

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
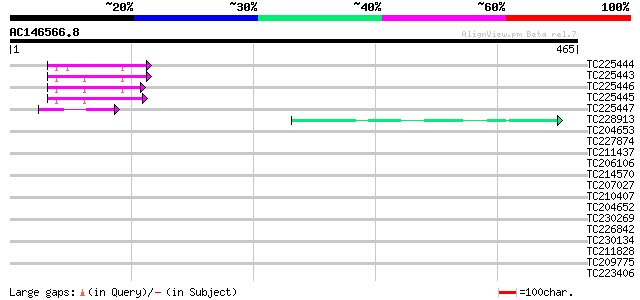
Query= AC146566.8 + phase: 0
(465 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC225444 homologue to UP|RL24_CICAR (O65743) 60S ribosomal prote... 55 7e-08
TC225443 homologue to UP|RL24_CICAR (O65743) 60S ribosomal prote... 52 4e-07
TC225446 similar to UP|RL24_HORVU (P50888) 60S ribosomal protein... 51 1e-06
TC225445 homologue to UP|RL24_CICAR (O65743) 60S ribosomal prote... 50 2e-06
TC225447 homologue to UP|RL24_CICAR (O65743) 60S ribosomal prote... 45 9e-05
TC228913 weakly similar to UP|Q8IP68 (Q8IP68) CG31813-PA, partia... 43 3e-04
TC204653 similar to UP|Q9AT20 (Q9AT20) Histone H1 (Fragment), pa... 41 0.001
TC227874 weakly similar to UP|O22670 (O22670) Ag13 protein precu... 40 0.002
TC211437 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%) 40 0.002
TC206106 weakly similar to UP|Q7RK82 (Q7RK82) Maebl (Fragment), ... 40 0.003
TC214570 homologue to PIR|T05766|T05766 peptidylprolyl isomerase... 39 0.004
TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%) 38 0.008
TC210407 similar to UP|O49656 (O49656) Predicted protein, partia... 38 0.008
TC204652 similar to UP|H1_PEA (P08283) Histone H1, partial (81%) 37 0.019
TC230269 weakly similar to UP|WSC2_YEAST (P53832) Cell wall inte... 37 0.019
TC226842 weakly similar to UP|Q84VG2 (Q84VG2) Senescence-associa... 37 0.019
TC230134 weakly similar to UP|NUCL_HUMAN (P19338) Nucleolin (Pro... 37 0.024
TC211828 similar to UP|RPIA_ARATH (Q9ZU38) Probable-ribose 5-pho... 36 0.032
TC209775 weakly similar to GB|AAM47330.1|21360429|AY113022 At1g7... 35 0.071
TC223406 weakly similar to UP|Q9FG99 (Q9FG99) Similarity to GTP-... 35 0.071
>TC225444 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protein L24,
complete
Length = 750
Score = 55.1 bits (131), Expect = 7e-08
Identities = 43/102 (42%), Positives = 59/102 (57%), Gaps = 17/102 (16%)
Frame = +2
Query: 32 KPSKRS-----RKAEKETVA--APKPKKGKTVKTES-STVSASEEVIQKKRNREPEVQDA 83
KPSK + RK K+ +A A K ++ T K S S V A+ EVIQKKR +PEV+DA
Sbjct: 182 KPSKLTWTAMYRKQHKKDIAQEAVKKRRRATKKPYSRSIVGATLEVIQKKRTEKPEVRDA 361
Query: 84 AREAALRE---------DATKREAALQAIRSKRAKGERSIQE 116
AREA LRE D K + A A +S++++G+ SI +
Sbjct: 362 AREAQLREIKERIKKTKDDKKAKKAEVAAKSQKSQGKGSISK 487
