

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146575.4 - phase: 0
(165 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
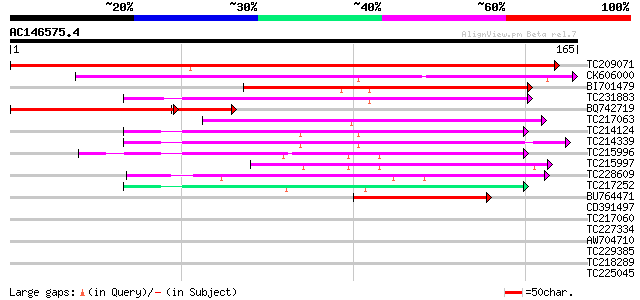
Score E
Sequences producing significant alignments: (bits) Value
TC209071 239 4e-64
CK606000 83 7e-17
BI701479 81 3e-16
TC231883 weakly similar to UP|Q7QBQ3 (Q7QBQ3) AgCP4056 (Fragment... 77 4e-15
BQ742719 69 1e-14
TC217063 56 9e-09
TC214124 55 1e-08
TC214339 similar to GB|CAA71502.1|2769566|ATCTPP chloroplast thy... 55 2e-08
TC215996 similar to UP|Q9LV44 (Q9LV44) Similarity to signal pept... 54 3e-08
TC215997 similar to UP|Q9LV44 (Q9LV44) Similarity to signal pept... 51 3e-07
TC228609 similar to UP|Q84VZ6 (Q84VZ6) At2g31140, partial (71%) 46 7e-06
TC217252 weakly similar to GB|CAA71502.1|2769566|ATCTPP chloropl... 42 1e-04
BU764471 39 9e-04
CD391497 33 0.062
TC217060 33 0.062
TC227334 weakly similar to UP|Q9LXX7 (Q9LXX7) Cytochrome P450-li... 29 1.2
AW704710 28 2.6
TC229385 similar to UP|Q9SGN7 (Q9SGN7) F3M18.17 (At1g28390/F3M18... 26 7.6
TC218289 26 7.6
TC225045 similar to UP|Q8L7C3 (Q8L7C3) Expressed protein, partia... 26 9.9
>TC209071
Length = 953
Score = 239 bits (610), Expect = 4e-64
Identities = 107/164 (65%), Positives = 127/164 (77%), Gaps = 4/164 (2%)
Frame = +2
Query: 1 MGTRNLVWNVTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFT----DDYVF 56
MGT + +WN TKK + G++T TV+D + TV+PVRG SMSPTFNPK S DDYV
Sbjct: 179 MGTSSFLWNCTKKFISAGIVTVTVTDLFVTVIPVRGGSMSPTFNPKAGSLMGGVFDDYVL 358
Query: 57 VEKLCLDKFKFSHGDIVIFSSPSNFKETHIKRIIALPGEWFVNRHNQDVLKVPEGHCWVE 116
VEK CL +KFSHGD+V+F SP N KETH+KRI ALPGEWF DV+++P GHCWVE
Sbjct: 359 VEKFCLHSYKFSHGDVVVFRSPQNRKETHVKRIAALPGEWFGTHQKNDVIQIPLGHCWVE 538
Query: 117 GDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIGAVKNTTPE 160
GDN ASS DS S+GP+PLG++RGRVTHVVWPPQRIGAVKNT P+
Sbjct: 539 GDNTASSLDSNSFGPIPLGIIRGRVTHVVWPPQRIGAVKNTPPQ 670
>CK606000
Length = 629
Score = 82.8 bits (203), Expect = 7e-17
Identities = 51/151 (33%), Positives = 74/151 (48%), Gaps = 5/151 (3%)
Frame = -1
Query: 20 ITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPS 79
+ + V VRG SM P N N YV + + G IV F +P
Sbjct: 539 VCYFVESHVVGTTRVRGPSMYPYLNENYNETRFGYVCLTWKLYAQQNLQRGMIVTFYNPM 360
Query: 80 NFKETHIKRIIALPGEWFVNR--HNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLV 137
+ IKRII LPG+ + ++VP H WVEGD S DS +YGP+ + LV
Sbjct: 359 KPESMAIKRIIGLPGDMIHTKAPFPTPYIRVPPNHIWVEGDGT-KSLDSNTYGPISIRLV 183
