

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146571.2 - phase: 0
(527 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
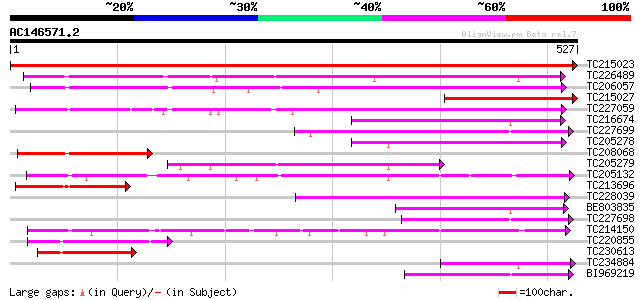
Score E
Sequences producing significant alignments: (bits) Value
TC215023 similar to UP|Q6PBW6 (Q6PBW6) Chaperonin containing TCP... 963 0.0
TC226489 homologue to UP|Q8H9B2 (Q8H9B2) T-complex polypeptide 1... 306 1e-83
TC206057 homologue to UP|Q9ZRX1 (Q9ZRX1) Cytosolic chaperonin, d... 293 1e-79
TC215027 weakly similar to UP|Q8WSY6 (Q8WSY6) CCT chaperonin bet... 223 1e-58
TC227059 similar to UP|O81503 (O81503) F9D12.18 protein, partial... 217 9e-57
TC216674 homologue to GB|AAM26704.1|20857172|AY102137 AT3g11830/... 139 4e-33
TC227699 similar to UP|TCPZ_HUMAN (P40227) T-complex protein 1, ... 124 8e-29
TC205278 similar to PRF|2206327A.0|1587206|2206327A T complex pr... 122 5e-28
TC208068 homologue to PRF|2206327A.0|1587206|2206327A T complex ... 117 1e-26
TC205279 homologue to PRF|2206327A.0|1587206|2206327A T complex ... 108 4e-24
TC205132 homologue to UP|CH62_CUCMA (Q05046) Chaperonin CPN60-2,... 105 4e-23
TC213696 homologue to GB|AAM26704.1|20857172|AY102137 AT3g11830/... 104 1e-22
TC228039 similar to GB|AAQ56831.1|34098897|BT010388 At3g03960 {A... 97 2e-20
BE803835 similar to GP|24371053|db t-complex polypeptide 1 {Brug... 97 2e-20
TC227698 similar to UP|TCPZ_HUMAN (P40227) T-complex protein 1, ... 89 6e-18
TC214150 homologue to UP|Q9ZTV1 (Q9ZTV1) Chaperonin 60 alpha sub... 87 2e-17
TC220855 similar to GB|AAQ56831.1|34098897|BT010388 At3g03960 {A... 84 2e-16
TC230613 homologue to UP|Q8L7N0 (Q8L7N0) TCP-1 chaperonin-like p... 79 4e-15
TC234884 similar to UP|TCPA_DROME (P12613) T-complex protein 1, ... 70 2e-12
BI969219 65 1e-10
>TC215023 similar to UP|Q6PBW6 (Q6PBW6) Chaperonin containing TCP1, subunit 2
(Beta), partial (98%)
Length = 1992
Score = 963 bits (2490), Expect = 0.0
Identities = 496/527 (94%), Positives = 515/527 (97%)
Frame = +3
Query: 1 MTIGNIFKDEASEEKGERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTN 60
M I NIFK+EASEEKGERARM+SFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTN
Sbjct: 96 MAIDNIFKNEASEEKGERARMASFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTN 275
Query: 61 DGATILKSLHIDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMT 120
DGATILKSLHIDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVA KIHPMT
Sbjct: 276 DGATILKSLHIDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVATKIHPMT 455
Query: 121 IIAGFRMAAECARSALVEKVVDNKGDAEKFRSDLMNIARTTLSSKILSQDKEHFAKLAVD 180
II+GFRMAAECAR+AL+EKVVDNK D+EKFRSDL+NIA TTLSSKILSQDKEHFAKLAVD
Sbjct: 456 IISGFRMAAECARNALLEKVVDNKADSEKFRSDLLNIAMTTLSSKILSQDKEHFAKLAVD 635
Query: 181 AVMRLKGSTNLESVQIIKKPGGSLIDSFLDEGFILDKKIGLGQPKRIENAKILVANTAMD 240
AVMRLKGSTNLES+QIIKKPGGSL+DSFLDEGFILDKKIG+GQPKRIENAKILVANTAMD
Sbjct: 636 AVMRLKGSTNLESIQIIKKPGGSLMDSFLDEGFILDKKIGIGQPKRIENAKILVANTAMD 815
Query: 241 TDKVKIYGARVRVDSMAKVAEIEGAEKEKMKEKVNKIIGHGINCFVNRQLIYNFPEELFA 300
TDKVKIYGARVRVDSMA+VA+IE AEKEKM+EKV KIIGHGINCFVNRQLIYNFPEELFA
Sbjct: 816 TDKVKIYGARVRVDSMARVAQIETAEKEKMREKVQKIIGHGINCFVNRQLIYNFPEELFA 995
Query: 301 DAGILAIEHADFDGIERLALVTGGEIASTFDNPESVKLGQCELIEEIMIGEDKLIKFSGV 360
DAGILAIEHADFDGIERLALVTGGEIASTFDNPESVKLG C+LIEEIMIGEDKLI FSGV
