

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146566.2 + phase: 2 /partial
(358 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
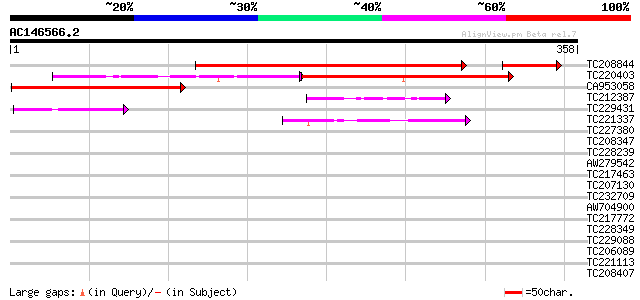
Score E
Sequences producing significant alignments: (bits) Value
TC208844 weakly similar to UP|Q76KB3 (Q76KB3) Makorin ring-zinc-... 323 e-100
TC220403 similar to UP|Q7X7W8 (Q7X7W8) Makorin ring-zinc-finger ... 208 2e-79
CA953058 196 1e-50
TC212387 weakly similar to UP|Q9ZW89 (Q9ZW89) F5A8.9 protein, pa... 45 7e-05
TC229431 44 9e-05
TC221337 similar to UP|Q9SD57 (Q9SD57) RNA-binding protein-like ... 44 1e-04
TC227380 weakly similar to UP|Q7EYM0 (Q7EYM0) Zinc finger protei... 40 0.002
TC208347 similar to UP|Q9XF63 (Q9XF63) RING-H2 zinc finger prote... 39 0.003
TC228239 weakly similar to UP|Q6IVC4 (Q6IVC4) Zinc finger protei... 39 0.004
AW279542 similar to GP|28558782|gb| RING/C3HC4/PHD zinc finger-l... 39 0.004
TC217463 similar to PIR|E86378|E86378 protein F21J9.10 [imported... 39 0.005
TC207130 similar to GB|AAH34435.1|21707915|BC034435 ZC3HDC3 prot... 39 0.005
TC232709 weakly similar to UP|Q9LZJ6 (Q9LZJ6) RING-H2 zinc finge... 39 0.005
AW704900 similar to GP|10178249|d DNA repair protein RAD5 protei... 38 0.006
TC217772 similar to GB|AAM51588.1|21436021|AY116954 AT5g19430/F7... 38 0.008
TC228349 37 0.011
TC229088 similar to UP|Q40221 (Q40221) Protein containing C-term... 37 0.014
TC206089 UP|Q39902 (Q39902) C-terminal zinc-finger (Fragment), c... 37 0.014
TC221113 weakly similar to UP|Q9FWS3 (Q9FWS3) F1B16.12 protein, ... 37 0.014
TC208407 similar to UP|Q84KA9 (Q84KA9) RING/C3HC4/PHD zinc finge... 37 0.014
>TC208844 weakly similar to UP|Q76KB3 (Q76KB3) Makorin ring-zinc-finger
protein (Fragment), partial (34%)
Length = 1264
Score = 323 bits (827), Expect(2) = e-100
Identities = 147/171 (85%), Positives = 159/171 (92%)
Frame = +1
Query: 118 LREDDVVQSVITSPSELPICSFAAAGNCPRGEQCPHVHGDLCPSCGRQCLHPFRPEEREE 177
L ED+V Q++ TSPSEL ICSFAAAGNCPRGE+CPH+HGDLCP+CG+ CLHPFRPEEREE
Sbjct: 7 LGEDNVGQTIGTSPSELSICSFAAAGNCPRGEKCPHIHGDLCPTCGKHCLHPFRPEEREE 186
Query: 178 HMMSCRNKQKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWR 237
HM SC NK KHL+ALKRSQEIECSVCLE VLSKPTAAE KFGLLSECDHPFC+SCIRNWR
Sbjct: 187 HMKSCENKHKHLDALKRSQEIECSVCLELVLSKPTAAECKFGLLSECDHPFCISCIRNWR 366
Query: 238 SSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSEEKMEIIDTYKAKL 288
SSNPTLGMDVNSTLRACPICRKLSYFV+PSVIWY+T+EEK EIID YKAKL
Sbjct: 367 SSNPTLGMDVNSTLRACPICRKLSYFVIPSVIWYSTTEEKQEIIDNYKAKL 519
Score = 62.4 bits (150), Expect(2) = e-100
Identities = 29/37 (78%), Positives = 32/37 (86%)
Frame = +3
Query: 312 KHAYRDGRLEEVALRHLGAADGDTIIAKDIRFLSTLS 348
+HAYRDGRLE+V LRHLGAADGDT+IAKDIR LS
Sbjct: 519 QHAYRDGRLEQVVLRHLGAADGDTVIAKDIRLSDFLS 629
