

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146567.11 - phase: 0
(331 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
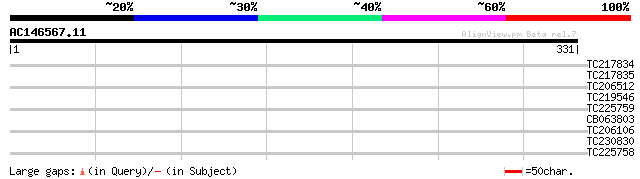
Score E
Sequences producing significant alignments: (bits) Value
TC217834 similar to UP|D111_ARATH (P42698) DNA-damage-repair/tol... 35 0.061
TC217835 homologue to UP|D111_ARATH (P42698) DNA-damage-repair/t... 34 0.10
TC206512 similar to UP|Q9FV81 (Q9FV81) Glu-tRNA(Gln) amidotransf... 32 0.52
TC219546 UP|HS70_SOYBN (P26413) Heat shock 70 kDa protein, complete 31 0.68
TC225759 similar to UP|Q9SMK5 (Q9SMK5) Plasma membrane intrinsic... 31 0.89
CB063803 30 1.2
TC206106 weakly similar to UP|Q7RK82 (Q7RK82) Maebl (Fragment), ... 28 5.7
TC230830 similar to UP|Q940E5 (Q940E5) Wound-responsive protein ... 27 9.8
TC225758 similar to UP|Q9SMK5 (Q9SMK5) Plasma membrane intrinsic... 27 9.8
>TC217834 similar to UP|D111_ARATH (P42698) DNA-damage-repair/toleration
protein DRT111, chloroplast precursor, partial (57%)
Length = 1006
Score = 34.7 bits (78), Expect = 0.061
Identities = 18/60 (30%), Positives = 36/60 (60%), Gaps = 4/60 (6%)
Frame = +2
Query: 143 QIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIP--ESIRHNFEQME--EERTKVLIAIE 198
++D++++D + +C +Y ++ +T+PN P E++R F Q E EE TK L+ ++
Sbjct: 449 EVDDELEDEVGSECAKYGTVTRVLIFEITEPNFPVHEAVR-IFVQFERSEETTKALVDLD 625
>TC217835 homologue to UP|D111_ARATH (P42698) DNA-damage-repair/toleration
protein DRT111, chloroplast precursor, partial (37%)
Length = 695
Score = 33.9 bits (76), Expect = 0.10
Identities = 18/60 (30%), Positives = 36/60 (60%), Gaps = 4/60 (6%)
Frame = +2
Query: 143 QIDEKMKDALQVDCTRYAPGIEIIGVRVTKPNIP--ESIRHNFEQME--EERTKVLIAIE 198
++D++++D + +C +Y ++ +T+PN P E++R F Q E EE TK L+ ++
Sbjct: 158 EVDDELEDEVGSECAKYGIVTRVLIFEITEPNFPVHEAVR-IFVQFERSEETTKALVDLD 334
>TC206512 similar to UP|Q9FV81 (Q9FV81) Glu-tRNA(Gln) amidotransferase
subunit B (At1g48520/T1N15_12), partial (74%)
Length = 1626
Score = 31.6 bits (70), Expect = 0.52
Identities = 35/165 (21%), Positives = 73/165 (44%), Gaps = 5/165 (3%)
Frame = +1
Query: 167 GVRVTKPNIPESIRHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKN---AN 223
G++ + P +PE R +EQM VL + ++E T+ K A S+ N ++
Sbjct: 667 GIKNSLPELPEIKRRRYEQMGLSMQDVLFLANDKNIAEFFDATLAKGADSKLVANWIMSD 846
Query: 224 VSKILMEQKLSEKDSARRQEEIENAMYLAREKSLADADFYRVIKE--AEANRLKLTPEFL 281
++ + +KL+ + EE+ + + +++ ++ E A+ +K E +
Sbjct: 847 IAAFMKNEKLTINEIKLTPEELSELIASIKGGTISGKIGKEILFELLAKGGTVK---ELI 1017
Query: 282 ELKFIESIANNTKIFFGDKIPNMILDQRLLGNFLVEEVPRGAATK 326
+ K + IA+ +I ++D+ ++ N E RG TK
Sbjct: 1018QKKDLVQIADPVEI-------EKMVDKVIVENPKQVEQYRGGKTK 1131
>TC219546 UP|HS70_SOYBN (P26413) Heat shock 70 kDa protein, complete
Length = 2099
Score = 31.2 bits (69), Expect = 0.68
Identities = 41/189 (21%), Positives = 81/189 (42%), Gaps = 13/189 (6%)
Frame = +1
Query: 143 QIDEKMKDALQVDCTRYAPGIEIIGVRVT-----KPNIPESIRHNFEQMEEERTKVLIAI 197
Q DEK++D L +D T + G+E G +T IP F + + VLI +
Sbjct: 1162 QGDEKVQDLLLLDVTPLSLGLETAGGVMTVLIPRNTTIPTKKEQIFSTYSDNQPGVLIQV 1341