Sbjct: 996 DAGILAIEHADFDGIERLALVTGGEIASTFDNPESVKLGHCDLIEEIMIGEDKLIHFSGV 1175
Query: 361 AMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALA 420
AMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKE+DALA
Sbjct: 1176AMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEVDALA 1355
Query: 421 RKTPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCTAGIDVISGSV 480
+KTPGKKSLA+EAFSRALLAIPT IADNAGLDSAELISQLRAEHQ EGCTAGIDVISGSV
Sbjct: 1356KKTPGKKSLAIEAFSRALLAIPTIIADNAGLDSAELISQLRAEHQKEGCTAGIDVISGSV 1535
Query: 481 GDMVERGICEAFKVKQAVLLSATEAAEMILRVDDIITCAPRRREDRM 527
GDM ERGICEAFKVKQAVLLS+TEAAEMILRVD+IITCAPRRREDRM
Sbjct: 1536GDMAERGICEAFKVKQAVLLSSTEAAEMILRVDEIITCAPRRREDRM 1676
>TC226489 homologue to UP|Q8H9B2 (Q8H9B2) T-complex polypeptide 1, complete
Length = 2038
Score = 306 bits (785), Expect = 1e-83
Identities = 189/528 (35%), Positives = 306/528 (57%), Gaps = 25/528 (4%)
Frame = +1
Query: 14 EKGERARMSSFVGAMAIADLVKTTLGPKGMDKIL-QSTGRGREVTVTNDGATILKSLHID 72
+ G+ R + V A+A++VK++LGP G+DK+L G +VT+TNDGATILK L ++
Sbjct: 112 QSGQDVRTQNVVACQAVANIVKSSLGPVGLDKMLVDDIG---DVTITNDGATILKMLEVE 282
Query: 73 NPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAECA 132
+PAAKVLV+++++QD EVGDGTTSVV++A ELL+ A LV KIHP +II+G+R+A A
Sbjct: 283 HPAAKVLVELAELQDREVGDGTTSVVIVAAELLKRANDLVRNKIHPTSIISGYRLAMREA 462
Query: 133 RSALVEKVVDNKGDAEKFRSD-LMNIARTTLSSKILSQDKEHFAKLAVDAVMRLKGSTN- 190
+ EK+ EK D L+N A+T++SSK+++ D + FA L VDAV +K TN
Sbjct: 463 CKYVEEKLAVK---VEKLGKDSLINCAKTSMSSKLIAGDSDFFAILVVDAVQAVK-MTNA 630
Query: 191 -------LESVQIIKKPGGSLIDSFLDEGFILDK-KIGLGQPKRIENAKILVANTAMDTD 242
++ + I+K G S DSFL G+ L+ + G P R+ A+I + +
Sbjct: 631 RGEVKYPIKGINILKAHGKSARDSFLMNGYALNTGRAAQGMPLRVAPARIACLDFNLQKT 810
Query: 243 KVKIYGARVRVDSMAKVAEIEGAEKEKMKEKVNKIIGHGINCFVNRQLIYNFPEELFADA 302
K+++ G +V V ++ +I E + KE++ K++ G N + + I + + F +A
Sbjct: 811 KMQL-GVQVLVTDPRELEKIRQREADMTKERIEKLLKAGANVILTTKGIDDMALKYFVEA 987
Query: 303 GILAIEHADFDGIERLALVTGGEIASTFDNPESVK------LGQCELIEEIMIGEDKLIK 356
G +A+ + + +A TG + STF + E + LG + + E I +D ++
Sbjct: 988 GAIAVRRVRKEDMRHVAKATGATLVSTFADMEGEETFEPSFLGYADEVVEERISDDAVVM 1167
Query: 357 FSGVAMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEI 416
G A T++LRGA+ H+LDE +R+LHDAL ++ +T+ + V+ GGG E ++ +
Sbjct: 1168IKGTKTTSAVTLILRGANDHMLDEMDRALHDALSIVKRTLESNTVVAGGGAVEAALSVYL 1347
Query: 417 DALARKTPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCTA----- 471
+ LA ++ LA+ F+ +LL IP ++ NA D+ EL+++LRA H + A
Sbjct: 1348EYLATTLGSREQLAIAEFAESLLIIPKVLSVNAAKDATELVAKLRAYHHSAQTKADKKHL 1527
Query: 472 ---GIDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDII 516
G+D+ G + + +E G+ E K ++ ATEAA ILR+DD+I
Sbjct: 1528SSMGLDLSEGKIRNNLEAGVIEPAMSKVKIIQFATEAAITILRIDDMI 1671
>TC206057 homologue to UP|Q9ZRX1 (Q9ZRX1) Cytosolic chaperonin,
delta-subunit, complete
Length = 1930
Score = 293 bits (750), Expect = 1e-79
Identities = 183/511 (35%), Positives = 289/511 (55%), Gaps = 13/511 (2%)
Frame = +2
Query: 20 RMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATILKSLHIDNPAAKVL 79
R ++ V A ++A+ V+T+LGPKGMDK++ ++ EV +TNDGATIL + + PAAK+L
Sbjct: 161 RHANIVAARSVANAVRTSLGPKGMDKMISTSSD--EVIITNDGATILNKMQVLQPAAKML 334
Query: 80 VDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAECARSALVEK 139