>TC220403 similar to UP|Q7X7W8 (Q7X7W8) Makorin ring-zinc-finger protein
(Makorin RING finger protein), partial (62%)
Length = 1109
Score = 208 bits (529), Expect(2) = 2e-79
Identities = 94/139 (67%), Positives = 110/139 (78%), Gaps = 5/139 (3%)
Frame = +1
Query: 185 KQKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLG 244
K K+L+ALK SQE++C+VCLERVLSKP A+ KFGLL ECDH FC+SCIRNWR+S PT G
Sbjct: 451 KGKYLQALKDSQEVQCNVCLERVLSKPKPADCKFGLLPECDHAFCLSCIRNWRNSAPTSG 630
Query: 245 MDV-----NSTLRACPICRKLSYFVVPSVIWYATSEEKMEIIDTYKAKLKSIDCKHFDFG 299
MD+ +T+R CP+C KLSYFV+PS IWY+T EEK EIID YKA K IDCKHF+FG
Sbjct: 631 MDIINAGTANTVRTCPVCLKLSYFVIPSGIWYSTKEEKQEIIDNYKANCKLIDCKHFNFG 810
Query: 300 EGNCPFGTSCFYKHAYRDG 318
GNCPFG SCFYKH + G
Sbjct: 811 NGNCPFGASCFYKHTVKPG 867
Score = 105 bits (263), Expect(2) = 2e-79
Identities = 66/161 (40%), Positives = 91/161 (55%), Gaps = 2/161 (1%)
Frame = +2
Query: 28 NNICTFYQKGVCAYGSRCRYDHVKASRAQSSTPSSSITEHQPLVSESAVLGNTRVTSNGV 87
++ICT+Y+KG CAYGSRCRY HVKAS+A SS H P V + V TS+ V
Sbjct: 17 DDICTYYKKGSCAYGSRCRYKHVKASQASSSANG----RHSP-VLDPVVNHTITGTSSWV 181
Query: 88 ATAAEFSLFSTPFVLPSEQAWNQESAQLDFLREDDVVQSVITS--PSELPICSFAAAGNC 145
A + S ++A + + L L E+DV QS +S PSE C+FAAA NC
Sbjct: 182 PKAVKSS-------SSDKRARSSQQKNLPSL-ENDVGQSSTSSVIPSEHLFCAFAAA-NC 334
Query: 146 PRGEQCPHVHGDLCPSCGRQCLHPFRPEEREEHMMSCRNKQ 186
P ++C +HG+ C C + CLHP +E+E H+ +C K+
Sbjct: 335 PLEDKCSRIHGNQCLYCRKFCLHPTDRKEKENHLRTCEKKE 457
>CA953058
Length = 421
Score = 196 bits (498), Expect = 1e-50
Identities = 88/110 (80%), Positives = 96/110 (87%)
Frame = +1
Query: 2 LCKFFAHGACLKGEHCEFSHDWKAPPNNICTFYQKGVCAYGSRCRYDHVKASRAQSSTPS 61
LCKFFAHGACLKGEHCEFSHDWK PPNN+CTFYQKGVCAYGSRCRYDHVKASR QSS PS
Sbjct: 82 LCKFFAHGACLKGEHCEFSHDWKDPPNNVCTFYQKGVCAYGSRCRYDHVKASREQSSAPS 261
Query: 62 SSITEHQPLVSESAVLGNTRVTSNGVATAAEFSLFSTPFVLPSEQAWNQE 111
SS+ E Q LVS++ GNTR SNGVAT AEFSL S+P++LPSE AWN+E
Sbjct: 262 SSVIEPQTLVSDTVAFGNTRAASNGVATGAEFSLSSSPYLLPSEPAWNRE 411
>TC212387 weakly similar to UP|Q9ZW89 (Q9ZW89) F5A8.9 protein, partial (16%)
Length = 1107
Score = 44.7 bits (104), Expect = 7e-05
Identities = 30/91 (32%), Positives = 46/91 (49%)
Frame = +1
Query: 188 HLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDV 247
++ L S E+ C +CL S G+L C H FC CI+NW ++ T M
Sbjct: 511 NVAGLPTSTEMSCVICLTEFSSTR-------GILP-CGHRFCFPCIQNW--ADHTTSMRK 660
Query: 248 NSTLRACPICRKLSYFVVPSVIWYATSEEKM 278
ST CP+C K S+ ++ V AT+++K+
Sbjct: 661 TST---CPLC-KASFMMIKKVEHAATADQKI 741
>TC229431
Length = 525
Score = 44.3 bits (103), Expect = 9e-05
Identities = 26/75 (34%), Positives = 36/75 (47%), Gaps = 2/75 (2%)
Frame = +3
Query: 3 CKFFAHGACLKGEHCEFSHDWKAPPNNICT-FYQKGVCAYGSRCRYDHVKASRAQSSTPS 61
C FA +C+KG+ C F H P CT F KG C G C + H +++ T S
Sbjct: 156 CTHFARQSCMKGDDCPFDHQLSKYP---CTNFVSKGSCYKGDACLFSHQVSTKEDIPTAS 326
