

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
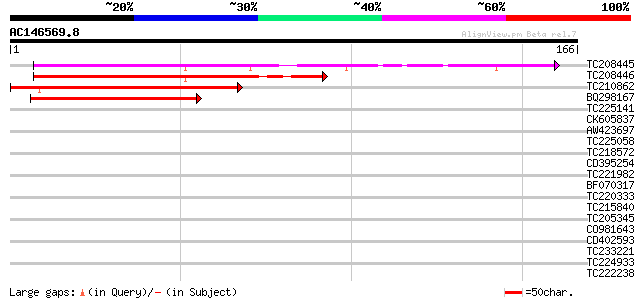
Query= AC146569.8 - phase: 0
(166 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC208445 similar to UP|Q876D1 (Q876D1) MNN4 (Fragment), partial ... 147 2e-36
TC208446 similar to UP|MNN4_YEAST (P36044) MNN4 protein, partial... 95 2e-20
TC210862 weakly similar to UP|Q9EST5 (Q9EST5) Proliferation rela... 61 3e-10
BQ298167 weakly similar to GP|5732079|gb|A T32N4.11 gene product... 47 5e-06
TC225141 type 2 metallothionein [Glycine max] 36 0.007
CK605837 34 0.037
AW423697 34 0.037
TC225058 UP|PSD6_ARATH (Q93Y35) Probable 26S proteasome non-ATPa... 33 0.048
TC218572 similar to UP|Q9FHE4 (Q9FHE4) Arabidopsis thaliana geno... 33 0.063
CD395254 similar to GP|17979464|gb| At1g68530/T26J14_10 {Arabido... 33 0.063
TC221982 similar to UP|O81326 (O81326) F3D13.3 protein, partial ... 33 0.063
BF070317 homologue to PIR|T11847|T118 gibberellin 20-oxidase (EC... 33 0.063
TC220333 weakly similar to UP|MNN4_YEAST (P36044) MNN4 protein, ... 33 0.082
TC215840 homologue to UP|Q41122 (Q41122) Proline-rich protein pr... 32 0.11
TC205345 similar to UP|Q9SZV5 (Q9SZV5) Proteinase-like protein (... 32 0.11
CO981643 32 0.14
CD402593 32 0.14
TC233221 UP|Q9NK49 (Q9NK49) Dac protein (Fragment), partial (5%) 32 0.18
TC224933 UP|Q6K853 (Q6K853) 40S ribosomal protein S30-like, comp... 32 0.18
TC222238 32 0.18
>TC208445 similar to UP|Q876D1 (Q876D1) MNN4 (Fragment), partial (9%)
Length = 1040
Score = 147 bits (371), Expect = 2e-36
Identities = 85/181 (46%), Positives = 108/181 (58%), Gaps = 27/181 (14%)
Frame = +1
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTT--------------EEV 53
GVT+VSLEGDD DRV V+G NVD+VCLA QLKKKF++VTILT EE
Sbjct: 139 GVTTVSLEGDDNDRVAVSGVNVDMVCLAYQLKKKFSSVTILTVVDLVKEEEAKKKKDEEE 318
Query: 54 KKKTEAEKKKEKEEEK-KKMIEACHAVLHGSCKCNNFNCHGKCNS--CTKCESSKCDGTN 110
KKK EAEKK+++EEE+ KKM+ + KC + +CHGKC++ CTKCES C G +
Sbjct: 319 KKKEEAEKKRKEEEERLKKMLRSVLCK-----KCKSSSCHGKCDTACCTKCESIHCGG-D 480
Query: 111 CVTICSFKCGNSKCDGNKCNGCTPKVTSPCF----------QRCPQWCTCSKCCAPYQPY 160
C +C C + KC+G+ C C ++S C CP+WC C KC PYQ
Sbjct: 481 CFIVC-VNCDSPKCEGD-CKPCINCLSSKCECECEPCPKPPSPCPKWCNCHKCYVPYQQP 654
Query: 161 C 161
C
Sbjct: 655 C 657
>TC208446 similar to UP|MNN4_YEAST (P36044) MNN4 protein, partial (3%)
Length = 565
Score = 94.7 bits (234), Expect = 2e-20
Identities = 56/101 (55%), Positives = 68/101 (66%), Gaps = 15/101 (14%)
Frame = +2
Query: 8 GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTT---------------EE 52
GVT+VSLEGDD DRV V+G NVD+VCLANQLKKKF++VTILT EE
Sbjct: 116 GVTTVSLEGDDNDRVAVSGVNVDMVCLANQLKKKFSSVTILTVVDLVKEEEAKKKKDEEE 295
Query: 53 VKKKTEAEKKKEKEEEKKKMIEACHAVLHGSCKCNNFNCHG 93
KKK EAEKK+++EEE+ K I +VL KC + +CHG
Sbjct: 296 KKKKEEAEKKRKEEEERLKKI--LRSVL--CKKCKSSSCHG 406
>TC210862 weakly similar to UP|Q9EST5 (Q9EST5) Proliferation related acidic
leucine rich protein PAL31 (Acidic nuclear
phosphoprotein 32 family, member B), partial (7%)
Length = 855