V++SK QD GDGTT+VVV+AG LL + L++ IHP + AA A L
Sbjct: 335 VELSKSQDSAAGDGTTTVVVIAGALLEQCLLLLSHGIHPTVVSDALHKAAVKAVDVLTAM 514
Query: 140 VVDNKGDAEKFRSDLMNIARTTLSSKILSQDKEHFAKLAVDAVMRLKGS-----TNLESV 194
V + R L+ A T+L+SK++SQ A LAVDAV+ + + +L V
Sbjct: 515 AVPVELSD---RDSLVKSASTSLNSKVVSQYSTLLAPLAVDAVLSVVDAPKPDMVDLRDV 685
Query: 195 QIIKKPGGSLIDSFLDEGFILDKKIG--LGQPKRIENAKILVANTAMDTDKVKIYGARVR 252
+I+KK GG++ D+ L +G + DKK+ G P R+ENAKI V + K I + V
Sbjct: 686 KIVKKLGGTVDDTELVKGLVFDKKVSHAAGGPTRMENAKIAVIQFQISPPKTDIEQSIV- 862
Query: 253 VDSMAKVAEIEGAEKEKMKEKVNKIIGHGINCF-----VNRQLIYNFPEELFADAGILAI 307
V +++ I E+ + + KI G N + R + + A A IL I
Sbjct: 863 VSDYSQMDRILKEERSYILSMIKKIKATGCNVLLIQKSILRDAVTDLSLHYLAKAKILVI 1042
Query: 308 EHADFDGIERLALVTGGEIASTFDNPESVKLGQCELIEEIMIGEDKLIKFSGVA-MGQAC 366
+ + D IE + + ++ + KLG +L+EE +G+ K++K +G+ MG+
Sbjct: 1043KDVERDEIEFITKTLNCLPIANIEHFRTEKLGYADLVEEFSLGDGKIVKITGIKEMGKTT 1222
Query: 367 TIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALARKTPGK 426
T+++RG++ VLDEAERSLHDALCV+ V ++ GGG PE+ +++++ A A+ G
Sbjct: 1223TVLVRGSNQLVLDEAERSLHDALCVVRCLVAKRFLIAGGGAPEIELSRQLGAWAKVLHGM 1402
Query: 427 KSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCTAGIDVISGSVGDMVER 486
+ + AF+ AL IP T+A+NAGL+ ++++LR H AGI+V G + +++E
Sbjct: 1403EGYCVRAFAEALEVIPYTLAENAGLNPIAIVTELRNRHAQGEINAGINVRKGQITNILEE 1582
Query: 487 GICEAFKVKQAVLLSATEAAEMILRVDDIIT 517
+ + V + + ATE MIL++DDI+T
Sbjct: 1583NVVQPLLVSTSAITLATECVRMILKIDDIVT 1675
>TC215027 weakly similar to UP|Q8WSY6 (Q8WSY6) CCT chaperonin beta subunit,
partial (20%)
Length = 695
Score = 223 bits (569), Expect = 1e-58
Identities = 113/123 (91%), Positives = 119/123 (95%)
Frame = +3
Query: 405 GGWPEMVMAKEIDALARKTPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEH 464
GGWPEMVMAKE+DALA+KTPGKKSLA+EAFSRALLAIPT IADNAGLDSAELISQLRAEH
Sbjct: 3 GGWPEMVMAKEVDALAKKTPGKKSLAIEAFSRALLAIPTIIADNAGLDSAELISQLRAEH 182
Query: 465 QNEGCTAGIDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDIITCAPRRRE 524
Q EGCT+GIDVISGSVGDM ERGI EAFKVKQAVLLS+TEAAEMILRVD+IITCAPRRRE
Sbjct: 183 QKEGCTSGIDVISGSVGDMAERGISEAFKVKQAVLLSSTEAAEMILRVDEIITCAPRRRE 362
Query: 525 DRM 527
DRM
Sbjct: 363 DRM 371
>TC227059 similar to UP|O81503 (O81503) F9D12.18 protein, partial (92%)
Length = 2147
Score = 217 bits (553), Expect = 9e-57
Identities = 160/529 (30%), Positives = 276/529 (51%), Gaps = 17/529 (3%)
Frame = +2
Query: 6 IFKDEASEEKGERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATI 65
+ KD E G + R + A A+AD+V+TTLGP+ M K+L G + VTNDG I
Sbjct: 218 VLKDSLKRESGSKVRYAIIQAAKAVADVVRTTLGPRSMLKMLLDAQGG--IVVTNDGNAI 391
Query: 66 LKSLHIDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGF 125
L+ L + +PAAK ++++S+ QD+EVGDGTTSV++LAGE+L A+ + KIHP I +
Sbjct: 392 LRELDLAHPAAKSMIELSRTQDEEVGDGTTSVIILAGEMLHVADAFI-DKIHPTVICRAY 568
Query: 126 RMAAECARSALVEKVV--DNKGDAEKFRSDLMNIARTTLSSKILSQDKEHFAKLAVDAVM 183
A E A A+++K+ N D R ++ + ++ + +K + + A LA+DA
Sbjct: 569 AKALEDA-IAVLDKIAMPINAQD----RGIMLGLVKSCIGTKFTGRFGDLIADLAIDATT 733
Query: 184 RL-----KGSTNLE---SVQIIKKPGGSLIDSFLDEGFILDKK-IGLGQPKR-IENAKIL 233
+ +G +++ +++ K PGG L DS + +G +++K + G+ +R I N +I+
Sbjct: 734 TVGVEVGQGLRDVDIKNYIKVEKVPGGQLEDSRVLKGVMINKDVVAPGKMRRKIVNPRII 913