>TC225443 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protein L24,
complete
Length = 767
Score = 52.4 bits (124), Expect = 4e-07
Identities = 39/102 (38%), Positives = 56/102 (54%), Gaps = 17/102 (16%)
Frame = +2
Query: 32 KPSKRS-----RKAEKETVAAPKPKKGKTVKTES---STVSASEEVIQKKRNREPEVQDA 83
KPSK + RK K+ +A +K + + S V A+ EVIQKKR +PEV+DA
Sbjct: 227 KPSKLTWTAMYRKQHKKDIAQEAVRKRRRAAKKPYSRSIVGATLEVIQKKRAEKPEVRDA 406
Query: 84 AREAALRE---------DATKREAALQAIRSKRAKGERSIQE 116
AREAALRE D K + A A +S++A G+ ++ +
Sbjct: 407 AREAALREIKERIKKTKDEKKAKKAEVASKSQKAGGKGNVSK 532
>TC225446 similar to UP|RL24_HORVU (P50888) 60S ribosomal protein L24,
partial (96%)
Length = 865
Score = 51.2 bits (121), Expect = 1e-06
Identities = 39/97 (40%), Positives = 52/97 (53%), Gaps = 17/97 (17%)
Frame = +2
Query: 32 KPSKRS-----RKAEKETVAAPKPKKGKTVKTES---STVSASEEVIQKKRNREPEVQDA 83
KPSK + RK K+ +A KK + + S V A+ EVIQKKR +PEV+DA
Sbjct: 233 KPSKLTWTAMYRKQHKKDIAQEAVKKRRRAAKKPYSRSIVGATLEVIQKKRAEKPEVRDA 412
Query: 84 AREAALRE---------DATKREAALQAIRSKRAKGE 111
AREAALRE D K + A +S++A G+
Sbjct: 413 AREAALREIKERIKKTKDEKKAKKAEVTAKSQKAGGK 523
>TC225445 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protein L24,
complete
Length = 723
Score = 50.4 bits (119), Expect = 2e-06
Identities = 35/90 (38%), Positives = 50/90 (54%), Gaps = 8/90 (8%)
Frame = +2
Query: 32 KPSKRS-----RKAEKETVAAPKPKKGKTVKTES---STVSASEEVIQKKRNREPEVQDA 83
KPSK + RK K+ +A KK + + S V A+ EVIQKKR +PEV+DA
Sbjct: 194 KPSKLTWTAMYRKQHKKDIAQEAVKKRRRAAKKPYSRSIVGATLEVIQKKRAEKPEVRDA 373
Query: 84 AREAALREDATKREAALQAIRSKRAKGERS 113
REAALRE + + ++K+A+ +S
Sbjct: 374 NREAALREIKERNKKTKDEKKAKKAEFAKS 463
>TC225447 homologue to UP|RL24_CICAR (O65743) 60S ribosomal protein L24,
partial (61%)
Length = 687
Score = 44.7 bits (104), Expect = 9e-05
Identities = 27/67 (40%), Positives = 37/67 (54%)
Frame = +1
Query: 24 DLVEEATRKPSKRSRKAEKETVAAPKPKKGKTVKTESSTVSASEEVIQKKRNREPEVQDA 83
D+ +EA RK + ++K ++ V A+ EVIQKKR +PEV+DA
Sbjct: 7 DIAQEAVRKRRRAAKKPYSRSI-----------------VGATLEVIQKKRAEKPEVRDA 135
Query: 84 AREAALR 90
AREAALR
Sbjct: 136AREAALR 156
>TC228913 weakly similar to UP|Q8IP68 (Q8IP68) CG31813-PA, partial (19%)
Length = 692
Score = 43.1 bits (100), Expect = 3e-04
Identities = 51/223 (22%), Positives = 76/223 (33%), Gaps = 1/223 (0%)