Query: 198 -EKQKVSEKEAETMKKMAIS------EAEKNANVSKILMEQKLSEKDSARRQEEIENAMY 250
E ++ K+ + K ++ NV + + + + ++N +
Sbjct: 1342 FEGERARTKDNNLLGKFELTGIPPAPRGVPQINVCFDIDANGILNVSAEDKTAGVKNKIT 1521
Query: 251 LAREKS-LADADFYRVIKEAEANRLKLTPEFLELKFIESIANNTKIFFGDKIPNMILDQR 309
+ +K L+ + +++K+AE R K E ++ K +E A N+ + + N I D++
Sbjct: 1522 ITNDKGRLSKEEIEKMVKDAE--RYKAEDEEVKKK-VE--AKNSLENYAYNMRNTIKDEK 1686
Query: 310 LLGNFLVEE 318
+ G +E
Sbjct: 1687 IGGKLSPDE 1713
>TC225759 similar to UP|Q9SMK5 (Q9SMK5) Plasma membrane intrinsic
polypeptide, partial (74%)
Length = 1051
Score = 30.8 bits (68), Expect = 0.89
Identities = 17/41 (41%), Positives = 25/41 (60%)
Frame = +1
Query: 184 EQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANV 224
E+ EEE+ +V+ +++KV EK AET K AE+ A V
Sbjct: 619 EKKEEEKPQVIETSDEKKVEEKPAETSAKGEEKPAEEAAAV 741
>CB063803
Length = 411
Score = 30.4 bits (67), Expect = 1.2
Identities = 13/36 (36%), Positives = 21/36 (58%)
Frame = +3
Query: 92 VNRLHKESVYETLLNYGVQYDKTWIYDKIHHEINQF 127
++ + K +V + N G +Y KT+IY +HH QF
Sbjct: 12 LDAVDKVTVMDLF*NLGYEYFKTYIYTVVHHICTQF 119
>TC206106 weakly similar to UP|Q7RK82 (Q7RK82) Maebl (Fragment), partial (7%)
Length = 1040
Score = 28.1 bits (61), Expect = 5.7
Identities = 24/109 (22%), Positives = 52/109 (47%)
Frame = +2
Query: 176 PESIRHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKLSE 235
PE + + +E++ +V ++ +V+E++ K+ ++E +K A V++ E +++E
Sbjct: 659 PEKKKTELKALEKKEEEVSEEKKEVEVTEEK----KEAEVTEEKKEAEVTEEKKEVEVAE 826
Query: 236 KDSARRQEEIENAMYLAREKSLADADFYRVIKEAEANRLKLTPEFLELK 284
+ E + + +A EK + + + KE E K E E K
Sbjct: 827 EKKEVEGTEEKKEVEVAEEKK--ETEVIKEKKEVEVTEEKKEVEVTEEK 967
>TC230830 similar to UP|Q940E5 (Q940E5) Wound-responsive protein 10.1
(Fragment), partial (36%)
Length = 771
Score = 27.3 bits (59), Expect = 9.8
Identities = 20/59 (33%), Positives = 32/59 (53%)
Frame = +1
Query: 175 IPESIRHNFEQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVSKILMEQKL 233
IPES FE+ E+E E + +S++E E K+M EK +++S I +QK+
Sbjct: 103 IPESEEEEFEEQEDE--------EPEVISQRENEN-KEM----GEKYSSISNIGTDQKV 240
>TC225758 similar to UP|Q9SMK5 (Q9SMK5) Plasma membrane intrinsic
polypeptide, partial (74%)
Length = 1087
Score = 27.3 bits (59), Expect = 9.8
Identities = 21/56 (37%), Positives = 29/56 (51%), Gaps = 6/56 (10%)
Frame = +1
Query: 184 EQMEEERTKVLIAIEKQKVSEKEAETMKKMAISEAEKNANVS------KILMEQKL 233
E+ EEE+ + +++KV EK+AET A E EK A + K EQKL
Sbjct: 670 EKKEEEKPQADETSDEKKVEEKQAET----AAKEEEKPAEPAEPPKP*KFRSEQKL 825
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.318 0.135 0.378
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,546,252
Number of Sequences: 63676
Number of extensions: 133692
Number of successful extensions: 729
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 724
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 727
length of query: 331
length of database: 12,639,632
effective HSP length: 98
effective length of query: 233
effective length of database: 6,399,384
effective search space: 1491056472
effective search space used: 1491056472
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146567.11