Query: 234 VANTAMDTDKVKIYGARVRVDSMAKVAE---IEGAEKEKMKEKVNKIIGHGINCFVNRQL 290
+ + ++ K G + K + + E+E ++E +I+ + + +
Sbjct: 914 LLDCPLEYKK----GENQTNAELLKEEDWSLLLKMEEEYIEELCMQILKFKPDLVITEKG 1081
Query: 291 IYNFPEELFADAGILAIEHADFDGIERLALVTGGEIASTFDNPESVKLGQ-CELIEEIMI 349
+ + + G+ AI R+A G I + D + +G L E I
Sbjct: 1082LSDLACHYLSKHGVSAIRRLRKTDNNRIAKACGAVIVNRPDELQESDVGTGAGLFEVKKI 1261
Query: 350 GEDKLIKFSGVAMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPE 409
G++ +ACT++LRGAS +L+E ER+L DA+ V + + +++ GGG E
Sbjct: 1262GDEYFAFIVDCKEPKACTVLLRGASKDLLNEVERNLQDAMSVARNIIKNPKLVPGGGATE 1441
Query: 410 MVMAKEIDALARKTPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQN-EG 468
+ ++ + + G + EA + A AIP T+A N G++ ++ L+ +H N E
Sbjct: 1442LTVSAALKQKSSSIEGIEKWPYEAAALAFEAIPRTLAQNCGVNVIRTMTALQGKHANGEN 1621
Query: 469 CTAGIDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDIIT 517
GID +GS+ DM E I +A+ VK +A EAA M+LR+DDI++
Sbjct: 1622AWIGIDGNTGSITDMKECKIWDAYNVKAQAFKTAIEAACMLLRIDDIVS 1768
>TC216674 homologue to GB|AAM26704.1|20857172|AY102137 AT3g11830/F26K24_12
{Arabidopsis thaliana;} , partial (42%)
Length = 1008
Score = 139 bits (349), Expect = 4e-33
Identities = 73/202 (36%), Positives = 120/202 (59%), Gaps = 3/202 (1%)
Frame = +2
Query: 318 LALVTGGEIASTFDNPESVKLGQCELIEEIMIGEDKLIKFSGVAMGQACTIVLRGASHHV 377
+A TGG + ++ +N LG CE+ EE +G ++ F+G GQ TIVLRG +
Sbjct: 2 VAAATGGTVQTSVNNIIDEVLGTCEIFEERQVGNERFNIFNGCPSGQTATIVLRGGADQF 181
Query: 378 LDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALARKTPGKKSLAMEAFSRA 437
++EAERSLHDA+ ++ + + +S V+ GGG +M +++ + AR GK L + ++++A
Sbjct: 182 IEEAERSLHDAIMIVRRALKNSTVVAGGGAIDMEISRYLRQHARTIAGKSQLFINSYAKA 361
Query: 438 LLAIPTTIADNAGLDSAELISQLRAEH---QNEGCTAGIDVISGSVGDMVERGICEAFKV 494
L IP + DNAG D+ +++++LR +H EG G+D+ +G + D + E V
Sbjct: 362 LEVIPRQLCDNAGFDATDVLNKLRQKHALPSGEGAPYGVDIATGGIADSFANFVWEPAVV 541
Query: 495 KQAVLLSATEAAEMILRVDDII 516
K + +ATEAA +IL VD+ I
Sbjct: 542 KINAINAATEAACLILSVDETI 607
>TC227699 similar to UP|TCPZ_HUMAN (P40227) T-complex protein 1, zeta subunit
(TCP-1-zeta) (CCT-zeta) (CCT-zeta-1) (Tcp20) (HTR3),
partial (38%)
Length = 1136
Score = 124 bits (312), Expect = 8e-29
Identities = 83/270 (30%), Positives = 141/270 (51%), Gaps = 10/270 (3%)
Frame = +1
Query: 265 AEKEKMKEKVNKII--------GHGINCFVNRQLIYNFPE-ELFADAGILAIEHADFDGI 315
AE+ ++ EKV +II G+ N V Q + P +L A GI+A+ A +
Sbjct: 37 AERRQVDEKVKRIIELKNKVCSGNDSNFVVINQKGIDPPSLDLLAREGIIALRRAKRRNM 216
Query: 316 ERLALVTGGEIASTFDNPESVKLGQCELIEEIMIGEDKLIKFSGVAMGQACTIVLRGASH 375
ERL L GGE ++ D+ LG L+ E ++GE+K V +CTI+++G +
Sbjct: 217 ERLVLACGGEAVNSVDDLTPECLGWAGLVYEHVLGEEKYTFVENVKNPFSCTILIKGPND 396
Query: 376 HVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALARKT-PGKKSLAMEAF 434
H + + + ++ D L + T+ D V+LG G E+ + + +KT G+ L +EAF
Sbjct: 397 HTIAQIKDAVRDGLRAVKNTLEDESVVLGAGAFEVAARQYLMNEVKKTVQGRAQLGVEAF 576
Query: 435 SRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCTAGIDVISGSVGDMVERGICEAFKV 494
+ ALL +P T+A+N+GLD+ ++I L EH ++G G+ + +G D GI + + V
Sbjct: 577 ADALLVVPKTLAENSGLDTQDVIIALTGEH-DKGNIVGLSLNTGEPIDPAMEGIFDNYSV 753
Query: 495 KQAVLLSATEAAEMILRVDDIITCAPRRRE 524
K+ ++ S +L VD++I R+
Sbjct: 754 KRQIINSGPVIVSQLLVVDEVIRAGRNMRK 843
>TC205278 similar to PRF|2206327A.0|1587206|2206327A T complex protein.