Frame = +2
Query: 232 SSEPASEAPKSEATRPEAHPSGIP-SVPKASVNIQTPILPSSPSSSSSSSSTNSDDIPLS 290
SS P S P T + P+ P S P AS +QTP PSS +S+S ++ S
Sbjct: 86 SSSPPSSPPSPSTTPTSSAPANAPPSSPSASTRLQTPSGPSSDASTSPKPTSTS------ 247
Query: 291 QHIKQCLPNFKPSTSTFQDDYEQMQIHFSEQRIKICENLNLSADHFFQPPLVEPLNIQHP 350
+ P+ PST M S Q P V +
Sbjct: 248 ---SRAAPSKSPSTWPSASLATSMSSPASPQ------------------PRVPSASTSST 364
Query: 351 ETTPTPQKASEVALEAATSEDPQQHETSTLHNFEKHLGGEMQPTSTKASKTVPEKTVLEN 410
+ +P AS VA A+ + P P ST ++ T P T
Sbjct: 365 TSAASPASASSVANTASVTTAP-------------------SPPSTPSTTTTPPPTA--R 481
Query: 411 QPETSTVPEQSVPEHTVPEKVVSDLPQPNTEQQTENLNHSPTQ 453
+S+ P S P++ P P++ + + SP +
Sbjct: 482 STPSSSNPTLSTSPTATPKRTRVSSPTPSSSSTSRSSRPSPKE 610
>TC204653 similar to UP|Q9AT20 (Q9AT20) Histone H1 (Fragment), partial (68%)
Length = 1501
Score = 40.8 bits (94), Expect = 0.001
Identities = 36/151 (23%), Positives = 64/151 (41%), Gaps = 6/151 (3%)
Frame = +1
Query: 13 KTKSKALTKEDDLVEEATRKPSKRSRKAEKETVAAPKPKKGKTVKTESSTVSASEEVIQK 72
K K K K D ++ P+K A+ + KPK K ++ S + K
Sbjct: 559 KPKPKPAAKAKDTKATKSKAPAKPKAAAKPKAKVPAKPKPAPKPKAAAAAASKPKPAAAK 738
Query: 73 KRNREPEVQDAAREAALREDATKREAA---LQAIRSKRAKGERSIQERIREAAEQLRKEE 129
+ +P + + AA + A + AA + ++ R S ++ AA+ K+
Sbjct: 739 PKGTKPAAKPKPKPAAKTKAAAAKPAAKPKAKPAKASRTSTRTSPGKKAAPAAKPAAKKV 918
Query: 130 ADPSLK---KAQKPVSLESPCFRLTPEQEER 157
A P+ K K+ KP S++SP + T ++ R
Sbjct: 919 A-PAKKAPVKSVKPKSVKSPAKKATAKRGGR 1008
>TC227874 weakly similar to UP|O22670 (O22670) Ag13 protein precursor,
partial (21%)
Length = 894
Score = 40.4 bits (93), Expect = 0.002
Identities = 42/160 (26%), Positives = 75/160 (46%), Gaps = 7/160 (4%)
Frame = +1
Query: 8 VAKSRKTKSKAL-TKEDDLVEEATRKPSKRS---RKAEKETVAAPKPKKGKTVKT-ESST 62
VA TK +L E+ EE P+ + + ++E+ AP+P + + E T
Sbjct: 103 VALPEDTKEASLKVDEEKNTEEVVETPTPETVVTEQTKEESTEAPRPAASEDKPSVEVQT 282
Query: 63 VSASEEVIQKKRNREPEVQDAAREAALREDATKREAALQAIRSKRAKGERSIQERIREAA 122
+EEV+ + +EP V+ +EAA + TK A I++ A + +++E +EA
Sbjct: 283 KEVAEEVVSETVTQEPTVE--KQEAA---EGTKENADPAEIKTPEAAEDVNVKEE-KEAE 444
Query: 123 EQLRKEE--ADPSLKKAQKPVSLESPCFRLTPEQEERTRE 160
+ ++EE A+ S +K + S E + E E +E
Sbjct: 445 AEAKEEEAPAEKSEEKVETEGSAEKKEEEVIEEAEAEKKE 564
>TC211437 similar to UP|Q6VBJ4 (Q6VBJ4) Epa5p, partial (7%)
Length = 857
Score = 40.0 bits (92), Expect = 0.002