Score = 60.8 bits (146), Expect = 3e-10
Identities = 30/71 (42%), Positives = 49/71 (68%), Gaps = 3/71 (4%)
Frame = +1
Query: 1 MHIYIYI---GVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKT 57
++IYIYI GV S++ EG+ D++ VTG+ +D V L N+++KKF+ T+L+ EEVK++
Sbjct: 292 IYIYIYIKHSGVQSLAFEGETSDQLVVTGEGIDAVFLTNRMRKKFSYATLLSVEEVKEEK 471
Query: 58 EAEKKKEKEEE 68
+ + EEE
Sbjct: 472 DEGGQDGGEEE 504
>BQ298167 weakly similar to GP|5732079|gb|A T32N4.11 gene product
{Arabidopsis thaliana}, partial (32%)
Length = 421
Score = 46.6 bits (109), Expect = 5e-06
Identities = 21/50 (42%), Positives = 34/50 (68%)
Frame = +3
Query: 7 IGVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKK 56
+GV SV++EG+ KDRV V G++VD L + ++KK + ++T E+K K
Sbjct: 147 VGVNSVNIEGESKDRVVVIGEDVDAANLTSCMRKKVGHTELVTVVELKGK 296
>TC225141 type 2 metallothionein [Glycine max]
Length = 912
Score = 36.2 bits (82), Expect = 0.007
Identities = 24/81 (29%), Positives = 39/81 (47%)
Frame = +2
Query: 52 EVKKKTEAEKKKEKEEEKKKMIEACHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNC 111
E +++ E E+++E+E E+++++ +L C C F H K + C G NC
Sbjct: 8 ERERERERERERERERERERVLL*LVLLLLLFCVCVCFFSHLKMSCC---------GGNC 160
Query: 112 VTICSFKCGNSKCDGNKCNGC 132
CG+S GN C GC
Sbjct: 161 ------GCGSSCKCGNGCGGC 205
>CK605837
Length = 498
Score = 33.9 bits (76), Expect = 0.037
Identities = 17/51 (33%), Positives = 29/51 (56%)
Frame = +1
Query: 21 RVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKK 71
R+ +T +D+ Q KKK ++ KKK + +KKK+K+++KKK
Sbjct: 13 RIKITAKGIDMASSFKQKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKK 165
Score = 26.9 bits (58), Expect = 4.5
Identities = 15/40 (37%), Positives = 24/40 (59%)
Frame = +2
Query: 32 VCLANQLKKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKK 71
V L + KKK ++ KKK + +KKK+K+++KKK
Sbjct: 50 VHLNKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKK 169
Score = 26.2 bits (56), Expect = 7.7
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = +3
Query: 39 KKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKK 71
KKK ++ KKK + +KKK+K+++KKK
Sbjct: 75 KKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKK 173
Score = 26.2 bits (56), Expect = 7.7
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = +3
Query: 39 KKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKK 71
KKK ++ KKK + +KKK+K+++KKK
Sbjct: 81 KKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKK 179
Score = 26.2 bits (56), Expect = 7.7
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = +2
Query: 39 KKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKK 71
KKK ++ KKK + +KKK+K+++KKK
Sbjct: 83 KKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKK 181
Score = 26.2 bits (56), Expect = 7.7
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = +3
Query: 39 KKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKK 71
KKK ++ KKK + +KKK+K+++KKK
Sbjct: 87 KKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKK 185
Score = 26.2 bits (56), Expect = 7.7
Identities = 13/33 (39%), Positives = 21/33 (63%)