{Cucumis sativus;} , partial (41%)
Length = 915
Score = 122 bits (305), Expect = 5e-28
Identities = 68/203 (33%), Positives = 114/203 (55%), Gaps = 3/203 (1%)
Frame = +2
Query: 318 LALVTGGEIASTFDNPESVKLGQCELIEEIMIG--EDKLIKFSGVAMGQACTIVLRGASH 375
+A+ GG I F KLG+ ++ E G +D+++ A +A TI +RG +
Sbjct: 44 IAIXXGGXIVPRFQELSPEKLGKAGMVREKSFGTTKDRMLYIEHCANSRAVTIFIRGGNK 223
Query: 376 HVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALARKTPGKKSLAMEAFS 435
+++E +RSLHDALCV + ++ ++ GGG E+ + ++A A + PG + A+ AF
Sbjct: 224 MIIEETKRSLHDALCVARNLIRNNSIVYGGGSAEISCSIAVEAAADRYPGVEQYAIRAFG 403
Query: 436 RALLAIPTTIADNAGLDSAELISQLRAEH-QNEGCTAGIDVISGSVGDMVERGICEAFKV 494
AL AIP +A+N+GL E +S ++++ ++ GID DM E+ + E
Sbjct: 404 DALEAIPMALAENSGLQPIETLSAVKSQQIKDNNPHFGIDCNDVGTNDMREQNVFETLIG 583
Query: 495 KQAVLLSATEAAEMILRVDDIIT 517
KQ LL AT+ +MIL++DD+I+
Sbjct: 584 KQQQLLLATQVVKMILKIDDVIS 652
>TC208068 homologue to PRF|2206327A.0|1587206|2206327A T complex protein.
{Cucumis sativus;} , partial (27%)
Length = 541
Score = 117 bits (294), Expect = 1e-26
Identities = 59/125 (47%), Positives = 89/125 (71%)
Frame = +1
Query: 8 KDEASEEKGERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATILK 67
+++ S +G A+ ++ A+A +++T+LGPKGMDK+LQS +VT+TNDGATIL
Sbjct: 154 QEQKSRLRGLDAQKANISAGKAVARILRTSLGPKGMDKMLQSPDG--DVTITNDGATILD 327
Query: 68 SLHIDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRM 127
+ +DN AK++V++S+ QD E+GDGTT VVV+AG LL +AE+L+ IHP+ I G+ M
Sbjct: 328 QMDVDNQIAKLMVELSRSQDYEIGDGTTGVVVMAGALLEKAERLLERGIHPIRIAEGYEM 507
Query: 128 AAECA 132
A+ A
Sbjct: 508 ASRIA 522
>TC205279 homologue to PRF|2206327A.0|1587206|2206327A T complex protein.
{Cucumis sativus;} , partial (52%)
Length = 838
Score = 108 bits (271), Expect = 4e-24
Identities = 78/271 (28%), Positives = 134/271 (48%), Gaps = 13/271 (4%)
Frame = +3
Query: 147 AEKFRSDLMNI------ARTTLSSKILSQDKEHFAKLAVDAVMRL----KGSTNLESVQI 196
A KF D N+ TTLSSKI+++ K A++AV AV+ + + NL+ +++
Sbjct: 27 ANKFEFDESNLEPLIQTCMTTLSSKIVNRCKRSLAEIAVKAVLAVADLARKDVNLDLIKV 206
Query: 197 IKKPGGSLIDSFLDEGFILDKKIGLGQ-PKRIENAKILVANTAMDTDKVKIYGARVRVDS 255
K GG L D+ L G ++DK + Q PK+IE+AKI + + K K +V +D+
Sbjct: 207 EGKVGGKLEDTELIYGIVVDKDMSHPQMPKQIEDAKIAILTCPFEPPKPKTKH-KVDIDT 383
Query: 256 MAKVAEIEGAEKEKMKEKVNKIIGHGINCFVNRQLIYNFPEELFADAGILAIEHADFDGI 315
+ K + E++ + V K G + + + L + A+ +
Sbjct: 384 VEKFQTLRLQEQKYFDDMVQKCKDVGATLVICQWGFDDEANHLLMHRNLPAVRWVGGVEL 563
Query: 316 ERLALVTGGEIASTFDNPESVKLGQCELIEEIMIG--EDKLIKFSGVAMGQACTIVLRGA 373
E +A+ TGG I F KLG+ ++ E G +D+++ A +A TI +RG
Sbjct: 564 ELIAIATGGRIVPRFQELSPEKLGKAGMVREKSFGTTKDRMLYIEHCANSRAVTIFIRGG 743
Query: 374 SHHVLDEAERSLHDALCVLSQTVNDSRVLLG 404
+ +++E +RSLHDALCV + ++ ++ G
Sbjct: 744 NKMIIEETKRSLHDALCVARNLIRNNSIVYG 836
>TC205132 homologue to UP|CH62_CUCMA (Q05046) Chaperonin CPN60-2,
mitochondrial precursor (HSP60-2), complete
Length = 2120
Score = 105 bits (263), Expect = 4e-23
Identities = 135/540 (25%), Positives = 227/540 (42%), Gaps = 30/540 (5%)
Frame = +3
Query: 16 GERARMSSFVGAMAIADLVKTTLGPKGMDKIL-QSTGRGREVTVTNDGATILKSLH---- 70
G AR G +AD VK T+GPKG + ++ QS G + VT DG T+ KS+
Sbjct: 222 GVEARALMLKGVEELADAVKVTMGPKGRNVVIEQSFGAPK---VTKDGVTVAKSIEFKDK 392
Query: 71 IDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAE 130
+ N A ++ ++ +D GDGTT VL + E K +AA ++ M + G MA
Sbjct: 393 VKNVGASLVKQVANATNDVAGDGTTCATVLTRAIFTEGCKSIAAGMNAMDLRRGISMA-- 566