Identities = 25/59 (42%), Positives = 36/59 (60%), Gaps = 2/59 (3%)
Frame = -1
Query: 228 SGATSSEPASEAPKSEATRPEAH--PSGIPSVPKASVNIQTPILPSSPSSSSSSSSTNS 284
S ++SS P+S + S + P + PS +PS P AS + P PSSP +SSSS S++S
Sbjct: 587 SSSSSSSPSSSSSSSPLSSPSSSSSPSSLPS-PLASSSGSPPSSPSSPENSSSSPSSSS 414
Score = 28.1 bits (61), Expect = 8.7
Identities = 16/33 (48%), Positives = 19/33 (57%)
Frame = -1
Query: 271 SSPSSSSSSSSTNSDDIPLSQHIKQCLPNFKPS 303
SS SSSSSS S++S PLS P+ PS
Sbjct: 593 SSSSSSSSSPSSSSSSSPLSSPSSSSSPSSLPS 495
>TC206106 weakly similar to UP|Q7RK82 (Q7RK82) Maebl (Fragment), partial (7%)
Length = 1040
Score = 39.7 bits (91), Expect = 0.003
Identities = 40/156 (25%), Positives = 75/156 (47%), Gaps = 8/156 (5%)
Frame = +2
Query: 9 AKSRKTKSKALTKEDDLV---EEATRKPSKRSRKAEKETV-----AAPKPKKGKTVKTES 60
A+ + ++S + +E ++V EA RK ++ +E + APKP+ K KTE
Sbjct: 506 AEDKISQSVSFKEETNVVGDLPEAQRKALDELKRLVQEALNNHELTAPKPEPEKK-KTEL 682
Query: 61 STVSASEEVIQKKRNREPEVQDAAREAALREDATKREAALQAIRSKRAKGERSIQERIRE 120
+ EE + +++ +E EV + +EA + E+ K+EA + K E + E +E
Sbjct: 683 KALEKKEEEVSEEK-KEVEVTEEKKEAEVTEE--KKEAEV-----TEEKKEVEVAEEKKE 838
Query: 121 AAEQLRKEEADPSLKKAQKPVSLESPCFRLTPEQEE 156
K+E + + +K + V E +T E++E
Sbjct: 839 VEGTEEKKEVEVAEEKKETEVIKEKKEVEVTEEKKE 946
>TC214570 homologue to PIR|T05766|T05766 peptidylprolyl isomerase M4E13.20 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(13%)
Length = 944
Score = 39.3 bits (90), Expect = 0.004
Identities = 34/125 (27%), Positives = 59/125 (47%), Gaps = 11/125 (8%)
Frame = +2
Query: 24 DLVEEATRKPSKRSRKAEKETVAA-----PKPKKGKTVKTESSTVSASEEVIQKKRNR-E 77
D VEE ++ + +K ++ETV K KK K K S S+ EE +KK+ + +
Sbjct: 404 DEVEEVEKQEKVKVKKVDEETVEVEVEKKEKKKKKKKNKENSEAASSDEEKTEKKKKKHK 583
Query: 78 PEVQDAAREAALREDATK----REAALQAIRSKRAKGERSIQERIREAAEQLRK-EEADP 132
+V+D++ E E K +EAA K + S ++ + + +K ++AD
Sbjct: 584 DKVEDSSPELDKSEKKKKKKKDKEAAAATAEISNGKEDDSNADKSEKKKHKKKKNKDADE 763
Query: 133 SLKKA 137
+KA
Sbjct: 764 E*RKA 778
>TC207027 weakly similar to UP|Q9LW95 (Q9LW95) KED, partial (17%)
Length = 1005
Score = 38.1 bits (87), Expect = 0.008
Identities = 26/131 (19%), Positives = 64/131 (48%), Gaps = 4/131 (3%)
Frame = +3
Query: 10 KSRKTKSKALTKEDDLVEEATRKPSKRSRKAEKETVAAPKPKKGKTVKTESSTVSASEEV 69
K +K K K ++ E+ ++ K+ K +K+ +KGK + E + ++
