

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146567.10 + phase: 0
(393 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
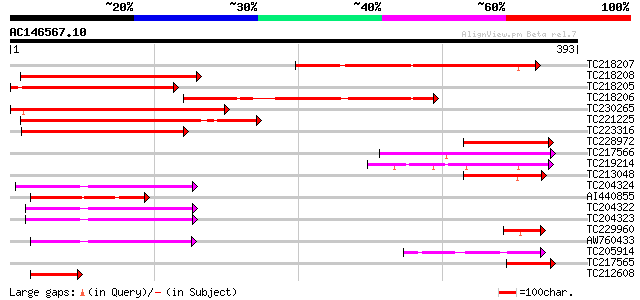
Score E
Sequences producing significant alignments: (bits) Value
TC218207 weakly similar to GB|AAO23575.1|27764908|BT003010 At5g6... 235 3e-62
TC218208 weakly similar to GB|AAO23575.1|27764908|BT003010 At5g6... 214 7e-56
TC218205 weakly similar to GB|AAO23575.1|27764908|BT003010 At5g6... 193 1e-49
TC218206 192 2e-49
TC230265 similar to GB|AAO23575.1|27764908|BT003010 At5g60980/MS... 179 3e-45
TC221225 weakly similar to UP|Q8VYJ4 (Q8VYJ4) AT3g25150/MJL12_9,... 161 5e-40
TC223316 weakly similar to GB|AAO23575.1|27764908|BT003010 At5g6... 160 1e-39
TC228972 similar to UP|Q8LEI7 (Q8LEI7) Ras-GTPase-activating pro... 83 2e-16
TC217566 similar to UP|Q8VYJ4 (Q8VYJ4) AT3g25150/MJL12_9, partia... 80 1e-15
TC219214 72 4e-13
TC213048 62 6e-10
TC204324 similar to UP|NTF2_ARATH (Q9C7F5) Nuclear transport fac... 57 1e-08
AI440855 57 1e-08
TC204322 similar to UP|NTF2_ARATH (Q9C7F5) Nuclear transport fac... 56 3e-08
TC204323 similar to UP|NTF2_ARATH (Q9C7F5) Nuclear transport fac... 56 3e-08
TC229960 similar to GB|AAO23575.1|27764908|BT003010 At5g60980/MS... 51 1e-06
AW760433 similar to PIR|H86398|H863 protein F17L21.10 [imported]... 49 3e-06
TC205914 weakly similar to UP|NAM8_YEAST (Q00539) NAM8 protein, ... 45 4e-05
TC217565 similar to UP|Q8VYJ4 (Q8VYJ4) AT3g25150/MJL12_9, partia... 44 1e-04
TC212608 43 2e-04
>TC218207 weakly similar to GB|AAO23575.1|27764908|BT003010
At5g60980/MSL3_100 {Arabidopsis thaliana;} , partial
(10%)
Length = 509
Score = 235 bits (599), Expect = 3e-62
Identities = 125/173 (72%), Positives = 141/173 (81%), Gaps = 3/173 (1%)
Frame = +2
Query: 199 ISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTSIEKVASNTQ-EDTPKKSFA 257
+SKEVS PLENG +SVTE V+PVNHVKESSH + EK ASN EDTPKKSFA
Sbjct: 2 VSKEVSQPLENGNVSVTEKVVPVNHVKESSH---QEHSHYHAEKAASNNALEDTPKKSFA 172
Query: 258 SIVNALKDNSAPFHLRASPAKPAVHPPRVHSVPAPEAPTPNMDIPLEKNNENAGRAHAIF 317
SIVNALK+N+APFH+R SP K V PRV S+PAPEAP P+++ P EKNNEN G+A+AIF
Sbjct: 173 SIVNALKENAAPFHVRVSPVK-LVEQPRVSSIPAPEAPAPSIESPPEKNNENGGKAYAIF 349
Query: 318 VANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNK--GSCFGFVEFESAASMQS 368
VANLPM+ATVEQL+RAFKKFGPIK+DGIQVRSNK SCFGFVEFESA SMQS