Query: 138 RGRVTHVVWPPQRIGAVK---NTTPERLPSS 165
+G+VTH++WP ++ VK + P+RL ++
Sbjct: 182 QGQVTHILWPFRKAERVKWWEHQLPDRLTAA 90
>BI701479
Length = 422
Score = 80.9 bits (198), Expect = 3e-16
Identities = 39/89 (43%), Positives = 56/89 (62%), Gaps = 5/89 (5%)
Frame = +2
Query: 69 HGDIVIFSSPSNFKETHIKRIIALPGE---WFVNRHNQ--DVLKVPEGHCWVEGDNAASS 123
HGD+V+ SP N K KR++A+ G+ +F H++ V VP+GH W++GDN +S
Sbjct: 14 HGDLVLVRSPLNPKIRLTKRVVAVEGDTVTYFDPLHSEAAQVAVVPKGHVWIQGDNIYAS 193
Query: 124 TDSKSYGPVPLGLVRGRVTHVVWPPQRIG 152
DS+ +GPVP GL+ G+V VWPP G
Sbjct: 194 RDSRHFGPVPYGLIEGKVFFRVWPPDSFG 280
>TC231883 weakly similar to UP|Q7QBQ3 (Q7QBQ3) AgCP4056 (Fragment), partial
(17%)
Length = 1263
Score = 77.0 bits (188), Expect = 4e-15
Identities = 44/121 (36%), Positives = 61/121 (50%), Gaps = 2/121 (1%)
Frame = +1
Query: 34 VRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKETHIKRIIALP 93
V G M PT N D + + L HGD+V+ SP N K +KR+
Sbjct: 499 VYGVRMLPTLNA-----AGDVLLTDPLSPRLGNIGHGDLVLLRSPLNPKIRLMKRVEGDN 663
Query: 94 GEWFVNRHNQ--DVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRI 151
+F H++ V VP+ H W++GDN +S DS+ +GPVP GL+ G+V VWPP
Sbjct: 664 VTYFDALHSKAAQVAVVPKRHVWIQGDNIYASRDSRHFGPVPYGLIEGKVFFRVWPPDSF 843
Query: 152 G 152
G
Sbjct: 844 G 846
>BQ742719
Length = 426
Score = 68.6 bits (166), Expect(2) = 1e-14
Identities = 29/49 (59%), Positives = 37/49 (75%)
Frame = +1
Query: 1 MGTRNLVWNVTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNS 49
MGT + +WN TKK + G++T TV+D + TV+PVRG SMSPTFNPK S
Sbjct: 133 MGTSSFLWNCTKKFISAGIVTVTVTDLFVTVIPVRGGSMSPTFNPKAGS 279
Score = 27.3 bits (59), Expect(2) = 1e-14
Identities = 11/19 (57%), Positives = 12/19 (62%)
Frame = +3
Query: 48 NSFTDDYVFVEKLCLDKFK 66
N DDYV VEK CL +K
Sbjct: 369 NYTADDYVLVEKFCLHSYK 425
>TC217063
Length = 861
Score = 55.8 bits (133), Expect = 9e-09
Identities = 34/122 (27%), Positives = 55/122 (44%), Gaps = 22/122 (18%)
Frame = +3
Query: 57 VEKLCLDKFKFSHGDIVIFSSPSNFKETHIKRIIALPGEWFV------------------ 98
+EK+ K + GDIV+ +P + + KR++ L G+
Sbjct: 249 MEKISPRFGKVTCGDIVVLRNPQHPRHFMTKRVVGLEGDSVTYISNPETNEYEGDSFTHI 428
Query: 99 ----NRHNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIGAV 154
N + VP+G WVEGDN +S DS+ +GPVP L+ G++ + P ++ G
Sbjct: 429 SSPDNGDKSKTIVVPKGAVWVEGDNKYNSNDSRKFGPVPYDLIDGKMFWRITPLKKFGPF 608
Query: 155 KN 156
N
Sbjct: 609 WN 614
>TC214124
Length = 1778
Score = 55.5 bits (132), Expect = 1e-08
Identities = 43/146 (29%), Positives = 62/146 (42%), Gaps = 28/146 (19%)
