2
Query: 330 VLAILIILL*LLLLILVLRIVLLLSIVFLLWVL 362
V +L++LL LLLL+L+L + LLL ++ LL +L
Sbjct: 371 VTVLLLLLLLLLLLLLLLLLSLLLLLLLLLLLL 469
>CO985160
Length = 777
Score = 32.7 bits (73), Expect = 1.2
Identities = 28/79 (35%), Positives = 46/79 (57%)
Frame = +3
Query: 317 LVLRQRTRIV*SEVLAILIILL*LLLLILVLRIVLLLSIVFLLWVLICLI*MERWLSKLQ 376
L+L Q +++ ++L +L+ L LLLL+L LR +LLL + +L+ L + L LQ
Sbjct: 435 LLLLQLRKLLLLDLLLLLLQLRKLLLLLLQLRKLLLLLLQLRKLLLLLLELRKLLLLLLQ 614
Query: 377 LRVR*LLLSYV*NVLCLYL 395
LR LLL + +L L++
Sbjct: 615 LRKLLLLLLELRKLLLLHV 671
Score = 31.6 bits (70), Expect = 2.7
Identities = 34/93 (36%), Positives = 52/93 (55%), Gaps = 6/93 (6%)
Frame = +3
Query: 298 VHSVLSLRGRRLQVESLL*LVLRQRTRIV*SEVLAILIILL*LLLLILVLRIVLLLSIVF 357
+ +L L+ R+L + LL L+L+ R ++L +L+ L LLLL+L LR +LLL +
Sbjct: 426 LRKLLLLQLRKLLLLDLLLLLLQLR------KLLLLLLQLRKLLLLLLQLRKLLLLLLEL 587
Query: 358 LLWVLICLI*MERWLSKLQLR------VR*LLL 384
+L+ L + L L+LR VR*LLL
Sbjct: 588 RKLLLLLLQLRKLLLLLLELRKLLLLHVR*LLL 686
Score = 30.4 bits (67), Expect = 5.9
Identities = 29/100 (29%), Positives = 57/100 (57%)
Frame = +3
Query: 273 RKVML*LIAIVRTLFALIATERVTLVHSVLSLRGRRLQVESLL*LVLRQRTRIV*SEVLA 332
RK++L L+ + + L L+ ++ L+ +L LR + LL L+ ++ ++ E+
Sbjct: 498 RKLLLLLLQLRKLLLLLLQLRKLLLL--LLELR------KLLLLLLQLRKLLLLLLELRK 653
Query: 333 ILIILL*LLLLILVLRIVLLLSIVFLLWVLICLI*MERWL 372
+L++ + LLL+L LR +LLL + L+ +L L+ + R L
Sbjct: 654 LLLLHVR*LLLLLQLRKMLLLVLHLLMLLLHLLLLLTRQL 773
>BE807600 similar to GP|24461867|gb CTV.22 {Poncirus trifoliata}, partial
(4%)
Length = 414
Score = 32.3 bits (72), Expect = 1.6
Identities = 24/56 (42%), Positives = 36/56 (63%)
Frame = -2
Query: 336 ILL*LLLLILVLRIVLLLSIVFLLWVLICLI*MERWLSKLQLRVR*LLLSYV*NVL 391
ILL LLLL+L+L ++LLL ++ LL +L + + L + LR+ *LLL + N L
Sbjct: 407 ILLLLLLLLLLLLLLLLLLLLLLLELLPQKLMLHLLLVDILLRIL*LLLLLLGNNL 240
>BF071504
Length = 422
Score = 31.6 bits (70), Expect = 2.7
Identities = 18/53 (33%), Positives = 36/53 (66%)
Frame = -3
Query: 313 SLL*LVLRQRTRIV*SEVLAILIILL*LLLLILVLRIVLLLSIVFLLWVLICL 365
SLL ++L +++* +++ ++++LL LL L+L ++LLL ++ LL +CL
Sbjct: 357 SLLLMLLLLLLQLL*EQIMLLMLLLLLLL*QKLLLLLLLLLLLLLLLVQCLCL 199
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.365 0.164 0.591
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 68,303,604
Number of Sequences: 63676
Number of extensions: 1052157
Number of successful extensions: 15199
Number of sequences better than 10.0: 49
Number of HSP's better than 10.0 without gapping: 6035
Number of HSP's successfully gapped in prelim test: 584
Number of HSP's that attempted gapping in prelim test: 8237
Number of HSP's gapped (non-prelim): 7708
length of query: 1452
length of database: 12,639,632
effective HSP length: 109
effective length of query: 1343
effective length of database: 5,698,948
effective search space: 7653687164
effective search space used: 7653687164
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.6 bits)
S2: 65 (29.6 bits)
Medicago: description of AC146584.2


