

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146574.11 + phase: 0 /pseudo
(738 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
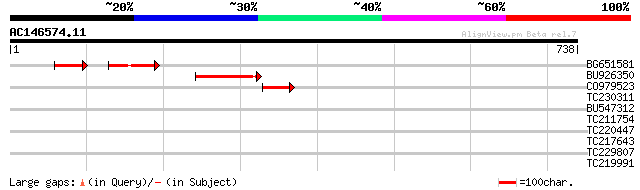
Score E
Sequences producing significant alignments: (bits) Value
BG651581 75 6e-26
BU926350 103 2e-22
CO979523 75 8e-14
TC230311 homologue to GB|AAF03534.1|6102610|AF187317 CAF protein... 37 0.041
BU547312 homologue to GP|11559645|gb short integuments 1 {Arabid... 35 0.092
TC211754 similar to UP|Q9M378 (Q9M378) TATA box binding protein ... 34 0.20
TC220447 32 1.0
TC217643 similar to UP|Q40593 (Q40593) Violaxanthin de-epoxidase... 30 3.9
TC229807 weakly similar to UP|O80872 (O80872) Expressed protein ... 30 5.0
TC219991 weakly similar to UP|Q892K6 (Q892K6) ATP-dependent DNA ... 29 8.6
>BG651581
Length = 325
Score = 75.5 bits (184), Expect(2) = 6e-26
Identities = 37/67 (55%), Positives = 52/67 (77%)
Frame = +3
Query: 129 KLIVGLRPALLIYMKFTQKPDQTDLVDVEQQLSLTEKAIASSCDICTCLDASKNVPNIEG 188
+L + LRP + ++MK KPD+T+LV +QLSLTE I ++C+ICTCL+A KNVP++EG
Sbjct: 129 RLALVLRPTISVHMKSMLKPDETELV---KQLSLTEGKILNACNICTCLNA*KNVPDVEG 299
Query: 189 WPVSKVA 195
P+SKVA
Sbjct: 300 LPISKVA 320
Score = 61.2 bits (147), Expect(2) = 6e-26
Identities = 31/44 (70%), Positives = 37/44 (83%), Gaps = 1/44 (2%)
Frame = +2
Query: 59 DVCPTADAVRAFVEHLVDPMLPAKASIRD-APSISQQQKVAKQV 101
DVCP DA++AF+E+LVDP+LPAK SIRD +PS SQ Q VAKQV
Sbjct: 2 DVCPAEDALKAFLEYLVDPVLPAKPSIRDNSPSPSQHQLVAKQV 133
>BU926350
Length = 464
Score = 103 bits (258), Expect = 2e-22
Identities = 55/86 (63%), Positives = 63/86 (72%)
Frame = +3
Query: 242 YQKRRVIQNPTKNGLNVDDDEFLQVGYSAVKEATGVNSNDIMLLESYTVYSQRKEKTASR 301
Y+KRR I+ PTKN N +++ LQ+GYSAVK+A G+N DI LESYTVYSQ K KTASR
Sbjct: 6 YKKRRAIKKPTKNESNSEEEGILQIGYSAVKKAAGINKTDITSLESYTVYSQSKAKTASR 185
Query: 302 FYIMKCSQSTADGSIQVPIKDLIESF 327
FYIMKCSQ IQ PIK LIE F
Sbjct: 186 FYIMKCSQLINLEFIQ-PIKSLIERF 260
>CO979523
Length = 829
Score = 75.5 bits (184), Expect = 8e-14
Identities = 34/42 (80%), Positives = 40/42 (94%)
Frame = -2
Query: 329 GPLLKKSSSSWTITSVVEYFHVLPYSEIISDWISRLV*FTFP 370
GPL+K+SSSSWT+TSVVEYFHVL YSE+I +WISRLV*F+FP
Sbjct: 474 GPLVKRSSSSWTVTSVVEYFHVLLYSEVILEWISRLV*FSFP 349
>TC230311 homologue to GB|AAF03534.1|6102610|AF187317 CAF protein
{Arabidopsis thaliana;} , partial (6%)
Length = 842
Score = 36.6 bits (83), Expect = 0.041
Identities = 26/74 (35%), Positives = 36/74 (48%), Gaps = 8/74 (10%)
Frame = +3
Query: 667 NGTWLRK-LDGICHENNWILPTYSVSLSDGEFHA-----TVRVKGVDFEYS--CEGNTCP 718
N T+ R+ L+ IC NW +P Y G HA VRV D ++ C G P
Sbjct: 147 NQTFTRQTLNDICLRRNWPMPFYRCVNEGGPAHAKRFTFAVRVNTTDKGWTDECVGEPMP 326
Query: 719 FPREARDSAAAQML 732
++A+DSAA +L
Sbjct: 327 SVKKAKDSAAVLLL 368
>BU547312 homologue to GP|11559645|gb short integuments 1 {Arabidopsis
thaliana}, partial (3%)
Length = 595
Score = 35.4 bits (80), Expect = 0.092
Identities = 23/66 (34%), Positives = 31/66 (46%), Gaps = 7/66 (10%)
Frame = -2