Frame = +1
Query: 34 VRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKE-------THI 86
+ +SM PT D V EK+ K DIVIF +P +E I
Sbjct: 856 IPSSSMYPTLE------VGDRVLTEKVSFFFRKPDVSDIVIFKAPPCLEEFGFSSSDVFI 1017
Query: 87 KRIIALPGEWFVNR---------------------HNQDVLKVPEGHCWVEGDNAASSTD 125
KRI+A G+ R + D + VPEG+ +V GDN +S D
Sbjct: 1018KRIVAKAGDTVEVRDGKLLVNGAAEERQFVVEPLAYEMDPMVVPEGYVFVMGDNRNNSFD 1197
Query: 126 SKSYGPVPLGLVRGRVTHVVWPPQRI 151
S ++GP+P+ + GR WPP ++
Sbjct: 1198SHNWGPLPVENIVGRSMFRYWPPSKV 1275
>TC214339 similar to GB|CAA71502.1|2769566|ATCTPP chloroplast thylakoidal
processing peptidase {Arabidopsis thaliana;} , partial
(55%)
Length = 1836
Score = 54.7 bits (130), Expect = 2e-08
Identities = 47/158 (29%), Positives = 65/158 (40%), Gaps = 28/158 (17%)
Frame = +3
Query: 34 VRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKE-------THI 86
+ +SM PT D V EK+ K DIVIF +P +E I
Sbjct: 876 IPSSSMYPTLE------VGDRVLTEKVSFFFRKPDVSDIVIFKAPPWLEEFGFSSSDVFI 1037
Query: 87 KRIIALPGEWFVNR---------------------HNQDVLKVPEGHCWVEGDNAASSTD 125
KRI+A G+ R + D + VPEG+ +V GDN S D
Sbjct: 1038 KRIVAKAGDTVEVRDGKLLINGAAEEQEFVLEALAYEMDPMVVPEGYVFVMGDNRNKSFD 1217
Query: 126 SKSYGPVPLGLVRGRVTHVVWPPQRIGAVKNTTPERLP 163
S ++GP+P+ + GR WPP + A T +LP
Sbjct: 1218 SHNWGPLPVENIVGRSMFRYWPPSK--ASDTDTHRKLP 1325
>TC215996 similar to UP|Q9LV44 (Q9LV44) Similarity to signal peptidase,
partial (57%)
Length = 1245
Score = 53.9 bits (128), Expect = 3e-08
Identities = 49/160 (30%), Positives = 68/160 (41%), Gaps = 29/160 (18%)
Frame = +1
Query: 21 TFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSP-- 78
TF RY + SM PTF+ D + EK+ K DIVIF SP
Sbjct: 499 TFVAEPRY-----IPSLSMYPTFD------VGDRIIAEKVSYYFRKPCASDIVIFKSPPV 645
Query: 79 ------SNFKETHIKRIIALPGEWF-----------VNRHNQDVL----------KVPEG 111
SNF + IKR++A G+ V ++ + +L +VPE
Sbjct: 646 LQEVGYSNF-DVFIKRMVAKEGDIVEVRKGHLVVNGVEKNEEYILEPPAYEMKPTRVPEN 822
Query: 112 HCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRI 151
+ +V GDN +S DS +GP+P + R WPP RI
Sbjct: 823 YVFVMGDNRNNSYDSHVWGPLPAKNIIDRSVFRYWPPNRI 942
>TC215997 similar to UP|Q9LV44 (Q9LV44) Similarity to signal peptidase,
partial (37%)
Length = 720
Score = 50.8 bits (120), Expect = 3e-07
Identities = 39/117 (33%), Positives = 53/117 (44%), Gaps = 29/117 (24%)
Frame = +3
Query: 71 DIVIFSSPSNFKET-------HIKRIIALPGEWF-----------VNRHNQDVL------ 106
DIVIF SP +E IKR++A G+ V R+ + +L
Sbjct: 12 DIVIFKSPPVLQEVGYSDDDVFIKRVVAKAGDIVEVRKGHLVVNGVERNEEYILEPPAYE 191
