

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146565.8 - phase: 0 /pseudo
(1145 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
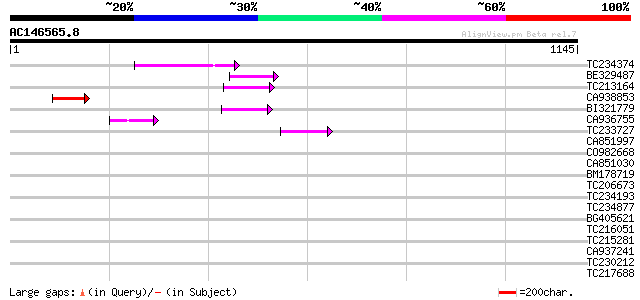
Score E
Sequences producing significant alignments: (bits) Value
TC234374 112 7e-25
BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase... 73 6e-13
TC213164 71 2e-12
CA938853 similar to GP|1684847|gb|A pinin {Homo sapiens}, partia... 68 2e-11
BI321779 53 9e-07
CA936755 48 3e-05
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 46 1e-04
CA851997 40 0.008
CO982668 37 0.065
CA851030 36 0.084
BM178719 36 0.084
TC206673 35 0.19
TC234193 33 0.93
TC234877 32 1.6
BG405621 31 2.7
TC216051 weakly similar to UP|Q8GTJ9 (Q8GTJ9) Cysteine proteinas... 31 3.5
TC215281 31 3.5
CA937241 similar to GP|6840918|gb|A hematopoietic antigen CD38 {... 30 7.9
TC230212 similar to GB|AAP78934.1|32306501|BT009666 At5g12120 {A... 30 7.9
TC217688 similar to UP|Q6K709 (Q6K709) DNA-binding protein-like,... 30 7.9
>TC234374
Length = 787
Score = 112 bits (281), Expect = 7e-25
Identities = 74/215 (34%), Positives = 117/215 (54%), Gaps = 3/215 (1%)
Frame = +2
Query: 253 HVDCRRLVLENWAKNVRGVGMVRLQGKLRILKTVFKQWNRSVFGDVDRQVRLATDEVSRI 312
+VDC R V + W+ +V G +V LQ KL++ K K W ++VFG+V + V AT+EV I
Sbjct: 5 NVDCVRFVXKVWSCHVLGCPIVVLQQKLKM*KC*LKTWIKNVFGNVHQLVL*ATNEVDSI 184
Query: 313 QQIIDSVGFSDDLYRQDLEAQLLLTNALNVQDQFWKEKSRHQSFIHGDRNTAYF---HRV 369
QQ I +VG+SD L+ Q++ AQ L L Q+ W+EKSR G++NT++F H +
Sbjct: 185 QQQIVNVGYSDALFNQEVLAQKELQTVLRFQECLWREKSRI*GHAQGEQNTSFFIK*HSI 364
Query: 370 ARIKASSKNISLLYDGATVLSDAGDIAAHILNYFQGIFDVVNNCIPNDLVARSIHALVTE 429
+ D ++ + F +V+ + + R+I +LV+
Sbjct: 365 ILYLRVYLSYVKGIDY*SLKRSSKQRCGISFPIFMRAATLVS--LTAL*IYRTIPSLVSG 538
Query: 430 DENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGG 464
+++ L +PL +E+KS VFD+NG+G PG DG G
Sbjct: 539 EDDAFLSNIPLNEEVKSLVFDMNGEGVPGTDGCNG 643
Score = 40.8 bits (94), Expect = 0.003
Identities = 39/143 (27%), Positives = 60/143 (41%), Gaps = 3/143 (2%)
Frame = +1
Query: 365 YFHRVARIKASSKNISLLYDGATVLSDAGDIAAHILNYFQGIFDVVNNCIPNDLVARSIH 424
+FH++A SK +S L G +L + A + ++F I+ ++CIPN V S H
Sbjct: 340 FFHKIALHHFVSKGLS*LRKGN*LLESQEEFEAEVRDFFPNIYAGSDSCIPNSFVDISYH 519
Query: 425 A-LVTEDENNMLIRLPLRDEIKSAVFDLNGD--GAPGPDGFGGHFYQSFWDIVGSDVVLS 481
+ L I+ P + +F + + G F+Q + D VG V
Sbjct: 520 SKLGIR*R*CFFIQHPFK*G--G*IFSV*HEW*GCSRDRWLQRIFFQHY*DAVGDTVYQL 693
Query: 482 VQDFFLQGSFADNINSNLLVLIP 504
V F N+NSN VL P
Sbjct: 694 VLQLFKSRWILPNLNSNTAVLFP 762
>BE329487 weakly similar to PIR|T14619|T146 reverse transcriptase - beet
retrotransposon (fragment), partial (15%)
Length = 376
Score = 73.2 bits (178), Expect = 6e-13
Identities = 36/99 (36%), Positives = 52/99 (52%)
Frame = -2
Query: 444 IKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNINSNLLVLI 503
IKSAV+ +PGPDG F + FW+ + D++ + +F G F N++ + LI