Sbjct: 350 VANLPMNATVEQLERAFKKFGPIKQDGIQVRSNKQQQSCFGFVEFESATSMQS 508
>TC218208 weakly similar to GB|AAO23575.1|27764908|BT003010
At5g60980/MSL3_100 {Arabidopsis thaliana;} , partial
(25%)
Length = 436
Score = 214 bits (544), Expect = 7e-56
Identities = 100/126 (79%), Positives = 117/126 (92%)
Frame = +3
Query: 8 QTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQ 67
++ T QM+GNAFV+QYYSILH++PDQVHRFY +SS++SRPEEDGTMT VTTT EI+KKI
Sbjct: 54 ESSTTQMIGNAFVQQYYSILHQEPDQVHRFYQESSILSRPEEDGTMTMVTTTLEINKKIL 233
Query: 68 SLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLND 127
SL+YTSFRVE+LSADAQPSY +GV+VVVTGCLTG+DN+KRKF QSFFLAPQDKG+ VLND
Sbjct: 234 SLDYTSFRVEILSADAQPSYKDGVIVVVTGCLTGSDNLKRKFTQSFFLAPQDKGYCVLND 413
Query: 128 VFRYVD 133
VFRYVD
Sbjct: 414 VFRYVD 431
>TC218205 weakly similar to GB|AAO23575.1|27764908|BT003010
At5g60980/MSL3_100 {Arabidopsis thaliana;} , partial
(22%)
Length = 432
Score = 193 bits (490), Expect = 1e-49
Identities = 94/117 (80%), Positives = 106/117 (90%)
Frame = +3
Query: 1 MAVPEAVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTA 60
MAV E +PTPQ VGNAFVEQYYSILH+ PDQVHRFYH+SS++SRPEEDGTMT VTTT
Sbjct: 87 MAVSEG--SPTPQTVGNAFVEQYYSILHQKPDQVHRFYHESSILSRPEEDGTMTMVTTTL 260
Query: 61 EIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAP 117
EI+KKI SL+YTSFRVE+LSADAQPSY +GV+VVVTGCLTG+DN+KRKF QSFFLAP
Sbjct: 261 EINKKILSLDYTSFRVEILSADAQPSYKDGVIVVVTGCLTGSDNLKRKFTQSFFLAP 431
>TC218206
Length = 503
Score = 192 bits (489), Expect = 2e-49
Identities = 107/179 (59%), Positives = 127/179 (70%), Gaps = 2/179 (1%)
Frame = +1
Query: 121 GFYVLNDVFRYVDAYKSIDIESVPAND-ADESAPSEAIITPEPEPVHVPEVIPPTQTVIP 179
G++VLNDVFRYVD YKS+DIESVPAND ADESAP++A + PEPE +HV E
Sbjct: 1 GYFVLNDVFRYVDEYKSVDIESVPANDAADESAPTDAFV-PEPEAIHVAE---------- 147
Query: 180 TAQTVIPPTQTVIADTETIISKEVSLPLENGKLSVTENVIPVNHVKESSHHVKEPEQPTS 239
+P +QT + D + +SKEVS PLENG LSVTE V+PV+HVKE SH E +
Sbjct: 148 ----DVPASQTDVVDADIGVSKEVSQPLENGNLSVTEKVVPVDHVKECSHQ----EHHSH 303
Query: 240 IEKVASNTQ-EDTPKKSFASIVNALKDNSAPFHLRASPAKPAVHPPRVHSVPAPEAPTP 297
EK ASN EDTPKKSFASIVNALK+N+APFH+R SP K + PRV S+PAPEAP P
Sbjct: 304 AEKAASNNSLEDTPKKSFASIVNALKENAAPFHVRVSPVK-LLEQPRVSSIPAPEAPAP 477
>TC230265 similar to GB|AAO23575.1|27764908|BT003010 At5g60980/MSL3_100
{Arabidopsis thaliana;} , partial (27%)
Length = 599
Score = 179 bits (453), Expect = 3e-45
Identities = 85/154 (55%), Positives = 117/154 (75%), Gaps = 2/154 (1%)