Query: 62 SSI-TEHQPLVSESA 75
+ E PL+S +A
Sbjct: 327 NMCRPELSPLLSGNA 371
Score = 40.0 bits (92), Expect = 0.002
Identities = 18/51 (35%), Positives = 28/51 (54%), Gaps = 4/51 (7%)
Frame = +3
Query: 3 CKFFAHGACLKGEHCEFSHD----WKAPPNNICTFYQKGVCAYGSRCRYDH 49
C+ + +G C +G+ C+FSHD K+ P CT + + C G C +DH
Sbjct: 69 CRHYLNGRCHEGDKCQFSHDVVPLTKS*P---CTHFARQSCMKGDDCPFDH 212
>TC221337 similar to UP|Q9SD57 (Q9SD57) RNA-binding protein-like protein,
partial (77%)
Length = 1151
Score = 43.5 bits (101), Expect = 1e-04
Identities = 29/126 (23%), Positives = 55/126 (43%), Gaps = 7/126 (5%)
Frame = +1
Query: 173 EEREEHMMSCRNKQK------HLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDH 226
+ +++H+ + + K K +L + +E+EC +CLE + SK +L C+H
Sbjct: 655 DRKQKHLCATKYKLKDLTSKGNLSEIDMERELECGICLE-INSKV--------VLPNCNH 807
Query: 227 PFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIW-YATSEEKMEIIDTYK 285
C+ C +W + + ++CP CR V +W Y +S E ++ K
Sbjct: 808 SMCMKCYEDWHARS-----------QSCPFCRDSLKRVNTDDLWIYISSSEINDLASINK 954
Query: 286 AKLKSI 291
K +
Sbjct: 955 ENFKRL 972
>TC227380 weakly similar to UP|Q7EYM0 (Q7EYM0) Zinc finger protein
family-like, partial (70%)
Length = 770
Score = 39.7 bits (91), Expect = 0.002
Identities = 27/117 (23%), Positives = 50/117 (42%)
Frame = +2
Query: 166 CLHPFRPEEREEHMMSCRNKQKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECD 225
C+ +R + EEH + + +E EC +C+E + SK +L +C+
Sbjct: 167 CMERYRKRDDEEH--------RQPSDIDIEREEECGICME-MNSKI--------VLPDCN 295
Query: 226 HPFCVSCIRNWRSSNPTLGMDVNSTLRACPICRKLSYFVVPSVIWYATSEEKMEIID 282
H C++C WR+ + ++CP CR V +W T + +++D
Sbjct: 296 HVMCLTCYHEWRTRS-----------QSCPFCRDSLKRVNSGDLWVFT--DNRDVVD 427
>TC208347 similar to UP|Q9XF63 (Q9XF63) RING-H2 zinc finger protein ATL3
(At1g72310) (RING-H2 zinc finger protein (ATL3);
86824-85850) (RING-H2 zinc finger protein ATL3;
90350-91324), partial (24%)
Length = 1229
Score = 39.3 bits (90), Expect = 0.003
Identities = 22/61 (36%), Positives = 30/61 (49%)
Frame = +3
Query: 198 IECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPIC 257
+EC+VCL V+ A LL +C+H F V+CI W S+ T CP+C
Sbjct: 489 LECAVCLSEVVEGEKAR-----LLPKCNHGFHVACIDMWFQSHST-----------CPLC 620
Query: 258 R 258
R
Sbjct: 621 R 623
>TC228239 weakly similar to UP|Q6IVC4 (Q6IVC4) Zinc finger protein
(Zinc-finger transcription factor), partial (32%)
Length = 1244
Score = 38.9 bits (89), Expect = 0.004
Identities = 30/88 (34%), Positives = 39/88 (44%), Gaps = 14/88 (15%)
Frame = +2
Query: 25 APPNNICTFY-QKGVCAYGSRCRYDHVKASRAQSSTPS-------------SSITEHQPL 70
APP CT Y Q+GVC +GS C++DH S + S + S SSI P
Sbjct: 74 APP---CTHYTQRGVCKFGSACKFDHPMGSLSYSPSASSLADMPVAPYPVGSSIGTLAPS 244
Query: 71 VSESAVLGNTRVTSNGVATAAEFSLFST 98
S S + SN + A+ S ST
Sbjct: 245 SSSSELRPELGAGSNKESVASRMSSMST 328