Query: 674 LDGICHENNWILPTYSVSLSDGEFHA-----TVRVKGVDFEYS--CEGNTCPFPREARDS 726
L+ IC NW +P Y G HA VRV D ++ C G P ++A+DS
Sbjct: 585 LNDICLRRNWPMPFYRCVNEGGPAHAKRFTXAVRVNTTDRGWTDECVGEPMPSVKKAKDS 406
Query: 727 AAAQML 732
AA +L
Sbjct: 405 AAVLLL 388
>TC211754 similar to UP|Q9M378 (Q9M378) TATA box binding protein (TBP)
associated factor (TAF)-like protein, partial (3%)
Length = 737
Score = 34.3 bits (77), Expect = 0.20
Identities = 11/17 (64%), Positives = 14/17 (81%)
Frame = +3
Query: 672 RKLDGICHENNWILPTY 688
++LD CHENNW+LP Y
Sbjct: 684 QELDNTCHENNWVLPAY 734
>TC220447
Length = 594
Score = 32.0 bits (71), Expect = 1.0
Identities = 30/118 (25%), Positives = 45/118 (37%), Gaps = 10/118 (8%)
Frame = +2
Query: 447 KHEVTESHVSSKGLYIDLDNK------PGSDTKVALNQKEKNGCGITKRCDSVKEDWDMD 500
KH +++ S Y +D + P SDT + +TK + V E
Sbjct: 101 KHSLSKPSPSLSLRYCAMDQRHGPQPLPESDT---IGNDSHPKTQLTKETERVVESLSDA 271
Query: 501 VDKSLVLPSKNKECQKHTANTLHISEDQKIENPSVQHHSNE----CTRPSKAEKAVSK 554
+ L S +K C TL +E +VQH S++ T P+ KA SK
Sbjct: 272 ESRHRELRSSDKPCSDSNLETLGAGVTSSVETLAVQHRSDDLQGSSTAPNSGGKAESK 445
>TC217643 similar to UP|Q40593 (Q40593) Violaxanthin de-epoxidase precursor,
partial (46%)
Length = 895
Score = 30.0 bits (66), Expect = 3.9
Identities = 17/32 (53%), Positives = 19/32 (59%)
Frame = -2
Query: 308 SQSTADGSIQVPIKDLIESFRGPLLKKSSSSW 339
S ST+ SI P + I SF PL K SSSSW
Sbjct: 627 SFSTSPASILSPSRISISSFDNPLKKFSSSSW 532
>TC229807 weakly similar to UP|O80872 (O80872) Expressed protein
(At2g30010/F23F1.7), partial (14%)
Length = 1369
Score = 29.6 bits (65), Expect = 5.0
Identities = 19/47 (40%), Positives = 24/47 (50%)
Frame = -2
Query: 95 QKVAKQVHSVVLLYNYYHRKRHPELEFVAFKDFCKLIVGLRPALLIY 141
Q+V+ + S VL HP EF FK F I+GL P LLI+
Sbjct: 264 QRVSNKSQSFVL---------HPSQEFRTFKFFHLTILGLPPKLLIF 151
>TC219991 weakly similar to UP|Q892K6 (Q892K6) ATP-dependent DNA helicase
recQ , partial (3%)
Length = 1030
Score = 28.9 bits (63), Expect = 8.6
Identities = 17/62 (27%), Positives = 28/62 (44%), Gaps = 11/62 (17%)
Frame = +2
Query: 466 NKPGSDTKVALNQKEKNG-----CGITKRCD------SVKEDWDMDVDKSLVLPSKNKEC 514
N+ G D + A N + C + + C+ S E WD+++D+ +LP E
Sbjct: 491 NQTGKDDEPAHNASNLSDPTLETCHVERYCEDGISAKSSLEKWDLEIDEVPILPVNGSEV 670
Query: 515 QK 516
QK
Sbjct: 671 QK 676
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.136 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 32,748,298
Number of Sequences: 63676
Number of extensions: 462991
Number of successful extensions: 2865
Number of sequences better than 10.0: 21
Number of HSP's better than 10.0 without gapping: 2832
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2864
length of query: 738
length of database: 12,639,632
effective HSP length: 104
effective length of query: 634
effective length of database: 6,017,328
effective search space: 3814985952
effective search space used: 3814985952
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146574.11