Sbjct: 318 KEKKDKEKGDGDGEEKKEKKDKEKEKKKEKKDKDEETDTLKEKGKNDEGEDDEGNKKKKK 497
Query: 70 IQKKRNREPEVQDAAREAALREDA----TKREAALQAIRSKRAKGERSIQERIREAAEQL 125
+K++ ++ + + +E +ED+ + R+ ++ I+ + K ++ ++ +E E+
Sbjct: 498 DKKEKEKDHKKEKKDKEEGEKEDSKVEVSVRDIDIEEIKKEGEKEDKG-KDGGKEVKEKK 674
Query: 126 RKEEADPSLKK 136
+KE+ D KK
Sbjct: 675 KKEDKDKKEKK 707
>TC210407 similar to UP|O49656 (O49656) Predicted protein, partial (33%)
Length = 1010
Score = 38.1 bits (87), Expect = 0.008
Identities = 26/72 (36%), Positives = 35/72 (48%), Gaps = 9/72 (12%)
Frame = +3
Query: 244 ATRPEAHPSGIPSVPKASVNIQTPIL--------PSSPS-SSSSSSSTNSDDIPLSQHIK 294
AT P PSG + P + + T PSSPS SSSS+SS + +P + K
Sbjct: 219 ATSPSRRPSGSTTPPPSPITSSTATTFCSSSSSSPSSPSPSSSSNSSASPSSLPTRSNQK 398
Query: 295 QCLPNFKPSTST 306
P KPS++T
Sbjct: 399 SACPWPKPSSAT 434
>TC204652 similar to UP|H1_PEA (P08283) Histone H1, partial (81%)
Length = 1347
Score = 37.0 bits (84), Expect = 0.019
Identities = 40/166 (24%), Positives = 69/166 (41%), Gaps = 6/166 (3%)
Frame = +3
Query: 7 PVAKSRKTK---SKALTKEDDLVEEATRKPSKRSRKAEKETVAAPKPKKGKTVKTESSTV 63
P AK++ TK SKA K + + P+K A+ + AA KPK +
Sbjct: 570 PAAKAKDTKATKSKAPAKPKPAAKPKAKAPAKAKPAAKPKASAAAKPKPAAAKPKAAKPA 749
Query: 64 SASEEVIQKKRNREPEVQDAAREAALREDATKREAALQAIRSKRAKGERSIQERIREAAE 123
+ S+ + K A + A K +A + +++ R S ++ AA+
Sbjct: 750 AKSKPAAKTK-------------AVPAKPAAKPKA--KPVKASRTSTRTSPGKKAAPAAK 884
Query: 124 QLRKEEADPSLK---KAQKPVSLESPCFRLTPEQEERTREIAANGL 166
K+ A + K K+ KP S++SP + T ++ R E G+
Sbjct: 885 PAAKKVAVAAKKTPVKSVKPKSVKSPAKKATAKRGGRK*ERVVGGV 1022
Score = 32.3 bits (72), Expect = 0.46
Identities = 27/125 (21%), Positives = 51/125 (40%), Gaps = 1/125 (0%)
Frame = +3
Query: 29 ATRKPSKRSRKAEKETVAAPKPKKGKTVKTESSTVSASEEVIQKKRNREPEVQDAAREAA 88
++ P+K+ + A ++ PKPK K +T S + +P + A+ A
Sbjct: 501 SSSSPAKKKKPAAAKSKPKPKPKPAAKAKDTKATKS------KAPAKPKPAAKPKAKAPA 662
Query: 89 LREDATKREAALQAIRSKRAKGERSIQERIR-EAAEQLRKEEADPSLKKAQKPVSLESPC 147
+ A K +A+ A A ++ + + + A + + A P+ K KPV
Sbjct: 663 KAKPAAKPKASAAAKPKPAAAKPKAAKPAAKSKPAAKTKAVPAKPAAKPKAKPVKASRTS 842
Query: 148 FRLTP 152
R +P
Sbjct: 843 TRTSP 857
>TC230269 weakly similar to UP|WSC2_YEAST (P53832) Cell wall integrity and
stress response component 2 precursor, partial (6%)
Length = 696
Score = 37.0 bits (84), Expect = 0.019
Identities = 29/121 (23%), Positives = 58/121 (46%), Gaps = 17/121 (14%)