Frame = +2
Query: 39 KKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKK 71
KKK ++ KKK + +KKK+K+++KKK
Sbjct: 77 KKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKKK 175
>AW423697
Length = 417
Score = 33.9 bits (76), Expect = 0.037
Identities = 27/72 (37%), Positives = 39/72 (53%), Gaps = 3/72 (4%)
Frame = +1
Query: 3 IYIYIGVTSVSLEGDDKDRVCVTGDNVDIVCLANQLKKKFNNVTILTTEEVKK---KTEA 59
I + GV SV + + D+V V G VD L + + K+ + +E KK K E
Sbjct: 88 IIVCAGVESVETDLAN-DQVIVKGV-VDPAKLVDHVYKRTKKQASIVKDEEKKEEEKKEE 261
Query: 60 EKKKEKEEEKKK 71
EK++EKEEEKK+
Sbjct: 262 EKREEKEEEKKE 297
>TC225058 UP|PSD6_ARATH (Q93Y35) Probable 26S proteasome non-ATPase
regulatory subunit 6, partial (13%)
Length = 588
Score = 33.5 bits (75), Expect = 0.048
Identities = 16/44 (36%), Positives = 26/44 (58%)
Frame = +1
Query: 48 LTTEEVKKKTEAEKKKEKEEEKKKMIEACHAVLHGSCKCNNFNC 91
+T E KKK + +KKK+K+++KKK L SC+ + +C
Sbjct: 49 ITPESYKKKKKKKKKKKKKKKKKKKNSRLVLSLSPSCRIRHEDC 180
Score = 30.0 bits (66), Expect = 0.53
Identities = 16/41 (39%), Positives = 26/41 (63%)
Frame = +3
Query: 34 LANQLKKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKKMIE 74
L N + K NN ++ KKK + +KKK+K+++KKK +E
Sbjct: 21 LCNLICKILNNTRVI*----KKKKKKKKKKKKKKKKKKKLE 131
>TC218572 similar to UP|Q9FHE4 (Q9FHE4) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K17O22 (AT5g45100/K17O22_9),
partial (43%)
Length = 716
Score = 33.1 bits (74), Expect = 0.063
Identities = 30/135 (22%), Positives = 44/135 (32%), Gaps = 5/135 (3%)
Frame = +1
Query: 24 VTGDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKKMIEACHAVLHGS 83
V + V +C+ NQ+ K+ T ++ E E+ + S
Sbjct: 208 VLQERVKNLCVENQIWKELAQTNEATANNLRNNLEQVLAHVSEDHHHNLHHTTVEAAESS 387
Query: 84 CKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSK-CDG----NKCNGCTPKVTS 138
C NN N H + E C G+ GN K DG CN C + +
Sbjct: 388 CASNNNNSH------HREEEEVCGGS----------GNGKQSDGVLGKRMCNQCGVRESI 519
Query: 139 PCFQRCPQWCTCSKC 153
C C C+ C
Sbjct: 520 VLLLPCRHLCLCTMC 564
>CD395254 similar to GP|17979464|gb| At1g68530/T26J14_10 {Arabidopsis
thaliana}, partial (9%)
Length = 670
Score = 33.1 bits (74), Expect = 0.063
Identities = 21/57 (36%), Positives = 31/57 (53%), Gaps = 6/57 (10%)
Frame = -2
Query: 54 KKKTEAEKKKEKEEEKKKMIEACHAVLHGSCKCNN-----FNCHGKCNSCT-KCESS 104
KKK + +KKK+K+++KKK + + L SC+ + F KCNS KC S
Sbjct: 423 KKKKKKKKKKKKKKKKKKKKKTQTSSLSPSCRIRHEGQIAFGSGFKCNSAVWKCNRS 253
Score = 27.7 bits (60), Expect = 2.6
Identities = 16/43 (37%), Positives = 26/43 (60%), Gaps = 7/43 (16%)
Frame = -3
Query: 36 NQLKKKFNN-------VTILTTEEVKKKTEAEKKKEKEEEKKK 71
N K+K NN T + ++ KKK + +KKK+K+++KKK
Sbjct: 494 NCRKRKSNNG*KI*YTSTYMGLKKKKKKKKKKKKKKKKKKKKK 366
Score = 27.7 bits (60), Expect = 2.6
Identities = 12/34 (35%), Positives = 24/34 (70%)
Frame = -1
Query: 38 LKKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKK 71
+ +K+N ++ + KKK + +KKK+K+++KKK
Sbjct: 469 MDEKYNIHQLIWA*KKKKKKKKKKKKKKKKKKKK 368
Score = 26.9 bits (58), Expect = 4.5
Identities = 11/25 (44%), Positives = 20/25 (80%)