Frame = +2
Query: 1 MAVPEAVQ--TPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTT 58
MA+PE + TP+ Q+VGNAFVEQYY ILH+ P+ VHRFY DSS ++R + +G MTTVTT
Sbjct: 137 MAMPETIPPTTPSAQVVGNAFVEQYYHILHQSPELVHRFYQDSSFLTRSDSNGVMTTVTT 316
Query: 59 TAEIDKKIQSLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQ 118
EI +KI SL+Y + E+ +ADAQ S+ GV+V+VTGCLTG DN++RKF+Q+FFLAPQ
Sbjct: 317 VQEIHEKIISLKYEDYTAEIKTADAQESHKGGVIVLVTGCLTGKDNVRRKFSQTFFLAPQ 496
Query: 119 DKGFYVLNDVFRYVDAYKSIDIESVPANDADESA 152
+KG+YVLNDVFR+++ + + S + +E+A
Sbjct: 497 EKGYYVLNDVFRFIEENDTPQLNSSTVSVINENA 598
>TC221225 weakly similar to UP|Q8VYJ4 (Q8VYJ4) AT3g25150/MJL12_9, partial
(27%)
Length = 745
Score = 161 bits (407), Expect = 5e-40
Identities = 84/167 (50%), Positives = 109/167 (64%)
Frame = +2
Query: 8 QTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQ 67
QTP +VGNAFV+QYY +LH P+ VHRFY D S + RPE++G M TT +I+KKI
Sbjct: 236 QTPAADIVGNAFVDQYYHMLHESPELVHRFYQDVSKLGRPEQNGIMGITTTMFDINKKIL 415
Query: 68 SLEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYVLND 127
SL Y E++S DAQ SY+ GV+V+VTG + G D+IK+KF Q FFLAPQ+KG++VLND
Sbjct: 416 SLGYGELSAEIVSVDAQESYDGGVIVLVTGFMIGKDDIKQKFTQCFFLAPQEKGYFVLND 595
Query: 128 VFRYVDAYKSIDIESVPANDADESAPSEAIITPEPEPVHVPEVIPPT 174
VFRYVD I+ A+D +AP + + P V E I T
Sbjct: 596 VFRYVD---ENGIQG-SAHDIGSAAPPDTVANPSVLETQVCEQISVT 724
>TC223316 weakly similar to GB|AAO23575.1|27764908|BT003010
At5g60980/MSL3_100 {Arabidopsis thaliana;} , partial
(25%)
Length = 475
Score = 160 bits (404), Expect = 1e-39
Identities = 75/116 (64%), Positives = 92/116 (78%)
Frame = +1
Query: 9 TPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQS 68
TP+ Q+VGNAFVEQYY ILH PD V+RFY DSSV+SRP+ G MT+VTT I++KI S
Sbjct: 127 TPSAQVVGNAFVEQYYHILHHSPDLVYRFYQDSSVISRPDSSGVMTSVTTMKGINEKILS 306
Query: 69 LEYTSFRVEVLSADAQPSYNNGVMVVVTGCLTGTDNIKRKFAQSFFLAPQDKGFYV 124
L + F+ E+ +ADAQ SY GV V+VTGCLTG DN++RKFAQSFFLAPQD G++V
Sbjct: 307 LNFKEFKAEIKTADAQKSYKEGVTVLVTGCLTGKDNLRRKFAQSFFLAPQDNGYFV 474
>TC228972 similar to UP|Q8LEI7 (Q8LEI7) Ras-GTPase-activating protein
SH3-domain binding protein-like, partial (19%)
Length = 918
Score = 82.8 bits (203), Expect = 2e-16
Identities = 41/64 (64%), Positives = 50/64 (78%), Gaps = 1/64 (1%)
Frame = +3
Query: 315 AIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNK-GSCFGFVEFESAASMQSALEVW 373