Query: 107 ----KVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVTHVVWPPQRI-GAVKNTT 158
+VPE + +V GDN +S DS +GP+P + GR WPP RI G V T
Sbjct: 192 MKPTRVPENYVFVMGDNRNNSYDSHVWGPLPAKNIIGRSVFRYWPPNRIAGTVSKET 362
>TC228609 similar to UP|Q84VZ6 (Q84VZ6) At2g31140, partial (71%)
Length = 1140
Score = 46.2 bits (108), Expect = 7e-06
Identities = 36/129 (27%), Positives = 60/129 (45%), Gaps = 6/129 (4%)
Frame = +1
Query: 35 RGASMSPTFNPKTNSFTDDYVFVEKL-CLDKFKFSHGDIVIFSSPSNFKETHIKRIIALP 93
+G M+PT + K V KL +D + GD+V+ P ++R+ A+
Sbjct: 238 KGEEMAPTIDGKA------VTLVRKLPVVDPTRVFVGDVVVVKDPEKPDNYLLRRLTAVE 399
Query: 94 GEWFVNRHNQDVLKVPE-GHCWVEGDN----AASSTDSKSYGPVPLGLVRGRVTHVVWPP 148
G V+ +D V E CWV +N A + DS+++GPV + + GRV + +
Sbjct: 400 GYEMVSTDEKDEAFVLEKDQCWVVAENEKLKAKEAKDSRTFGPVQMTDIVGRVIYCLRSA 579
Query: 149 QRIGAVKNT 157
G V+N+
Sbjct: 580 VDHGRVQNS 606
>TC217252 weakly similar to GB|CAA71502.1|2769566|ATCTPP chloroplast
thylakoidal processing peptidase {Arabidopsis thaliana;}
, partial (37%)
Length = 769
Score = 42.4 bits (98), Expect = 1e-04
Identities = 39/144 (27%), Positives = 54/144 (37%), Gaps = 26/144 (18%)
Frame = +1
Query: 34 VRGASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPS-----NFKETHIKR 88
+ +SM PT D + VEK + DIV F P+ N IKR
Sbjct: 241 IPSSSMYPTLR------VGDRIIVEKASYYIRSPAIHDIVTFKDPTQSSGENTDAVFIKR 402
Query: 89 IIALPGEWFVNRHN---------------------QDVLKVPEGHCWVEGDNAASSTDSK 127
I+A G+ H + VP GH +V GDN +S DS
Sbjct: 403 IVAKAGDTVEVNHGALYINGVAQQEDFIAEPPAYAMQLTHVPNGHVYVLGDNRNNSYDSH 582
Query: 128 SYGPVPLGLVRGRVTHVVWPPQRI 151
+GP+P+ + GR P+ I
Sbjct: 583 VWGPLPVKNIVGRYVTCYHRPRNI 654
>BU764471
Length = 386
Score = 39.3 bits (90), Expect = 9e-04
Identities = 17/40 (42%), Positives = 27/40 (67%)
Frame = +2
Query: 101 HNQDVLKVPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGR 140
+ D + VPEG+ +V GDN +S DS ++GP+P+ + GR
Sbjct: 254 YEMDPMVVPEGYVFVMGDNRNNSFDSHNWGPLPVENIVGR 373
>CD391497
Length = 594
Score = 33.1 bits (74), Expect = 0.062
Identities = 17/49 (34%), Positives = 28/49 (56%), Gaps = 4/49 (8%)
Frame = -1
Query: 113 CWVEGDN----AASSTDSKSYGPVPLGLVRGRVTHVVWPPQRIGAVKNT 157
CWVE +N A + DS+++GPV + + GRV + + G V+N+
Sbjct: 531 CWVEAENEKLNAKEAKDSRTFGPVQMTDIVGRVIYCLRSAVDHGRVQNS 385
>TC217060
Length = 477
Score = 33.1 bits (74), Expect = 0.062
Identities = 20/60 (33%), Positives = 31/60 (51%)
Frame = +1
Query: 36 GASMSPTFNPKTNSFTDDYVFVEKLCLDKFKFSHGDIVIFSSPSNFKETHIKRIIALPGE 95