>AW279542 similar to GP|28558782|gb| RING/C3HC4/PHD zinc finger-like protein
{Cucumis melo}, partial (12%)
Length = 434
Score = 38.9 bits (89), Expect = 0.004
Identities = 22/66 (33%), Positives = 32/66 (48%)
Frame = +3
Query: 193 KRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLR 252
K +Q +EC+VCL K + LL +C+H F CI +W +S+ T
Sbjct: 27 KANQTLECAVCLTDFTHKDS-----LRLLPKCNHVFHPHCIDSWLTSHVT---------- 161
Query: 253 ACPICR 258
CP+CR
Sbjct: 162 -CPVCR 176
>TC217463 similar to PIR|E86378|E86378 protein F21J9.10 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(46%)
Length = 687
Score = 38.5 bits (88), Expect = 0.005
Identities = 23/87 (26%), Positives = 34/87 (38%)
Frame = +3
Query: 196 QEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACP 255
+E EC +CLE +L C H C+ C R W N+ +CP
Sbjct: 237 REDECGICLEPCTKM---------VLPNCCHAMCIKCYRKW-----------NTRSESCP 356
Query: 256 ICRKLSYFVVPSVIWYATSEEKMEIID 282
CR V +W T +E +++D
Sbjct: 357 FCRGSLRRVNSEDLWVLTCDE--DVVD 431
>TC207130 similar to GB|AAH34435.1|21707915|BC034435 ZC3HDC3 protein {Homo
sapiens;} , partial (6%)
Length = 1005
Score = 38.5 bits (88), Expect = 0.005
Identities = 17/47 (36%), Positives = 23/47 (48%)
Frame = +1
Query: 3 CKFFAHGACLKGEHCEFSHDWKAPPNNICTFYQKGVCAYGSRCRYDH 49
C +F G C +C + H P +IC + KG CA G+ CR H
Sbjct: 307 CSYFLQGLC-SNRNCPYRHVNVNPKASICEGFLKGYCADGNECRKKH 444
>TC232709 weakly similar to UP|Q9LZJ6 (Q9LZJ6) RING-H2 zinc finger protein
ATL5 (At3g62690), partial (19%)
Length = 987
Score = 38.5 bits (88), Expect = 0.005
Identities = 23/73 (31%), Positives = 33/73 (44%), Gaps = 1/73 (1%)
Frame = +3
Query: 187 KHLEALKRSQEIE-CSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGM 245
K + K+ +E+ CSVCL ++ + +L C H F V C+ W +SN T
Sbjct: 342 KQTDQFKQGEEVV*CSVCLGTIVEDTISR-----VLPNCKHIFHVDCVDKWFNSNTT--- 497
Query: 246 DVNSTLRACPICR 258
CPICR
Sbjct: 498 --------CPICR 512
>AW704900 similar to GP|10178249|d DNA repair protein RAD5 protein
{Arabidopsis thaliana}, partial (13%)
Length = 428
Score = 38.1 bits (87), Expect = 0.006
Identities = 22/75 (29%), Positives = 36/75 (47%), Gaps = 1/75 (1%)
Frame = +1
Query: 186 QKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSC-IRNWRSSNPTLG 244
Q+ +E L++ ++ EC +CLE + +L+ C H C C + +WR
Sbjct: 199 QEVVEELRKGEQGECPICLEVF---------EDAVLTPCAHRLCRECLLSSWR------- 330
Query: 245 MDVNSTLRACPICRK 259
N+T CP+CRK
Sbjct: 331 ---NATSGLCPVCRK 366
>TC217772 similar to GB|AAM51588.1|21436021|AY116954 AT5g19430/F7K24_180
{Arabidopsis thaliana;} , partial (61%)
Length = 1136
Score = 37.7 bits (86), Expect = 0.008
Identities = 16/59 (27%), Positives = 31/59 (52%)
Frame = +2
Query: 200 CSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICR 258
C++CL++++ + TA L+ C+H +CV+CI +W + + CP C+