+I++ NLP +ATVEQL+ FKKFGPIK GIQVRS+K G CFGFVEFE +SM SALE
Sbjct: 6 SIYIRNLPFNATVEQLEEVFKKFGPIKHGGIQVRSSKHGFCFGFVEFEELSSMHSALEAS 185
Query: 374 SISI 377
I++
Sbjct: 186 PITV 197
>TC217566 similar to UP|Q8VYJ4 (Q8VYJ4) AT3g25150/MJL12_9, partial (20%)
Length = 1031
Score = 80.5 bits (197), Expect = 1e-15
Identities = 50/130 (38%), Positives = 67/130 (51%), Gaps = 8/130 (6%)
Frame = +3
Query: 257 ASIVNALKDNSAPFHLRASPAKPAVHPPRVHSVPAPEAPTPNMDI--------PLEKNNE 308
A IV +K+ +AP + + H + A P+ + + N E
Sbjct: 9 AYIVKVMKEGAAPSSTVTPVSVKSAHKSQEQQGIAAPPPSSISETNGSIINTNEVGNNQE 188
Query: 309 NAGRAHAIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNKGSCFGFVEFESAASMQS 368
++I+V LP +AT L+ FKKFGPIK GIQVRS KG +GFVEFE A++ QS
Sbjct: 189 TEAEGYSIYVKGLPPTATPAVLENEFKKFGPIKSGGIQVRSQKGFSYGFVEFEVASAAQS 368
Query: 369 ALEVWSISIN 378
ALE ISIN
Sbjct: 369 ALEASPISIN 398
>TC219214
Length = 1127
Score = 72.0 bits (175), Expect = 4e-13
Identities = 52/146 (35%), Positives = 77/146 (52%), Gaps = 17/146 (11%)
Frame = +2
Query: 249 EDTPKKSFASIVNALKD----NSAPFHLRASPAKPAVHPPRVHSVPAP---EAPTPNMDI 301
E+ PKK++ASI+ K ++AP H S K A PP ++ V P ++ + +M
Sbjct: 50 EEPPKKTYASILRVSKGLPVLSAAPKHAPHS-FKSAPPPPELNHVAQPAVQQSSSASMYA 226
Query: 302 PLEKNNENAGRAHA--------IFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSNK-- 351
P E E A +A ++V NLP + T ++D+ FK FG IK DGI +R K
Sbjct: 227 P-ESGTEAAEEGYALEEDEVTSVYVRNLPANVTEVEIDQEFKNFGRIKPDGIFIRVRKEI 403
Query: 352 GSCFGFVEFESAASMQSALEVWSISI 377
G C+ FVEFE +Q+AL+ I +
Sbjct: 404 GVCYAFVEFEDIIGVQNALQASPIQL 481
>TC213048
Length = 754
Score = 61.6 bits (148), Expect = 6e-10
Identities = 32/60 (53%), Positives = 41/60 (68%), Gaps = 2/60 (3%)
Frame = +1
Query: 315 AIFVANLPMSATVEQLDRAFKKFGPIKRDGIQVRSN--KGSCFGFVEFESAASMQSALEV 372
+I NLP++ T QL+ FKKFG I GIQVR+N +G CFGFVEF S SM SA+++
Sbjct: 1 SIXXXNLPLNVTAAQLELEFKKFGXIXXXGIQVRNNXXQGYCFGFVEFLSXNSMNSAIQL 180
>TC204324 similar to UP|NTF2_ARATH (Q9C7F5) Nuclear transport factor 2
(NTF-2), complete
Length = 772
Score = 57.0 bits (136), Expect = 1e-08
Identities = 41/129 (31%), Positives = 64/129 (48%), Gaps = 3/129 (2%)
Frame = +1
Query: 5 EAVQTPTPQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDK 64
EA P + AFVE YYS + + + Y + S++S + + + I
Sbjct: 88 EARSDMDPDALAKAFVEHYYSTFDTNRNNLANLYQEGSMLSFEGQ-----KIQGSHNIVA 252
Query: 65 KIQSLEYTSFRVEVLSADAQPSYNNGVMVV-VTGCL-TGTDNIKRKFAQSFFLAPQDKG- 121
