

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146568.12 + phase: 0 /pseudo
(1737 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
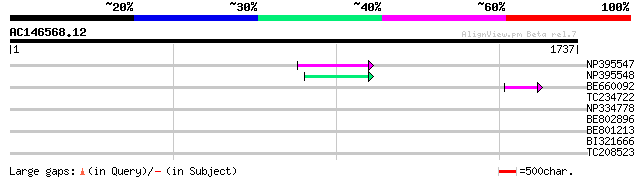
Sequences producing significant alignments: (bits) Value
NP395547 reverse transcriptase [Glycine max] 102 1e-21
NP395548 reverse transcriptase [Glycine max] 58 4e-08
BE660092 weakly similar to GP|9884624|dbj retroelement pol polyp... 48 3e-05
TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (... 43 0.001
NP334778 reverse transcriptase [Glycine max] 40 0.012
BE802896 40 0.012
BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis ... 38 0.044
BI321666 33 1.4
TC208523 similar to UP|TOB1_MOUSE (Q61471) Tob1 protein (Transdu... 30 7.0
>NP395547 reverse transcriptase [Glycine max]
Length = 762
Score = 102 bits (254), Expect = 1e-21
Identities = 92/235 (39%), Positives = 109/235 (46%)
Frame = +2
Query: 881 IVHG*VLFMLYQKKVE*WWW*MKKMS*YPPVLSLVGECA*TTGG*TPQQEKITFHYHSWI 940
I G V F L+QKK * W M +MS*+ S GEC G QEK H+ SWI
Sbjct: 56 IAPGLVRFKLFQKKEG*QW*KMIEMS*FLQEESPDGECVLIIGSSMKPQEKTITHFPSWI 235
Query: 941 K*LKG*LAKPFTVSWMGIRGIIR*RWPQKIKRRLPLLVHSVCLLIGECPSGYVVHQQPFN 1000
K L+ P TVSW + IR +W +IK++ L V SV LLI C S YV+ F
Sbjct: 236 KCLRDLQGNPSTVSWTDTQVTIRLQWILRIKKKQLLHVLSVFLLIAACRSVYVMPLLLFR 415
Query: 1001 GACSPSSLT**RRTLRYLWTISLYLVSRLTNACFISMLSLKDVPKLTLF*TGKNVILW*L 1060
+T *R L+ LW I LV L A I V L TGKNV LW
Sbjct: 416 DV*WQFLMTW*RNVLKSLWMIFRSLVHLLEIA*QI*RKCYNVVKNLIWCLTGKNVTLWYK 595
Query: 1061 KV*FWVIK*VLKA*RWIKLR*RL*KNFHHL*MLRG*GVFWGMPVSIGGL*RIFQK 1115
KV K + + RW+K * L NFH M + VFW M IG *R K
Sbjct: 596 KVLC*DTKSLKEELRWLKKN*MLLINFHPQLM*KAYTVFWVMLDFIGDS*RTSPK 760
>NP395548 reverse transcriptase [Glycine max]
Length = 762
Score = 57.8 bits (138), Expect = 4e-08
Identities = 69/214 (32%), Positives = 84/214 (39%)
Frame = +2
Query: 902 MKKMS*YPPVLSLVGECA*TTGG*TPQQEKITFHYHSWIK*LKG*LAKPFTVSWMGIRGI 961
MK+M+*Y LSLVG A T T Q K Y SWI+ + SWM I
Sbjct: 119 MKRMT*YQHELSLVGNYASITASSTKPQGKTISLYPSWIRCWRDLQDTLIIASWMHTLDI 298
Query: 962 IR*RWPQKIKRRLPLLVHSVCLLIGECPSGYVVHQQPFNGACSPSSLT**RRTLRYLWTI 1021
IR +I+RR P V L I G +H AC P * R+ +Y W
Sbjct: 299 IRLL*TPRIRRRWPSHALLVSLPIDGFHLGCAMHLPHSKCACWPFLQI*WRKASKYSWMT 478
Query: 1022 SLYLVSRLTNACFISMLSLKDVPKLTLF*TGKNVILW*LKV*FWVIK*VLKA*RWIKLR* 1081
YL KD K T +* G++V W K * IK + R K R
Sbjct: 479 FQYLCPH*KVV*RSWRWYYKDAWKQT*Y*IGRSVTSWFEKA*S*AIKFRPEELR*TKQRL 658
Query: 1082 RL*KNFHHL*MLRG*GVFWGMPVSIGGL*RIFQK 1115
K+ HH ML+ G P S R QK
Sbjct: 659 MSLKSCHHHQMLKASGAS*DKPGSTEDSSRTSQK 760
>BE660092 weakly similar to GP|9884624|dbj retroelement pol polyprotein-like
{Arabidopsis thaliana}, partial (13%)
Length = 378
Score = 48.1 bits (113), Expect = 3e-05
Identities = 44/114 (38%), Positives = 57/114 (49%)
Frame = -1
Query: 1517 CLKSTTLNIKLQLPTTLKLVVK*RFPTVS*NKFWRKL*RLLGKIGLGS*TTLFGLIGLLL 1576
CLKST TT + + + +F T +F R+L GKIG+ L G IGL
Sbjct: 366 CLKSTGWYTGYPHHTTPRPMDRQKFLTGKSREF*RRLCSQAGKIGVPGLMMLSGHIGLPT 187
Query: 1577 KRILGCHLINWSLAKRVIFQ*S*STKLIGPSKLLILIKL*LVRKDS*SSMS*KK 1630
K C LI SL + VIFQ STK G +L + + LVRK S + +S* K
Sbjct: 186 KHP*ECLLIGLSLERHVIFQWKLSTKHTGQ*RLATSLWIKLVRKGSCN*VS*MK 25
>TC234722 similar to UP|Q6WAY9 (Q6WAY9) Pol (Fragment), partial (32%)
Length = 482
Score = 42.7 bits (99), Expect = 0.001