G SM PT + KT F +EK+ K + GDIV+ +P + + KR++ L G+
Sbjct: 247 GPSMLPTIDLKTGVF-----LMEKISPRFGKVTCGDIVVLRNPQHPRHFMTKRVVGLEGD 411
>TC227334 weakly similar to UP|Q9LXX7 (Q9LXX7) Cytochrome P450-like protein,
partial (77%)
Length = 1754
Score = 28.9 bits (63), Expect = 1.2
Identities = 18/60 (30%), Positives = 25/60 (41%), Gaps = 2/60 (3%)
Frame = -1
Query: 108 VPEGHCWVEGDNAASSTDSKSYGPVPLGLVRGRVT--HVVWPPQRIGAVKNTTPERLPSS 165
+P+ H W + S P GL G V H + PP R+ V++ P PSS
Sbjct: 1484 LPQTHSWTRVKHRIPKRTLGSILQPPFGLKFGTVLPPHALHPPHRVRVVRHEPPLPNPSS 1305
>AW704710
Length = 404
Score = 27.7 bits (60), Expect = 2.6
Identities = 16/48 (33%), Positives = 24/48 (49%)
Frame = -1
Query: 9 NVTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVF 56
N +K+ I T + T++P+ + TFNPKTN F D+ F
Sbjct: 368 NFFRKVRHIDSTTRMIMMNNPTIIPIS----THTFNPKTNVFLDNPEF 237
>TC229385 similar to UP|Q9SGN7 (Q9SGN7) F3M18.17 (At1g28390/F3M18_17),
partial (34%)
Length = 1245
Score = 26.2 bits (56), Expect = 7.6
Identities = 11/28 (39%), Positives = 18/28 (64%)
Frame = +1
Query: 113 CWVEGDNAASSTDSKSYGPVPLGLVRGR 140
C+V DN ++ TD S+G + L ++ GR
Sbjct: 733 CYVTPDNLSTKTDVFSFGILLLEIISGR 816
>TC218289
Length = 1586
Score = 26.2 bits (56), Expect = 7.6
Identities = 9/23 (39%), Positives = 13/23 (56%)
Frame = -2
Query: 143 HVVWPPQRIGAVKNTTPERLPSS 165
H WPP A+ +T+P +P S
Sbjct: 421 HEPWPPSHFAAMLSTSPTAMPPS 353
>TC225045 similar to UP|Q8L7C3 (Q8L7C3) Expressed protein, partial (67%)
Length = 640
Score = 25.8 bits (55), Expect = 9.9
Identities = 22/74 (29%), Positives = 32/74 (42%)
Frame = +2
Query: 2 GTRNLVWNVTKKLATIGLITFTVSDRYATVVPVRGASMSPTFNPKTNSFTDDYVFVEKLC 61
GTR L+ +KL I +T T +ATV + GAS+S T + T+ F V
Sbjct: 8 GTRPLL---IQKLLGIAKMTTT----FATVAGLGGASLSSTSSRLTSGFVKSPVSARNTL 166
Query: 62 LDKFKFSHGDIVIF 75
+G + F
Sbjct: 167RQAVAMGNGRVTCF 208
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.136 0.422
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,809,795
Number of Sequences: 63676
Number of extensions: 108283
Number of successful extensions: 488
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 480
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 482
length of query: 165
length of database: 12,639,632
effective HSP length: 90
effective length of query: 75
effective length of database: 6,908,792
effective search space: 518159400
effective search space used: 518159400
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146575.4