Query: 131 CARSALVEKVVDNKGDAEKFRSDLMNIART-TLSS-------KILSQDKEHFAKLAVDAV 182
V+ VV N + S IA+ T+S+ +++++ E K V +
Sbjct: 567 ------VDAVVTNLKSRARMISTSEEIAQVGTISANGEREIGELIAKAMEKVGKEGVITI 728
Query: 183 MRLKGSTN-LESVQIIKKPGGSLIDSFL----DEGFILDKKIGLGQPKRIE--NAKILVA 235
K N LE V+ +K G + F+ ++ L+ + L K+I NA + V
Sbjct: 729 SDGKTLYNELEVVEGMKLDRGYISPYFITNDKNQKCELEDPLILIHEKKISSINAIVKVL 908
Query: 236 NTAMDTDKVKIYGARVRVDSMAKVAEIEGAEKEKMKEKVNKIIGHGINCFVNRQ-LIYNF 294
A+ + + A V+S A I + +K K G G N N Q L
Sbjct: 909 ELALKRQRSLLIIAE-DVESDALATLILNKLRAGIKVCAIKAPGFGENRKANLQDLAVLT 1085
Query: 295 PEELFADAGILAIEHADFDGIERLALVTGGEIASTFDNPESVKLGQCELIEEIMIG---- 350
L + L +E D D + +T + + + K E E+I
Sbjct: 1086GGALITEELGLKLEKVDLDMLGTCKKITVSKDDTVILDGAGDKKALEERCEQIRSAIENS 1265
Query: 351 -----EDKLIKFSGVAMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGG 405
++KL + G + + GAS + E + + DAL V + ++ GG
Sbjct: 1266TSDYDKEKLQERLAKLSGGVAVLKIGGASEAEVGEKKDRVTDALNATKAAVEEG-IIPGG 1442
Query: 406 GWPEMVMAKEIDALARKTPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQ 465
G + +KE+D L +K + ++ AL TIA NAG++ A ++ +L +
Sbjct: 1443GVALLYASKELDKLQTANFDQK-IGVQIIQNALKTPVLTIASNAGVEGAVVVGKLLEQEN 1619
Query: 466 NEGCTAGIDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDIITCAPRRRED 525
++ G D G DMV+ GI + KV + L+ A + ++ + +++ P +D
Sbjct: 1620HD---LGYDAAKGEYVDMVKAGIIDPLKVIRTALVDAASVSSLMTTTEAVVSELPNDDKD 1790
>TC213696 homologue to GB|AAM26704.1|20857172|AY102137 AT3g11830/F26K24_12
{Arabidopsis thaliana;} , partial (21%)
Length = 440
Score = 104 bits (259), Expect = 1e-22
Identities = 54/107 (50%), Positives = 77/107 (71%)
Frame = +1
Query: 6 IFKDEASEEKGERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATI 65
+ K+ +G+ +S+ A+AD+V+TTLGP+GMDK++ +G VT++NDGATI
Sbjct: 115 LLKEGTDTSQGKPQLVSNINACTAVADVVRTTLGPRGMDKLIHDD-KGT-VTISNDGATI 288
Query: 66 LKSLHIDNPAAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLV 112
+K L I +PAAK+L DI+K QD EVGDGTT+VV+LA E LREA+ +
Sbjct: 289 MKLLDIVHPAAKILADIAKSQDSEVGDGTTTVVLLAAEFLREAKPFI 429
>TC228039 similar to GB|AAQ56831.1|34098897|BT010388 At3g03960 {Arabidopsis
thaliana;} , partial (50%)
Length = 1141
Score = 96.7 bits (239), Expect = 2e-20
Identities = 68/256 (26%), Positives = 113/256 (43%), Gaps = 1/256 (0%)
Frame = +2
Query: 266 EKEKMKEKVNKIIGHGINCFVNRQLIYNFPEELFADAGILAIEHADFDGIERLALVTGGE 325
E+ K++E + + G V+ + ++ ++ + + R TG
Sbjct: 32 EEAKVEELIKAVADSGAKVIVSGGAVGEMALHFCERYKLMVLKISSKFELRRFCRTTGSV 211
Query: 326 IASTFDNPESVKLGQCELIEEIMIGEDKLIKFSGVAMGQA-CTIVLRGASHHVLDEAERS 384
P LG + + IG ++ G + T+VLRG++ +LD+ ER+
Sbjct: 212 AMLKLGQPNPDDLGYVDSVSVQEIGGVRVTIVKNEEGGNSVATVVLRGSTDSILDDLERA 391
Query: 385 LHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALARKTPGKKSLAMEAFSRALLAIPTT 444
+ D + DSR + G E+ +AK + + K G A+ F+ + IP T
Sbjct: 392 VDDGVNTYKAMCRDSRFVPGAAATEIELAKRVKDFSFKETGLDQYAIAKFAESFEMIPRT 571
Query: 445 IADNAGLDSAELISQLRAEHQNEGCTAGIDVISGSVGDMVERGICEAFKVKQAVLLSATE 504
+A+NAGL++ E+IS L AEH + GID+ G D+ I + K L A +
Sbjct: 572 LAENAGLNAMEIISSLYAEHASGNAKVGIDLEEGVCKDVSTLSIWDLHVTKLFALKYAAD 751
Query: 505 AAEMILRVDDIITCAP 520
AA +LRVD II P
Sbjct: 752 AACTVLRVDQIIMAKP 799
>BE803835 similar to GP|24371053|db t-complex polypeptide 1 {Bruguiera
sexangula}, partial (32%)
Length = 541
Score = 96.7 bits (239), Expect = 2e-20
Identities = 55/169 (32%), Positives = 95/169 (55%), Gaps = 8/169 (4%)
Frame = +2