Frame = -3
Query: 48 LTTEEVKKKTEAEKKKEKEEEKKKM 72
L ++ KKK + +KKK+K+++KKK+
Sbjct: 431 LKKKKKKKKKKKKKKKKKKKKKKKL 357
Score = 26.2 bits (56), Expect = 7.7
Identities = 12/27 (44%), Positives = 19/27 (69%)
Frame = -3
Query: 54 KKKTEAEKKKEKEEEKKKMIEACHAVL 80
KKK + +KKK+K+++KKK + VL
Sbjct: 425 KKKKKKKKKKKKKKKKKKKKKKLRLVL 345
>TC221982 similar to UP|O81326 (O81326) F3D13.3 protein, partial (5%)
Length = 679
Score = 33.1 bits (74), Expect = 0.063
Identities = 21/69 (30%), Positives = 28/69 (40%)
Frame = +3
Query: 81 HGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPC 140
H S +C F+C GKC C+ CG+ C C C P +PC
Sbjct: 225 H*SHRCILFSCCGKCGDIFCKHLVH*KERTCIC*LVQACGDCHCCLFDC--CLPW*NTPC 398
Query: 141 FQRCPQWCT 149
+Q C WC+
Sbjct: 399 WQCC--WCS 419
>BF070317 homologue to PIR|T11847|T118 gibberellin 20-oxidase (EC 1.14.11.-)
- kidney bean, partial (25%)
Length = 425
Score = 33.1 bits (74), Expect = 0.063
Identities = 17/57 (29%), Positives = 29/57 (50%), Gaps = 2/57 (3%)
Frame = -3
Query: 65 KEEEKKKM--IEACHAVLHGSCKCNNFNCHGKCNSCTKCESSKCDGTNCVTICSFKC 119
+EE K M + +HG NC G+ + C + E+SK +G + ++CS+ C
Sbjct: 348 REEPPKSMRGTPSSGTFMHGFSSSGQMNCFGRLSWCLRSEASKTNGASS-SLCSWWC 181
>TC220333 weakly similar to UP|MNN4_YEAST (P36044) MNN4 protein, partial
(3%)
Length = 536
Score = 32.7 bits (73), Expect = 0.082
Identities = 17/43 (39%), Positives = 28/43 (64%)
Frame = +3
Query: 29 VDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKEKEEEKKK 71
+D V + +LKK V I++ K++ + EKK+EK+EEKK+
Sbjct: 21 MDTVKIVKKLKK-VGKVDIISVGPAKEEKKEEKKEEKKEEKKR 146
Score = 28.5 bits (62), Expect = 1.5
Identities = 12/21 (57%), Positives = 18/21 (85%)
Frame = +1
Query: 51 EEVKKKTEAEKKKEKEEEKKK 71
EE K++ + EKK+EK+EEKK+
Sbjct: 145 EEKKEEKKEEKKEEKKEEKKE 207
Score = 28.1 bits (61), Expect = 2.0
Identities = 12/20 (60%), Positives = 17/20 (85%)
Frame = +1
Query: 51 EEVKKKTEAEKKKEKEEEKK 70
EE K++ + EKK+EK+EEKK
Sbjct: 157 EEKKEEKKEEKKEEKKEEKK 216
Score = 27.3 bits (59), Expect = 3.4
Identities = 11/18 (61%), Positives = 16/18 (88%)
Frame = +1
Query: 54 KKKTEAEKKKEKEEEKKK 71
+KK + EKK+EK+EEKK+
Sbjct: 130 RKKKKEEKKEEKKEEKKE 183
>TC215840 homologue to UP|Q41122 (Q41122) Proline-rich protein precursor,
partial (74%)
Length = 1171
Score = 32.3 bits (72), Expect = 0.11
Identities = 21/64 (32%), Positives = 28/64 (42%), Gaps = 6/64 (9%)
Frame = +1
Query: 83 SCKCNNFNCHGKCNSCTKCESSKCDGTNCV------TICSFKCGNSKCDGNKCNGCTPKV 136
SC C + +C G +S T C S T+C T CS G+ C + C+G
Sbjct: 361 SCSC*STSCSGSSSS*TSCSGSPSS*TSCSGSPSS*TSCS---GSPSC*TSGCSGSPSC* 531
Query: 137 TSPC 140
TS C
Sbjct: 532 TSGC 543
>TC205345 similar to UP|Q9SZV5 (Q9SZV5) Proteinase-like protein (AT4g30020
protein) (AT4g30020/F6G3_50), partial (62%)
Length = 1729
Score = 32.3 bits (72), Expect = 0.11
Identities = 15/45 (33%), Positives = 23/45 (50%), Gaps = 4/45 (8%)
Frame = +3
Query: 71 KMIEACHAVLHGSCKCNNFNCHGKCNSCTK----CESSKCDGTNC 111
K I+ H + H C N +CH C+ C + C+ S+ +GT C
Sbjct: 1209 KTIKFKHPINHNFLSCENSSCHAHCHKCCRGGNLCDHSQ-NGTCC 1340