K+ SL + + + + D+QPS N M+V V+G L + KF+Q F L P +G
Sbjct: 253 KLTSLPFQQCQHSITTVDSQPSGVNAAMLVFVSGNLQLAGEQHALKFSQMFHLIPTPQGS 432
Query: 122 FYVLNDVFR 130
+YVLND+FR
Sbjct: 433 YYVLNDIFR 459
>AI440855
Length = 361
Score = 57.0 bits (136), Expect = 1e-08
Identities = 31/83 (37%), Positives = 52/83 (62%)
Frame = +3
Query: 15 VGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSF 74
VG+ FV QYY IL + P+ VH+FY DSS M R + D ++ T +I + L +T+
Sbjct: 102 VGSYFVGQYYQILRQQPNLVHQFYSDSSSMIRVDGD-SVETAHDVLQIHSIVSLLNFTT- 275
Query: 75 RVEVLSADAQPSYNNGVMVVVTG 97
+E+ + ++ S++ GV+V+V+G
Sbjct: 276 -IEIKTINSLDSWDGGVLVMVSG 341
>TC204322 similar to UP|NTF2_ARATH (Q9C7F5) Nuclear transport factor 2
(NTF-2), partial (98%)
Length = 673
Score = 55.8 bits (133), Expect = 3e-08
Identities = 38/122 (31%), Positives = 61/122 (49%), Gaps = 3/122 (2%)
Frame = +1
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEY 71
P + AFVE YYS + + + Y + S+++ + + + I K+ SL +
Sbjct: 46 PDALAKAFVEHYYSTFDTNRNGLANLYQEGSMLTFEGQ-----KIQGASNIVAKLTSLPF 210
Query: 72 TSFRVEVLSADAQPS-YNNGVMVVVTGCL-TGTDNIKRKFAQSFFLAPQDKG-FYVLNDV 128
+ + D QPS N G++V V+G L + KF+Q F L P +G +YVLND+
Sbjct: 211 QQCHHSISTVDCQPSGVNAGMLVFVSGNLQLAGEQHTLKFSQMFHLIPTPQGSYYVLNDI 390
Query: 129 FR 130
FR
Sbjct: 391 FR 396
>TC204323 similar to UP|NTF2_ARATH (Q9C7F5) Nuclear transport factor 2
(NTF-2), partial (98%)
Length = 812
Score = 55.8 bits (133), Expect = 3e-08
Identities = 38/122 (31%), Positives = 61/122 (49%), Gaps = 3/122 (2%)
Frame = +2
Query: 12 PQMVGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEY 71
P + AFVE YYS + + + Y + S+++ + + + I K+ SL +
Sbjct: 83 PDALAKAFVEHYYSTFDTNRNGLANLYQEGSMLTFEGQ-----KIQGASSIVAKLTSLPF 247
Query: 72 TSFRVEVLSADAQPS-YNNGVMVVVTGCL-TGTDNIKRKFAQSFFLAPQDKG-FYVLNDV 128
+ + D QPS N G++V V+G L + KF+Q F L P +G +YVLND+
Sbjct: 248 QQCHHSISTVDCQPSGVNAGMLVFVSGNLQLAGEQHTLKFSQMFHLIPTPQGSYYVLNDI 427
Query: 129 FR 130
FR
Sbjct: 428 FR 433
>TC229960 similar to GB|AAO23575.1|27764908|BT003010 At5g60980/MSL3_100
{Arabidopsis thaliana;} , partial (14%)
Length = 763
Score = 50.8 bits (120), Expect = 1e-06
Identities = 27/31 (87%), Positives = 27/31 (87%), Gaps = 2/31 (6%)
Frame = +3
Query: 343 DGIQVRSNKG--SCFGFVEFESAASMQSALE 371
DGIQVRSNK SCFGFVEFESA SMQSALE
Sbjct: 36 DGIQVRSNKQQQSCFGFVEFESATSMQSALE 128
>AW760433 similar to PIR|H86398|H863 protein F17L21.10 [imported] -
Arabidopsis thaliana, partial (55%)