Query: 359 GVAMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDA 418
G +A T++LRGA+ H+LDE +R+LHDAL ++ +T+ + V+ GG E ++ ++
Sbjct: 11 GTKTTRAVTLILRGANDHMLDEMDRALHDALSIVKRTLESNTVVSAGGAVES*LSVYLEY 190
Query: 419 LARKTPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEH--------QNEGCT 470
LA + LA+ F+ +LL I ++ NA D+ EL+++L+A H + T
Sbjct: 191 LATPVGSTEQLAIAEFAESLLIISNVLSVNAANDATELVAKLQAYHHSVQSRADKKHLST 370
Query: 471 AGIDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDIITCA 519
G+D+ G + + ++ G+ E ++ ATEAA IL +DD+I A
Sbjct: 371 MGLDLSQGKIQNNLDAGVIEPAMSTVKIIQCATEAAITILTIDDMIKLA 517
>TC227698 similar to UP|TCPZ_HUMAN (P40227) T-complex protein 1, zeta subunit
(TCP-1-zeta) (CCT-zeta) (CCT-zeta-1) (Tcp20) (HTR3),
partial (17%)
Length = 713
Score = 88.6 bits (218), Expect = 6e-18
Identities = 49/161 (30%), Positives = 90/161 (55%), Gaps = 1/161 (0%)
Frame = +1
Query: 365 ACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALARKT- 423
+CTI+++G + H + + + ++ D L + T+ D V+LG G E+ + + +KT
Sbjct: 37 SCTILIKGPNDHTIAQIKDAVRDGLRAVKNTLEDESVVLGAGAFEVAARQYLMNEVKKTV 216
Query: 424 PGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCTAGIDVISGSVGDM 483
G+ L +EAF+ ALL +P T+A+N+GLD+ ++I L EH ++G G+ + +G D
Sbjct: 217 QGRAQLGVEAFADALLVVPKTLAENSGLDTQDVIIALTGEH-DKGNIVGLSLNTGEPIDP 393
Query: 484 VERGICEAFKVKQAVLLSATEAAEMILRVDDIITCAPRRRE 524
GI + + VK+ ++ S +L VD++I R+
Sbjct: 394 AMEGIFDNYSVKRQIINSGPVIVSQLLVVDEVIRAGRNMRK 516
>TC214150 homologue to UP|Q9ZTV1 (Q9ZTV1) Chaperonin 60 alpha subunit,
partial (98%)
Length = 2134
Score = 87.0 bits (214), Expect = 2e-17
Identities = 108/538 (20%), Positives = 230/538 (42%), Gaps = 33/538 (6%)
Frame = +3
Query: 17 ERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATILKSLHIDNP-- 74
+ +R + G +AD V TLGP+G + +L G + V NDG TI +++ + +P
Sbjct: 213 QHSRSAMQAGIDKLADAVGLTLGPRGRNVVLDEFGSPK---VVNDGVTIARAIELPDPME 383
Query: 75 --AAKVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAECA 132
A ++ +++ +D GDGTT+ VLA E+++ V + +P+++ G +
Sbjct: 384 NAGAALIREVASKTNDSAGDGTTTASVLAREIIKLGLLSVTSGANPVSLKRGIDKTVQ-- 557
Query: 133 RSALVEKVVDNKGDAEKFRSDLMNIARTTLSSKIL--SQDKEHFAKLAVDAVMRLKGSTN 190
LVE+ ++ K K D+ +A + + L E K+ D V+ ++ S++
Sbjct: 558 --GLVEE-LEKKARPVKGGDDIKAVASISAGNDELIGQMIAEAIDKVGPDGVLSIESSSS 728
Query: 191 LESVQIIKKPGGSLIDSFLDEGFILDKKIGLGQPKRIENAKILVANTAMDTDKVKI---- 246
E+ +++ G + ++ F+ + + + + ENA++L+ + + K I
Sbjct: 729 FETTVEVEE-GMEIDRGYISPQFVTNPEKLIVE---FENARVLITDQKISAIKDIIPLLE 896
Query: 247 YGARVRVDSMAKVAEIEGAEKEKMKEKVNKI-------------IGHGINCFVNRQLIYN 293
++R + ++ G + VNK+ G + I
Sbjct: 897 KTTQLRAPLLIIAEDVTGEALATL--VVNKLRGILNVAAIKAPGFGERRKALLQDIAILT 1070
Query: 294 FPEELFADAGILAIEHADFDGIERLALVTGGEIASTF----DNPESVKLGQCELIEEI-- 347
E +D G+L +E+ + + +T + ++T + ++ +L +E+
Sbjct: 1071GAEFQASDLGLL-VENTSVEQLGLARKITISKDSTTIIADAATKDELQARVAQLKKELSQ 1247
Query: 348 ---MIGEDKLIKFSGVAMGQACTIVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLG 404
+ +KL + G I + A+ L++ + + DA + + ++ G
Sbjct: 1248TDSVYDTEKLAERIAKLSGGVAVIKVGAATETELEDRKLRIEDAKNATFAAIEEG-IVPG 1424
Query: 405 GGWPEMVMAKEIDALARK-TPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAE 463
GG + ++ + A+ K + L + +AL+A IA NAG++ ++ ++++
Sbjct: 1425GGTALVHLSTHVPAIKDKLEDADERLGADIVQKALIAPAALIAQNAGIEGEVVVEKIKSG 1604
Query: 464 HQNEGCTAGIDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRVDDIITCAPR 521
G A D ++VE G+ + KV + L +A A M+L I+ P+