Identities = 30/74 (40%), Positives = 39/74 (52%)
Frame = -1
Query: 1534 KLVVK*RFPTVS*NKFWRKL*RLLGKIGLGS*TTLFGLIGLLLKRILGCHLINWSLAKRV 1593
K + K +F *+++ R L KIGL S L GL GL + CH NWS+ K
Sbjct: 224 KQMAKQKFQIKR*SEYLRILLCHQEKIGL*SWMMLSGLTGLHSRLP*ACHHFNWSMVKHA 45
Query: 1594 IFQ*S*STKLIGPS 1607
++ S STK IGPS
Sbjct: 44 TYRWSWSTKPIGPS 3
>NP334778 reverse transcriptase [Glycine max]
Length = 431
Score = 39.7 bits (91), Expect = 0.012
Identities = 27/70 (38%), Positives = 36/70 (50%)
Frame = +1
Query: 950 PFTVSWMGIRGIIR*RWPQKIKRRLPLLVHSVCLLIGECPSGYVVHQQPFNGACSPSSLT 1009
P+++SWM R IIR*RW Q+I +R L++ I C G + QP G S T
Sbjct: 103 PYSLSWMDFRVIIR*RWHQRIWKRQLSLLYGEPSAIR*CRLG*RILGQPTIGPWWHYSRT 282
Query: 1010 **RRTLRYLW 1019
* R R +W
Sbjct: 283 *CTRRSRPMW 312
>BE802896
Length = 416
Score = 39.7 bits (91), Expect = 0.012
Identities = 44/125 (35%), Positives = 57/125 (45%)
Frame = -3
Query: 1098 VFWGMPVSIGGL*RIFQK*PSPSITF*LKKLSLILMLHV*MLFL*LKTIW*LHQ*SLHPI 1157
+F M S G L* I +K*P T ++ SL LM++ L + K S HP
Sbjct: 414 LFLVMQDSTGAL*EILEK*PFHYPTCCKRR*SLTLMINANRLLIASKEH*LPPPSSRHPT 235
Query: 1158 GIYTLNLCVMQVIMLLVQS*ANTRTNFFMQFTMQARF*MKVK*TIQQLKKSCLQ*YLHWK 1217
G L+LCVM IM S N + + *M K I L+KS *+L K
Sbjct: 234 GQPLLSLCVMHQIMHWGLSLLRKLINCLG*YIILLGL*MLPKRIILLLRKSY*P*FLLLK 55
Query: 1218 NFVLI 1222
NF+LI
Sbjct: 54 NFILI 40
>BE801213 weakly similar to GP|6691193|gb| F7F22.17 {Arabidopsis thaliana},
partial (3%)
Length = 416
Score = 37.7 bits (86), Expect = 0.044
Identities = 29/63 (46%), Positives = 36/63 (57%)
Frame = +3
Query: 1551 RKL*RLLGKIGLGS*TTLFGLIGLLLKRILGCHLINWSLAKRVIFQ*S*STKLIGPSKLL 1610
+K+ L KIG +* LFG LL K L WS+ + VI+ *S*STK IGP +
Sbjct: 225 KKMWHLQEKIGHPN*RMLFGHARLLKKLQLV*LHSKWSIGRLVIYP*S*STKHIGP*SV* 404
Query: 1611 ILI 1613
ILI
Sbjct: 405 ILI 413
>BI321666
Length = 430
Score = 32.7 bits (73), Expect = 1.4
Identities = 20/45 (44%), Positives = 25/45 (55%)
Frame = +3
Query: 1402 FSRIAMIL*GGVTIAKEQVVFLRGMRCP*RESLKLNHLTVGALIL 1446
FS++ M + V AKE VF RGM C + LT+GALIL
Sbjct: 45 FSKMHMFMLSHVINAKELEVFPRGMNCHCIPFWRWRSLTIGALIL 179
>TC208523 similar to UP|TOB1_MOUSE (Q61471) Tob1 protein (Transducer of
erbB-2 1), partial (5%)
Length = 665
Score = 30.4 bits (67), Expect = 7.0
Identities = 18/53 (33%), Positives = 26/53 (48%), Gaps = 1/53 (1%)
Frame = -3
Query: 475 PKRRRRRKDNSRNSC-NSSHNFR*IFLLVMLLIKCRCMLNS*KKC*RGEENQK 526
PKR RRR+ R C H + LL++L ++ R L S K R + +K
Sbjct: 219 PKRHRRRRQLKRRRCLRRKHRLLLLLLLLLLRLRLRLRLKSQKSVWRLSQGRK 61
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.369 0.165 0.601
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 76,341,265
Number of Sequences: 63676
Number of extensions: 1071485
Number of successful extensions: 13089
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 5139
Number of HSP's successfully gapped in prelim test: 400
Number of HSP's that attempted gapping in prelim test: 7640
Number of HSP's gapped (non-prelim): 6139
length of query: 1737
length of database: 12,639,632
effective HSP length: 111
effective length of query: 1626
effective length of database: 5,571,596
effective search space: 9059415096
effective search space used: 9059415096
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 14 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 36 (21.7 bits)
S2: 66 (30.0 bits)
Medicago: description of AC146568.12


