

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146564.3 - phase: 0 /pseudo
(281 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
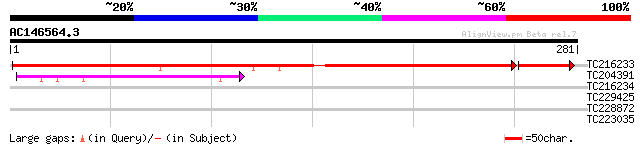
Score E
Sequences producing significant alignments: (bits) Value
TC216233 GB|CAA50573.1|402753|GMFUSA translation elongation fact... 261 7e-75
TC204391 homologue to UP|Q9ASR1 (Q9ASR1) At1g56070/T6H22_13 (Elo... 52 4e-07
TC216234 GB|CAA50573.1|402753|GMFUSA translation elongation fact... 34 0.065
TC229425 similar to UP|Q9FNM5 (Q9FNM5) GTP-binding protein LepA ... 28 3.6
TC228872 similar to UP|SEC8_ARATH (Q93YU5) Probable exocyst comp... 28 3.6
TC223035 similar to UP|Q73IE1 (Q73IE1) Elongation factor Tu fami... 28 4.6
>TC216233 GB|CAA50573.1|402753|GMFUSA translation elongation factor EF-G
{Glycine max;} , complete
Length = 2424
Score = 261 bits (667), Expect(2) = 7e-75
Identities = 150/267 (56%), Positives = 179/267 (66%), Gaps = 17/267 (6%)
Frame = +1
Query: 2 DIVTLVGLKDTIAGETLCDPKGPFVLDQMDFPVHVIKIAIEPKTKADMEKMEAGLIKLAE 61
DI+ L GLKDTI GETLCDP P VL++MDFP VIK+AIEPKTKAD++KM GLIKLA+
Sbjct: 1138 DIIALAGLKDTITGETLCDPDNPIVLERMDFPDPVIKVAIEPKTKADVDKMATGLIKLAQ 1317
Query: 62 EDPSFRVSQDKE-DRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESISEVTE- 119
EDPSF S+D+E ++T+I+G GELHLE IVDRLKREFKVEANVGAPQ NYRESIS++
Sbjct: 1318 EDPSFHFSRDEEINQTVIEGMGELHLEIIVDRLKREFKVEANVGAPQVNYRESISKICRS 1497
Query: 120 -------VRYVFNKPYILNP--------WTQVVDTNLRIKSKKKHFQKNSLYWGF*RIGK 164
+R+ + P W V + N R K+ W RIG+
Sbjct: 1498 EVCAQETIRWTGSVCRYHCPV*THGPR*WI*VQE*NQRRCCTKRIHS-----WCDERIGR 1662
Query: 165 LHEQWCSSRLSGCWCTRSTCG*YLP*CGFKCEGIPVGSKPSFYGRNEKSQTINA*TYNET 224
++EQWC+ LSG CT ST +LP C FKC G PVGSK SF GRN+KS T +A* YNE
Sbjct: 1663 VYEQWCACWLSGR*CTSSTHRWFLPRCRFKCVGFPVGSKRSF*GRNKKSWTKDA*AYNEG 1842
Query: 225 *SCYS*RTL*RCTW*SQQKKRPNHKCW 251
*SCYS*RT C W*SQ KKR + + W
Sbjct: 1843 *SCYS*RTSR*CNW*SQLKKRTDQQFW 1923
Score = 37.4 bits (85), Expect(2) = 7e-75
Identities = 16/28 (57%), Positives = 22/28 (78%)
Frame = +2
Query: 253 LTEGRASYSMQFDRFDTVSRNIQNELTT 280
+T+GRASY+MQ FD V ++IQN+L T
Sbjct: 2006 MTKGRASYTMQLAMFDVVPQHIQNQLAT 2089
>TC204391 homologue to UP|Q9ASR1 (Q9ASR1) At1g56070/T6H22_13 (Elongation factor
EF-2), complete
Length = 2941
Score = 51.6 bits (122), Expect = 4e-07
Identities = 37/119 (31%), Positives = 62/119 (52%), Gaps = 6/119 (5%)
Frame = +1
Query: 4 VTLVGLKDTIA-GETLCDPK--GPFVLDQMDFPVH-VIKIAIEPKTKADMEKMEAGLIKL 59
V +VGL I TL + K + M F V V+++A++ K +D+ K+ GL +L
Sbjct: 1474 VAMVGLDQFITKNATLTNEKEVDAHPIRAMKFSVSPVVRVAVQCKVASDLPKLVEGLKRL 1653
Query: 60 AEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLKREFKVEANV--GAPQANYRESISE 116
A+ DP + ++ I+ G GELHLE + L+ +F A + P ++RE++ E
Sbjct: 1654 AKSDPMVVCTIEESGEHIVAGAGELHLEICLKDLQDDFMGGAEIIKSDPVVSFRETVLE 1830
>TC216234 GB|CAA50573.1|402753|GMFUSA translation elongation factor EF-G
{Glycine max;} , partial (5%)
Length = 402
Score = 34.3 bits (77), Expect = 0.065
Identities = 15/25 (60%), Positives = 19/25 (76%)
Frame = +3
Query: 256 GRASYSMQFDRFDTVSRNIQNELTT 280
GRASY+MQ FD V ++IQN+L T
Sbjct: 18 GRASYTMQLAMFDVVPQHIQNQLAT 92
>TC229425 similar to UP|Q9FNM5 (Q9FNM5) GTP-binding protein LepA homolog,
partial (38%)
Length = 1009
Score = 28.5 bits (62), Expect = 3.6
Identities = 12/30 (40%), Positives = 18/30 (60%)
Frame = +2
Query: 82 GELHLETIVDRLKREFKVEANVGAPQANYR 111
G LH+E + +RL+RE+ + AP YR
Sbjct: 11 GLLHMEIVQERLEREYNLSLITTAPSVVYR 100
>TC228872 similar to UP|SEC8_ARATH (Q93YU5) Probable exocyst complex
component Sec8, partial (23%)
Length = 1260
Score = 28.5 bits (62), Expect = 3.6
Identities = 10/19 (52%), Positives = 13/19 (67%)
Frame = +3
Query: 169 WCSSRLSGCWCTRSTCG*Y 187
WCS+ LS C+C R+ G Y
Sbjct: 564 WCSANLSKCYCIRTGIGSY 620
>TC223035 similar to UP|Q73IE1 (Q73IE1) Elongation factor Tu family protein,
partial (13%)
Length = 443
Score = 28.1 bits (61), Expect = 4.6
Identities = 17/64 (26%), Positives = 31/64 (47%)
Frame = +2
Query: 56 LIKLAEEDPSFRVSQDKEDRTIIQGTGELHLETIVDRLKREFKVEANVGAPQANYRESIS 115
L+ AE + + V + +QG GEL L +++ ++RE E +V P+ Y+
Sbjct: 17 LMAEAETNLAINVLPGLSESFEVQGRGELQLGILIENMRRE-GFELSVSPPKVMYKTESG 193
Query: 116 EVTE 119
+ E
Sbjct: 194 QKLE 205
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.334 0.145 0.477
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 11,720,836
Number of Sequences: 63676
Number of extensions: 170784
Number of successful extensions: 1480
Number of sequences better than 10.0: 12
Number of HSP's better than 10.0 without gapping: 1458
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1478
length of query: 281
length of database: 12,639,632
effective HSP length: 96
effective length of query: 185
effective length of database: 6,526,736
effective search space: 1207446160
effective search space used: 1207446160
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.6 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146564.3