Sbjct: 375 IKSAVWQCGSVKSPGPDGLNFKFIKHFWERLKPDIIRFLNEFHANGIFPKGGNASFIALI 196
Query: 504 PKIPGAQAMGDFRPIALANFQFKIVTKILADRLASIAMC 542
PK+ QA+ DFRPI+L +KIV KI C
Sbjct: 195 PKVKHPQALNDFRPISLIGCVYKIVAKIXXQNXIEKXCC 79
>TC213164
Length = 446
Score = 71.2 bits (173), Expect = 2e-12
Identities = 32/104 (30%), Positives = 62/104 (58%)
Frame = +3
Query: 432 NNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSF 491
+++L+ + E+++ ++D +PGPDG F + FWDI+ +D++ + +F+ G F
Sbjct: 9 SDLLVARFEKKEVRATMWDRGNAKSPGPDGLNFKFIKEFWDILKTDLLRFLDEFYANGVF 188
Query: 492 ADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTKILADR 535
N+ L LIPK+ ++ +++PI+L +KIV K+L+ R
Sbjct: 189 LKRSNAFFLALIPKVHDP*SLNEYKPISLIGCIYKIVAKLLSGR 320
>CA938853 similar to GP|1684847|gb|A pinin {Homo sapiens}, partial (2%)
Length = 422
Score = 68.2 bits (165), Expect = 2e-11
Identities = 34/74 (45%), Positives = 49/74 (65%)
Frame = -3
Query: 87 SDQCIALEVTWQHSSVFLAAIYANTSYLFRRLLWADLTHLQGCFLGPWLFIGDFNAVMGA 146
S Q +A +++ + F AA+YA+T+Y+ RR LW++LT LQ LG W FIGDFN+ +GA
Sbjct: 225 SHQ*VAFKLSLGSKTSFFAAVYASTAYVKRRDLWSELTDLQQKHLGSWPFIGDFNSNVGA 46
Query: 147 HEKRGRWPPTAASC 160
EKRG + +C
Sbjct: 45 QEKRGGKMSCSIAC 4
>BI321779
Length = 421
Score = 52.8 bits (125), Expect = 9e-07
Identities = 29/103 (28%), Positives = 51/103 (49%)
Frame = +2
Query: 428 TEDENNMLIRLPLRDEIKSAVFDLNGDGAPGPDGFGGHFYQSFWDIVGSDVVLSVQDFFL 487
+ ++N MLI+ +EIK + D +PGPDG+ + FW+++ +V+ ++F
Sbjct: 62 SNEDNAMLIQDFEVEEIKKVI*DCESSKSPGPDGYSLLLIKIFWEVIKEEVINFFKEFHK 241
Query: 488 QGSFADNINSNLLVLIPKIPGAQAMGDFRPIALANFQFKIVTK 530
N + + LI K +G + PI+L +KIV K
Sbjct: 242 FDFLPRGTNRSFIALIAKCDNPLDIGQYCPISLVGCLYKIV*K 370
>CA936755
Length = 424
Score = 47.8 bits (112), Expect = 3e-05
Identities = 31/99 (31%), Positives = 51/99 (51%), Gaps = 1/99 (1%)
Frame = -2
Query: 202 ICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADVASVKNAISFKFYKAWTSHVDCRRLVL 261
+ +E+WL + +S TL R SDH P+++ F+ W +RLV
Sbjct: 291 LVSEQWLLSWPDSSQYTLPRDFSDHCPIIMQTKKVDW-GPKPFRVVDWWLHQKGYQRLVR 115
Query: 262 ENW-AKNVRGVGMVRLQGKLRILKTVFKQWNRSVFGDVD 299
E W A+ G G + L+ KLR+LK KQW++ +G+++
Sbjct: 114 ETWSAEQQPGWGGILLKNKLRMLKLSIKQWSKE-YGNIN 1
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 45.8 bits (107), Expect = 1e-04
Identities = 34/108 (31%), Positives = 48/108 (43%), Gaps = 3/108 (2%)
Frame = -2
Query: 547 FYSWS*YCGLHYLGV*GD*CVR*KTVWW*YCVKGGYSQGF*YAGLEFLNCGVASVWFFFC 606
F+ + Y GLH VR K++WW +K + +GF*Y G + +W
Sbjct: 853 FFIYLNYQGLHLCHFRSYKYVRQKSLWWELGLKDRHCEGF*YPGQGLPHLCPK*IWV*PH 674
Query: 607 FLQLDFGNFTLGQTFYFGEWQGAGVLLLCPWCSSRG---PFISSPFLS 651
L LD NF + + Y W+ G LLL C +R P SP +S
Sbjct: 673 LL*LDPSNFIICKALYLC*WRTCGFLLLSKGCKTRRSPIPLFVSPKMS 530
>CA851997
Length = 636
Score = 39.7 bits (91), Expect = 0.008
Identities = 25/83 (30%), Positives = 42/83 (50%), Gaps = 2/83 (2%)
Frame = +1