>CO981643
Length = 843
Score = 32.0 bits (71), Expect = 0.14
Identities = 22/76 (28%), Positives = 33/76 (42%), Gaps = 1/76 (1%)
Frame = -3
Query: 44 NVTILTTEEVKKKTEAEKKKEKEEEKKKMIEACHAVLHGSCKCNNF-NCHGKCNSCTKCE 102
NV + +E K + E E ++ EE K C C C +F +C G C C
Sbjct: 706 NVAEIANQEEKAR*EIEIPQDMEESVKCHFCGCF------CFCVDFLSCGGFC-----CG 560
Query: 103 SSKCDGTNCVTICSFK 118
++ CD C CS++
Sbjct: 559 ATGCDSLGCCIGCSYR 512
>CD402593
Length = 639
Score = 32.0 bits (71), Expect = 0.14
Identities = 16/44 (36%), Positives = 27/44 (61%)
Frame = -2
Query: 26 GDNVDIVCLANQLKKKFNNVTILTTEEVKKKTEAEKKKEKEEEK 69
GD +++ L N L N++ L E KKK + +KKK+K+++K
Sbjct: 461 GDKINVASLLNNL----NSLVSLFVHEKKKKKKKKKKKKKKKKK 342
>TC233221 UP|Q9NK49 (Q9NK49) Dac protein (Fragment), partial (5%)
Length = 651
Score = 31.6 bits (70), Expect = 0.18
Identities = 29/126 (23%), Positives = 39/126 (30%), Gaps = 19/126 (15%)
Frame = +1
Query: 48 LTTEEVKKKTEAEKKKEKEEEKKKMIEACHAVL--------HGSCKCNNFNCHGKCNSCT 99
+TTE+ KK + K C C C N + C C
Sbjct: 19 MTTEKRLKKIRQSANQSKSNTGCTCSRCCRVKCCRVKCPECSNCCCCMNVSWSINCPECC 198
Query: 100 KCESSKC----DGTNCV-TICSFKCGNSKCDGNKCNGCTPKVTSPCFQR------CPQWC 148
S KC +C +IC K C KC+ P C+ + C C
Sbjct: 199 WRPSCKCCRLPKFPSCFRSICRTKWSWPSCCATKCSLSLPSCCCCCYPKRARPSCCCMNC 378
Query: 149 TCSKCC 154
C +CC
Sbjct: 379 FCWQCC 396
Score = 30.0 bits (66), Expect = 0.53
Identities = 20/68 (29%), Positives = 26/68 (37%), Gaps = 3/68 (4%)
Frame = +1
Query: 97 SCTKCESSKCDGTNCVTICSFKCGNSKCDGNKCNGCTPKVTSPCFQRCPQWC---TCSKC 153
S T C S+C C KC KC +C+ C + CP+ C +C C
Sbjct: 73 SNTGCTCSRC--------CRVKCCRVKCP--ECSNCCCCMNVSWSINCPECCWRPSCKCC 222
Query: 154 CAPYQPYC 161
P P C
Sbjct: 223 RLPKFPSC 246
>TC224933 UP|Q6K853 (Q6K853) 40S ribosomal protein S30-like, complete
Length = 935
Score = 31.6 bits (70), Expect = 0.18
Identities = 14/29 (48%), Positives = 22/29 (75%)
Frame = +2
Query: 42 FNNVTILTTEEVKKKTEAEKKKEKEEEKK 70
FNN++IL KKK + +KKK+K+++KK
Sbjct: 497 FNNLSILLMLSSKKKKKKKKKKKKKKKKK 583
>TC222238
Length = 634
Score = 31.6 bits (70), Expect = 0.18
Identities = 21/52 (40%), Positives = 28/52 (53%), Gaps = 5/52 (9%)
Frame = -1
Query: 54 KKKTEAEKKKEKEEEKKKMIEACHAVL---HGSCKCNNFNCHG--KCNSCTK 100
K K + EK+K K +E K E + L +G C+ NNF HG K N+ TK
Sbjct: 418 KWKAKNEKRKGKTKEGSKAFE*FYQDLGTTNGKCRINNFAKHGIKKSNAWTK 263
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.134 0.459
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 12,767,018
Number of Sequences: 63676
Number of extensions: 307365
Number of successful extensions: 6192
Number of sequences better than 10.0: 229
Number of HSP's better than 10.0 without gapping: 2889
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 4273
length of query: 166
length of database: 12,639,632
effective HSP length: 90
effective length of query: 76
effective length of database: 6,908,792
effective search space: 525068192
effective search space used: 525068192
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 55 (25.8 bits)
Medicago: description of AC146569.8