Sbjct: 1605EWEVGYNAMAD----RYENLVEAGVIDPAKVTRCALQNAASVAGMVLTTQAIVVEKPK 1766
>TC220855 similar to GB|AAQ56831.1|34098897|BT010388 At3g03960 {Arabidopsis
thaliana;} , partial (28%)
Length = 575
Score = 83.6 bits (205), Expect = 2e-16
Identities = 45/135 (33%), Positives = 78/135 (57%)
Frame = +2
Query: 17 ERARMSSFVGAMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATILKSLHIDNPAA 76
+ A + + ++ + +T+LGP GM+K++ ++ VTND TI+ L + +PAA
Sbjct: 161 DEAVLKNIDACKQLSTITRTSLGPNGMNKMV--INHLDKLFVTNDAGTIVNELEVQHPAA 334
Query: 77 KVLVDISKVQDDEVGDGTTSVVVLAGELLREAEKLVAAKIHPMTIIAGFRMAAECARSAL 136
KVLV K Q +E+GDG + AGELL+ AE+L+ +HP II+G+ A +
Sbjct: 335 KVLVLAGKAQQEEIGDGANLTISFAGELLQGAEELIRMGLHPSEIISGYTKAIN-KTVQI 511
Query: 137 VEKVVDNKGDAEKFR 151
++++V+N ++ R
Sbjct: 512 LDELVENGSESMDVR 556
>TC230613 homologue to UP|Q8L7N0 (Q8L7N0) TCP-1 chaperonin-like protein
(At5g16070), partial (22%)
Length = 446
Score = 79.3 bits (194), Expect = 4e-15
Identities = 38/92 (41%), Positives = 58/92 (62%)
Frame = +3
Query: 27 AMAIADLVKTTLGPKGMDKILQSTGRGREVTVTNDGATILKSLHIDNPAAKVLVDISKVQ 86
A + D++KT LGPKG K+L G ++ +T DG T+LK + I NP A ++ + Q
Sbjct: 174 AKGLQDVLKTNLGPKGTIKML--VGGAGDIKLTKDGNTLLKEMQIQNPTAIMIARTAVAQ 347
Query: 87 DDEVGDGTTSVVVLAGELLREAEKLVAAKIHP 118
DD GDGTTS V+ GEL++++E+ + +HP
Sbjct: 348 DDASGDGTTSTVIFIGELMKQSERYIDEGMHP 443
>TC234884 similar to UP|TCPA_DROME (P12613) T-complex protein 1, alpha
subunit (TCP-1-alpha) (CCT-alpha), partial (27%)
Length = 673
Score = 70.1 bits (170), Expect = 2e-12
Identities = 43/134 (32%), Positives = 71/134 (52%), Gaps = 8/134 (5%)
Frame = +2
Query: 401 VLLGGGWPEMVMAKEIDALARKTPGKKSLAMEAFSRALLAIPTTIADNAGLDSAELISQL 460
V+ GGG E ++ ++ A ++ LA+ F+++LL IP T++ NA D+ +L+++L
Sbjct: 14 VVAGGGAVEAALSIYLENFATSLSSREQLAIAEFAKSLLVIPKTLSVNAAQDATDLVAKL 193
Query: 461 RAEHQNEGCTA--------GIDVISGSVGDMVERGICEAFKVKQAVLLSATEAAEMILRV 512
R+ H + G+D+ G V D + G+ E K L ATEAA ILR+
Sbjct: 194 RSYHNSSQTKVDHANLKWIGLDLTEGVVRDNRKAGVLEPAISKIKSLKFATEAAITILRI 373
Query: 513 DDIITCAPRRREDR 526
DD+I P +E +
Sbjct: 374 DDMIKLQPSPKEQQ 415
>BI969219
Length = 756
Score = 64.7 bits (156), Expect = 1e-10
Identities = 42/158 (26%), Positives = 77/158 (48%), Gaps = 1/158 (0%)
Frame = -1
Query: 368 IVLRGASHHVLDEAERSLHDALCVLSQTVNDSRVLLGGGWPEMVMAKEIDALARK-TPGK 426
I+++G + H + + + + D L + T+ D V+LG E+ ++ +K G+
Sbjct: 720 ILIKGLNVHPIAQIKDPVRDGLXXVKNTLEDESVVLGAXXFEVAASQXXMTEVKKPVQGR 541
Query: 427 KSLAMEAFSRALLAIPTTIADNAGLDSAELISQLRAEHQNEGCTAGIDVISGSVGDMVER 486
L +EAF+ ALL + A+N+GL + ++I L EH +G G+ + G D
Sbjct: 540 AQLGVEAFADALLVVLKXXAENSGLATQDVIIALTGEHA-KGNIVGLSLTPGEPIDPAME 364
Query: 487 GICEAFKVKQAVLLSATEAAEMILRVDDIITCAPRRRE 524
GI + VK+ ++ S +L VD++I R+
Sbjct: 363 GIFANYSVKRQIINSGPVIVSQLLVVDEVIRAGRTMRK 250
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.134 0.366
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,887,168
Number of Sequences: 63676
Number of extensions: 160837
Number of successful extensions: 778
Number of sequences better than 10.0: 66
Number of HSP's better than 10.0 without gapping: 743
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 750
length of query: 527
length of database: 12,639,632
effective HSP length: 102
effective length of query: 425
effective length of database: 6,144,680
effective search space: 2611489000
effective search space used: 2611489000
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146571.2