Query: 459 PDGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPI 518
PDGF FY FW + G DV + + +G+F ++N + +I +I +
Sbjct: 316 PDGFNLGFYHRFWGMCGEDVFQACCMWLAEGAFPSSVNDTTIAIILEI**S*RYERS*TN 495
Query: 519 ALANFQFKI--VTKILADRLASI 539
L N FKI ++++LA RL ++
Sbjct: 496 LLCNGGFKILFLSEVLAKRLKNV 564
>CO982668
Length = 870
Score = 36.6 bits (83), Expect = 0.065
Identities = 21/75 (28%), Positives = 34/75 (45%), Gaps = 2/75 (2%)
Frame = -2
Query: 193 YVALRLDRSICNEEWLAFYRVTSCCTLIRHQSDHHPLLLSADVASVKNA--ISFKFYKAW 250
+ +RLDR + N+EW + L + D P ++ DV N ++F AW
Sbjct: 275 FFQVRLDRVLMNQEWRIESPIVKVTHLFHCK*DQ*PHHINLDVG*PMNRKRCPYRFQNAW 96
Query: 251 TSHVDCRRLVLENWA 265
SH + L+ NW+
Sbjct: 95 LSHDEFDSLIRHNWS 51
>CA851030
Length = 358
Score = 36.2 bits (82), Expect = 0.084
Identities = 15/49 (30%), Positives = 30/49 (60%)
Frame = +1
Query: 336 LTNALNVQDQFWKEKSRHQSFIHGDRNTAYFHRVARIKASSKNISLLYD 384
L + L ++++W+++SR F D+NT +FHR + +S +I+ + D
Sbjct: 151 LDSLLKQEEEYWQQRSRAIWFKERDKNTEFFHRKTLQRKASNSITKIQD 297
>BM178719
Length = 378
Score = 36.2 bits (82), Expect = 0.084
Identities = 16/29 (55%), Positives = 19/29 (65%)
Frame = +2
Query: 132 GPWLFIGDFNAVMGAHEKRGRWPPTAASC 160
GP F+GD NA++G HEKRG P SC
Sbjct: 104 GP*CFLGDNNAILGEHEKRGGKLPLYISC 190
Score = 30.8 bits (68), Expect(2) = 0.28
Identities = 15/36 (41%), Positives = 20/36 (54%)
Frame = +2
Query: 215 SCCTLIRHQSDHHPLLLSADVASVKNAISFKFYKAW 250
SC + + SDHHPLL + S+ +A F F K W
Sbjct: 185 SCFFMSQT*SDHHPLLTNFQHCSMLHASHFFFIKMW 292
Score = 22.3 bits (46), Expect(2) = 0.28
Identities = 9/18 (50%), Positives = 12/18 (66%)
Frame = +1
Query: 253 HVDCRRLVLENWAKNVRG 270
+VDC R V E W+ +V G
Sbjct: 286 NVDCVRFVEEXWSGHVLG 339
>TC206673
Length = 742
Score = 35.0 bits (79), Expect = 0.19
Identities = 18/75 (24%), Positives = 34/75 (45%)
Frame = +3
Query: 299 DRQVRLATDEVSRIQQIIDSVGFSDDLYRQDLEAQLLLTNALNVQDQFWKEKSRHQSFIH 358
D+ + +++ ++ + + S D + E Q L A + ++KSR +
Sbjct: 12 DKGIHNIKKKLNEVEDLASNRSLSVDEIKAKKELQQQLWEASTAYESLLRQKSRDKWLKE 191
Query: 359 GDRNTAYFHRVARIK 373
GD N+AYFH V +
Sbjct: 192 GDNNSAYFHTVINFR 236
>TC234193
Length = 697
Score = 32.7 bits (73), Expect = 0.93
Identities = 17/61 (27%), Positives = 32/61 (51%)
Frame = -3
Query: 460 DGFGGHFYQSFWDIVGSDVVLSVQDFFLQGSFADNINSNLLVLIPKIPGAQAMGDFRPIA 519
+G+ +F + +++ DVV ++Q+F G + N++ L P +GDFRP+
Sbjct: 194 NGYNFNFIKKC*EVMKEDVVRAIQEFHSHGCLSRGTNASFLTHPP-----*GLGDFRPVT 30
Query: 520 L 520
L
Sbjct: 29 L 27
>TC234877
Length = 534
Score = 32.0 bits (71), Expect = 1.6
Identities = 18/74 (24%), Positives = 31/74 (41%)
Frame = +3
Query: 742 FVGFWRWLNSFYLSWLPDVQRKTEANLFPSYF**NQDEACYLERFLAFYYGSCSIG*IRY 801
+ G W +FYL + + + + **N+ LE F+YG C+ I Y
Sbjct: 12 YSGILSWFFAFYLPQCSIIPGEASTSAS*ANC**NKSGTSVLEGEFTFFYGLCAADYIYY 191
Query: 802 TWYVTFLLSCLYVA 815
W + + + +A
Sbjct: 192 RWNASLIFPLILLA 233
>BG405621
Length = 402