Sbjct: 311 CAICLDKIVLQETA------LVKGCEHAYCVTCILHWATYREKV---------TCPQCK 442
>TC228349
Length = 1112
Score = 37.4 bits (85), Expect = 0.011
Identities = 20/60 (33%), Positives = 28/60 (46%)
Frame = +2
Query: 199 ECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPICR 258
EC+VCL+ + S+ A L+ C+H F V C W S +P CP+CR
Sbjct: 533 ECAVCLDEIESEQPAR-----LVPGCNHGFHVHCADTWLSKHP-----------LCPVCR 664
>TC229088 similar to UP|Q40221 (Q40221) Protein containing C-terminal
RING-finger, partial (63%)
Length = 1240
Score = 37.0 bits (84), Expect = 0.014
Identities = 21/83 (25%), Positives = 33/83 (39%)
Frame = +3
Query: 176 EEHMMSCRNKQKHLEALKRSQEIECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRN 235
E+ + C + + + + E C +CLE + G L C H + VSCI+
Sbjct: 831 EDLLSKCLTETIYCSSEQSEDEGNCVICLEEYKNMDDV-----GTLQTCGHDYHVSCIKK 995
Query: 236 WRSSNPTLGMDVNSTLRACPICR 258
W S + CPIC+
Sbjct: 996 WLSMK-----------KLCPICK 1031
>TC206089 UP|Q39902 (Q39902) C-terminal zinc-finger (Fragment), complete
Length = 1401
Score = 37.0 bits (84), Expect = 0.014
Identities = 22/67 (32%), Positives = 30/67 (43%), Gaps = 1/67 (1%)
Frame = +3
Query: 193 KRSQEIE-CSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTL 251
++SQE E C++CLE + G L C H + V CIR W S
Sbjct: 1014 EQSQEEEACAICLEEYKNMDDV-----GTLKACGHDYHVGCIRKWLSMK----------- 1145
Query: 252 RACPICR 258
+ CPIC+
Sbjct: 1146 KVCPICK 1166
>TC221113 weakly similar to UP|Q9FWS3 (Q9FWS3) F1B16.12 protein, partial
(12%)
Length = 548
Score = 37.0 bits (84), Expect = 0.014
Identities = 14/26 (53%), Positives = 18/26 (68%)
Frame = +3
Query: 29 NICTFYQKGVCAYGSRCRYDHVKASR 54
+IC +Q+G C YG RCRY HV+ R
Sbjct: 156 DICRNFQRGSCQYGERCRYLHVQQQR 233
>TC208407 similar to UP|Q84KA9 (Q84KA9) RING/C3HC4/PHD zinc finger-like
protein, partial (36%)
Length = 644
Score = 37.0 bits (84), Expect = 0.014
Identities = 20/61 (32%), Positives = 27/61 (43%)
Frame = +2
Query: 198 IECSVCLERVLSKPTAAERKFGLLSECDHPFCVSCIRNWRSSNPTLGMDVNSTLRACPIC 257
+EC+VCL T L+ +CDH F CI W +S+ T CP+C
Sbjct: 305 LECAVCLNEFEDTETLR-----LIPKCDHVFHPECIDEWLASHTT-----------CPVC 436
Query: 258 R 258
R
Sbjct: 437 R 439
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.322 0.135 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 21,793,538
Number of Sequences: 63676
Number of extensions: 390043
Number of successful extensions: 2234
Number of sequences better than 10.0: 167
Number of HSP's better than 10.0 without gapping: 2096
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2212
length of query: 358
length of database: 12,639,632
effective HSP length: 98
effective length of query: 260
effective length of database: 6,399,384
effective search space: 1663839840
effective search space used: 1663839840
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146566.2