Frame = +2
Query: 24 DLVEEATRKPSKRSRKAEKETVAA-----PKPKKGKTVKTESSTVSASEEVIQKKRNR-- 76
D VEE + + +K ++ETV K KK K K S S+ EE +KK+N+
Sbjct: 50 DEVEELEKLEKVKVKKVDEETVEVEVEKKEKKKKKKKDKENSEVASSDEEKTEKKKNKNK 229
Query: 77 ----EPEVQDAAREAALREDATKREAA------LQAIRSKRAKGERSIQERIREAAEQLR 126
PE+ + ++ ++D AA + + +++ +++ +++ ++A E+ R
Sbjct: 230 IEDGSPELDKSEKKKKKKKDKAAAAAAEISNGKVDDSNADKSEKKKNKKKKNKDAKEE*R 409
Query: 127 K 127
K
Sbjct: 410 K 412
>TC226842 weakly similar to UP|Q84VG2 (Q84VG2) Senescence-associated
protein-like protein, partial (30%)
Length = 1116
Score = 37.0 bits (84), Expect = 0.019
Identities = 27/70 (38%), Positives = 36/70 (50%), Gaps = 6/70 (8%)
Frame = +2
Query: 226 HVSGATSSEPASEAP-KSEATRPEAHPSGIPSVPKASVNIQTPI-----LPSSPSSSSSS 279
H S TS+ P S P S A+ + P + SVP + + +P L SSPSSSS S
Sbjct: 101 HRSPTTSAWPTSGGPLSSSASSSSSFPWPVLSVPTGTNRVSSPSTSSVWLSSSPSSSSFS 280
Query: 280 SSTNSDDIPL 289
SS +S P+
Sbjct: 281 SSHSSSPDPM 310
>TC230134 weakly similar to UP|NUCL_HUMAN (P19338) Nucleolin (Protein C23),
partial (8%)
Length = 581
Score = 36.6 bits (83), Expect = 0.024
Identities = 25/78 (32%), Positives = 40/78 (51%)
Frame = -3
Query: 221 ALKAKHVSGATSSEPASEAPKSEATRPEAHPSGIPSVPKASVNIQTPILPSSPSSSSSSS 280
+L + S ++SS S P + + E+ S PS P+ S + + PSSPS +SSSS
Sbjct: 468 SLSSSSSSSSSSSSSLSPPPLASGSSSES-ASSPPSSPEKSSSSSSSSSPSSPSFTSSSS 292
Query: 281 STNSDDIPLSQHIKQCLP 298
S++S + + C P
Sbjct: 291 SSSSVSLASLFSSEVCFP 238
Score = 33.9 bits (76), Expect = 0.16
Identities = 22/63 (34%), Positives = 33/63 (51%), Gaps = 2/63 (3%)
Frame = -3
Query: 228 SGATSSEPASEAPKSEATRPEAHPSGIPSVPKASVNIQTPIL--PSSPSSSSSSSSTNSD 285
S ++SS P+S + S ++ + S + P AS + PSSP SSSSSS++S
Sbjct: 501 SSSSSSSPSSSSLSSSSSSSSSSSSSLSPPPLASGSSSESASSPPSSPEKSSSSSSSSSP 322
Query: 286 DIP 288
P
Sbjct: 321 SSP 313
>TC211828 similar to UP|RPIA_ARATH (Q9ZU38) Probable-ribose 5-phosphate
isomerase (Phosphoriboisomerase) , partial (47%)
Length = 729
Score = 36.2 bits (82), Expect = 0.032
Identities = 24/72 (33%), Positives = 37/72 (51%), Gaps = 1/72 (1%)
Frame = +2
Query: 235 PASEAPKSEATR-PEAHPSGIPSVPKASVNIQTPILPSSPSSSSSSSSTNSDDIPLSQHI 293
P S +PK+ + + P PS S +S + P LPS+PS++S+SSS + S+
Sbjct: 347 PPSSSPKTTSRKSPPTRPSSTSSPAWSSASAPAP-LPSTPSTASASSSAKENSKTSSESP 523
Query: 294 KQCLPNFKPSTS 305
P +PS S