Score = 31.2 bits (69), Expect = 2.7
Identities = 12/37 (32%), Positives = 23/37 (61%)
Frame = +2
Query: 331 EAQLLLTNALNVQDQFWKEKSRHQSFIHGDRNTAYFH 367
EAQ L N + ++ +W+++S+ +GD N+ +FH
Sbjct: 287 EAQAKLNNFIAQEESYWQQRSKIFWLKYGDLNSKFFH 397
>TC216051 weakly similar to UP|Q8GTJ9 (Q8GTJ9) Cysteine proteinase, partial
(58%)
Length = 1281
Score = 30.8 bits (68), Expect = 3.5
Identities = 11/25 (44%), Positives = 19/25 (76%)
Frame = +2
Query: 50 YWHSIGVKKCCLNNRGPRLPNLWAL 74
++H + K+CC +N+GPRL ++W L
Sbjct: 449 FYHGLEEKRCCHSNQGPRL-HMWKL 520
>TC215281
Length = 733
Score = 30.8 bits (68), Expect = 3.5
Identities = 19/51 (37%), Positives = 22/51 (42%), Gaps = 5/51 (9%)
Frame = -1
Query: 1090 VCGGALELALFTAAHWAG-----FLLH*GAAVLSPTPLFFSSGGLVCCCGC 1135
V GGA L L W FLL + T LFF+S +C CGC
Sbjct: 394 VAGGASILLLHLLEDWISL*QLVFLLQPSKDPIKKTLLFFASTSTLCRCGC 242
>CA937241 similar to GP|6840918|gb|A hematopoietic antigen CD38 {Canis
familiaris}, partial (4%)
Length = 203
Score = 29.6 bits (65), Expect = 7.9
Identities = 15/60 (25%), Positives = 32/60 (53%), Gaps = 1/60 (1%)
Frame = +2
Query: 365 YFHRVARIKASSKNI-SLLYDGATVLSDAGDIAAHILNYFQGIFDVVNNCIPNDLVARSI 423
+FH A+ + S+ NI SL D + ++ + +++ ++ FQ +F N C + + +I
Sbjct: 5 FFHSSAKARKSANNISSLTCDDSAIVHNQKEMSNTTISCFQNLF*ANNTCTDSQPIINNI 184
>TC230212 similar to GB|AAP78934.1|32306501|BT009666 At5g12120 {Arabidopsis
thaliana;} , partial (9%)
Length = 971
Score = 29.6 bits (65), Expect = 7.9
Identities = 17/61 (27%), Positives = 28/61 (45%)
Frame = +2
Query: 219 LIRHQSDHHPLLLSADVASVKNAISFKFYKAWTSHVDCRRLVLENWAKNVRGVGMVRLQG 278
L+R Q DH+PL++ AW +DC+ + V G+G++ L+G
Sbjct: 491 LLRRQLDHYPLMV-----------------AWFLCLDCKMV*YRLLKHQVLGIGVLHLKG 619
Query: 279 K 279
K
Sbjct: 620 K 622
>TC217688 similar to UP|Q6K709 (Q6K709) DNA-binding protein-like, partial
(61%)
Length = 722
Score = 29.6 bits (65), Expect = 7.9
Identities = 10/21 (47%), Positives = 15/21 (70%)
Frame = -3
Query: 352 RHQSFIHGDRNTAYFHRVARI 372
RH++ +HG +FHRVAR+
Sbjct: 186 RHRTLVHGKPQIRFFHRVARV 124
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.346 0.153 0.561
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 65,405,104
Number of Sequences: 63676
Number of extensions: 1186900
Number of successful extensions: 12196
Number of sequences better than 10.0: 40
Number of HSP's better than 10.0 without gapping: 8861
Number of HSP's successfully gapped in prelim test: 346
Number of HSP's that attempted gapping in prelim test: 2876
Number of HSP's gapped (non-prelim): 9846
length of query: 1145
length of database: 12,639,632
effective HSP length: 108
effective length of query: 1037
effective length of database: 5,762,624
effective search space: 5975841088
effective search space used: 5975841088
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 38 (21.7 bits)
S2: 64 (29.3 bits)
Medicago: description of AC146565.8