Length = 358
Score = 49.3 bits (116), Expect = 3e-06
Identities = 33/118 (27%), Positives = 59/118 (49%), Gaps = 3/118 (2%)
Frame = +2
Query: 15 VGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEEDGTMTTVTTTAEIDKKIQSLEYTSF 74
+ AFVE YYS + + + Y + S+++ + + +T+ I K+ SL + +
Sbjct: 5 LAKAFVEHYYSTFDTNRNGLANLYQEGSMLTFERQ-----KIQSTSNIVTKLTSLPFQXY 169
Query: 75 RVEVLSADAQPSY-NNGVMVVVTGCLTGTDNIKR-KFAQSFFLAP-QDKGFYVLNDVF 129
+ + + QPS+ N +++ V L TD KF Q F L P K +Y+LN++F
Sbjct: 170 HHSISTINYQPSHINTNILICVXRNLXLTDEQHTLKFNQIFHLIPTPQKSYYMLNNIF 343
>TC205914 weakly similar to UP|NAM8_YEAST (Q00539) NAM8 protein, partial (9%)
Length = 1513
Score = 45.4 bits (106), Expect = 4e-05
Identities = 33/99 (33%), Positives = 51/99 (51%), Gaps = 1/99 (1%)
Frame = +3
Query: 274 ASPAKPAVHPPRVHSVPAPEAPTPNMDIP-LEKNNENAGRAHAIFVANLPMSATVEQLDR 332
A+PA P P V+ VPA +P + P + NN IFV NL ++ + E+L +
Sbjct: 507 AAPAAPVPKP--VYPVPAYTSPVVQVQPPDYDVNNTT------IFVGNLDLNVSEEELKQ 662
Query: 333 AFKKFGPIKRDGIQVRSNKGSCFGFVEFESAASMQSALE 371
+FG I + V+ G FGFV+F + AS + A++
Sbjct: 663 NSLQFGEI----VSVKIQPGKGFGFVQFGTRASAEEAIQ 767
>TC217565 similar to UP|Q8VYJ4 (Q8VYJ4) AT3g25150/MJL12_9, partial (10%)
Length = 503
Score = 43.9 bits (102), Expect = 1e-04
Identities = 22/34 (64%), Positives = 26/34 (75%)
Frame = +2
Query: 345 IQVRSNKGSCFGFVEFESAASMQSALEVWSISIN 378
+QVRS KG FGFVEFE A+++QSALE I IN
Sbjct: 35 VQVRSQKGFSFGFVEFEVASAVQSALEASPILIN 136
>TC212608
Length = 421
Score = 43.1 bits (100), Expect = 2e-04
Identities = 20/36 (55%), Positives = 25/36 (68%)
Frame = +1
Query: 15 VGNAFVEQYYSILHRDPDQVHRFYHDSSVMSRPEED 50
VG+ FV QYY IL + P+ VH+FY DSS M R + D
Sbjct: 304 VGSYFVGQYYQILRQQPNLVHQFYSDSSSMIRVDGD 411
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.313 0.129 0.371
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,061,214
Number of Sequences: 63676
Number of extensions: 233979
Number of successful extensions: 2365
Number of sequences better than 10.0: 157
Number of HSP's better than 10.0 without gapping: 1832
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2285
length of query: 393
length of database: 12,639,632
effective HSP length: 99
effective length of query: 294
effective length of database: 6,335,708
effective search space: 1862698152
effective search space used: 1862698152
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146567.10