Sbjct: 524 PPPKPTTRPSPS 559
Score = 34.7 bits (78), Expect = 0.093
Identities = 26/75 (34%), Positives = 38/75 (50%), Gaps = 3/75 (4%)
Frame = +2
Query: 232 SSEPASEAPKSEATRPEAHPSGIPSVPKASVNIQTPILPSSPSSSSS---SSSTNSDDIP 288
SS P + + KS TRP + S P+ AS P PS+ S+SSS +S T+S+ P
Sbjct: 353 SSSPKTTSRKSPPTRPSSTSS--PAWSSASAPAPLPSTPSTASASSSAKENSKTSSESPP 526
Query: 289 LSQHIKQCLPNFKPS 303
+ + P+ PS
Sbjct: 527 PPKPTTRPSPSASPS 571
>TC209775 weakly similar to GB|AAM47330.1|21360429|AY113022
At1g79870/F19K16_17 {Arabidopsis thaliana;} , partial
(24%)
Length = 502
Score = 35.0 bits (79), Expect = 0.071
Identities = 24/70 (34%), Positives = 37/70 (52%)
Frame = +3
Query: 235 PASEAPKSEATRPEAHPSGIPSVPKASVNIQTPILPSSPSSSSSSSSTNSDDIPLSQHIK 294
P+S+APKSE+T P PS+P + + P + S +++S+STN+D Q
Sbjct: 147 PSSQAPKSESTPP-------PSIPFPTSRLS----PPTASDTTTSTSTNAD---TEQSPS 284
Query: 295 QCLPNFKPST 304
P F P+T
Sbjct: 285 PTRPTF*PTT 314
>TC223406 weakly similar to UP|Q9FG99 (Q9FG99) Similarity to GTP-binding
regulatory protein and WD-repeat protein, partial (14%)
Length = 422
Score = 35.0 bits (79), Expect = 0.071
Identities = 29/101 (28%), Positives = 37/101 (35%), Gaps = 2/101 (1%)
Frame = +3
Query: 359 ASEVALEAATSEDPQQHETSTLHNFEKHLGGEMQPTSTKASKT--VPEKTVLENQPETST 416
AS L T QQH +LHN + P++T + T +P T +QP T
Sbjct: 60 ASLPLLTPTTINKGQQHRNPSLHN---AASSQSPPSTTTTTSTPRIPPSTTTASQPSRVT 230
Query: 417 VPEQSVPEHTVPEKVVSDLPQPNTEQQTENLNHSPTQNQTI 457
P S P P S P P T + T TI
Sbjct: 231 PPPTSPP---*PSPANSSTPAPPTAKSAPGTASQKTHQTTI 344
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.303 0.119 0.314
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,122,155
Number of Sequences: 63676
Number of extensions: 241695
Number of successful extensions: 3025
Number of sequences better than 10.0: 218
Number of HSP's better than 10.0 without gapping: 2206
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2637
length of query: 465
length of database: 12,639,632
effective HSP length: 101
effective length of query: 364
effective length of database: 6,208,356
effective search space: 2259841584
effective search space used: 2259841584
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 17 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 43 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC146566.8


