

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146564.14 - phase: 0 /pseudo
(312 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
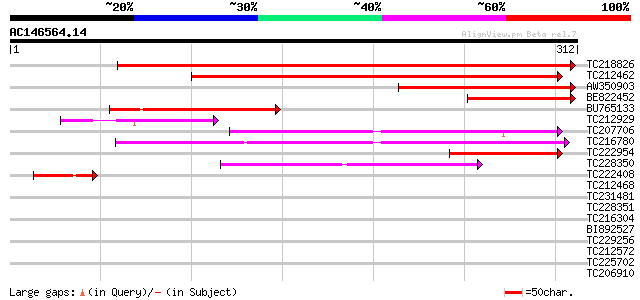
Score E
Sequences producing significant alignments: (bits) Value
TC218826 weakly similar to UP|Q6NQK3 (Q6NQK3) At1g12450, partial... 419 e-117
TC212462 similar to UP|Q9ZVQ5 (Q9ZVQ5) Expressed protein (At2g02... 228 3e-60
AW350903 137 8e-33
BE822452 80 9e-16
BU765133 similar to GP|13605571|gb| At2g02370/T16F16.16 {Arabido... 77 8e-15
TC212929 72 2e-13
TC207706 72 3e-13
TC216780 similar to UP|Q8H3D8 (Q8H3D8) Integral membrane protein... 72 4e-13
TC222954 similar to UP|Q9ZVQ5 (Q9ZVQ5) Expressed protein (At2g02... 64 9e-11
TC228350 similar to UP|Q75IA4 (Q75IA4) Expressed protein, partia... 51 8e-07
TC222408 43 2e-04
TC212468 similar to UP|Q8H3D8 (Q8H3D8) Integral membrane protein... 34 0.075
TC231481 similar to UP|Q9MAL1 (Q9MAL1) F27F5.5, partial (44%) 34 0.075
TC228351 similar to UP|Q75IA4 (Q75IA4) Expressed protein, partia... 30 1.4
TC216304 weakly similar to UP|Q8VWX5 (Q8VWX5) Ribosome-like prot... 29 2.4
BI892527 29 2.4
TC229256 28 4.1
TC212572 28 4.1
TC225702 UP|Q9ZRD8 (Q9ZRD8) GMFP5 (Fragment), partial (98%) 28 4.1
TC206910 weakly similar to UP|Q84ZX0 (Q84ZX0) HEN4, partial (9%) 28 4.1
>TC218826 weakly similar to UP|Q6NQK3 (Q6NQK3) At1g12450, partial (75%)
Length = 1078
Score = 419 bits (1076), Expect = e-117
Identities = 200/252 (79%), Positives = 225/252 (88%)
Frame = +1
Query: 60 ECSLPSRRSVVWYWVKMVLLFLSLGFLAVAVLKWVGPYLIDKEVIPIINWETETFSPPVL 119
E S P+ R+ VWYWVK+V+LFL LGFLAV VL WVGPY IDKE+IPIINWETETFS PVL
Sbjct: 31 EPSPPTIRAAVWYWVKLVVLFLYLGFLAVVVLVWVGPYFIDKEIIPIINWETETFSTPVL 210
Query: 120 TILLFASVAIFPTILLPSTPSMWVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSIFHHKI 179
T+L+F SVAIFPT+LLPST SMWVAG+T GYGFGFLLII+AAAIGVSLPF+IG +FHHKI
Sbjct: 211 TVLVFTSVAIFPTLLLPSTTSMWVAGMTFGYGFGFLLIISAAAIGVSLPFVIGKLFHHKI 390
Query: 180 EGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISPFPYMVFNYCAVATNVKYGPYIIGS 239
EGWLEKYPKKASIL+SAG G+WFHQFRAVA IRISPFPY++FNYCAVATNVKYGPY++GS
Sbjct: 391 EGWLEKYPKKASILRSAGGGSWFHQFRAVAFIRISPFPYLIFNYCAVATNVKYGPYMVGS 570
Query: 240 LVGMVPEIFVAIYTGILIKTLADASNERQSLSAQQIILNALGFCLTVVTTVIITVYAKRR 299
LVGMVPEIFVAIYTGILI+TLADAS E+ SLSA QIILN GFC+TV TT+ T YA+RR
Sbjct: 571 LVGMVPEIFVAIYTGILIRTLADASYEKHSLSAPQIILNVAGFCITVATTIFFTAYARRR 750
Query: 300 LKELQEEDKMLL 311
L ELQ E++ LL
Sbjct: 751 LDELQREEEPLL 786
>TC212462 similar to UP|Q9ZVQ5 (Q9ZVQ5) Expressed protein
(At2g02370/T16F16.16), partial (66%)
Length = 712
Score = 228 bits (581), Expect = 3e-60
Identities = 101/204 (49%), Positives = 144/204 (70%)
Frame = +3
Query: 101 KEVIPIINWETETFSPPVLTILLFASVAIFPTILLPSTPSMWVAGVTLGYGFGFLLIITA 160
K + PI+ WE F PVL ++L AS+A+FP L+PS PSMW+AG+ GYG GF +I+
Sbjct: 12 KVLYPIMEWEATAFGRPVLALVLVASLALFPVFLIPSGPSMWLAGMIFGYGLGFFIIMVG 191
Query: 161 AAIGVSLPFIIGSIFHHKIEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISPFPYMV 220
IG+ LP++IG +F +I WL+++P+ A++++ AG GNW QF+ VAL R+SPFPY +
Sbjct: 192 TTIGMVLPYLIGLLFRDRIHQWLKRWPQNAAMIRLAGEGNWSRQFQVVALFRVSPFPYTI 371
Query: 221 FNYCAVATNVKYGPYIIGSLVGMVPEIFVAIYTGILIKTLADASNERQSLSAQQIILNAL 280
FNY V TN+++ PY+ GS+ GMVPE F+ IY+G LIKTLADA + L+ +I+ N +
Sbjct: 372 FNYAVVVTNMRFWPYLCGSVAGMVPEAFIYIYSGRLIKTLADAQYGKHHLTTVEIVYNII 551
Query: 281 GFCLTVVTTVIITVYAKRRLKELQ 304
F + +VTT+ TVYAKR L EL+
Sbjct: 552 SFIIAIVTTIAFTVYAKRTLNELK 623
>AW350903
Length = 689
Score = 137 bits (344), Expect = 8e-33
Identities = 62/97 (63%), Positives = 82/97 (83%)
Frame = -3
Query: 215 PFPYMVFNYCAVATNVKYGPYIIGSLVGMVPEIFVAIYTGILIKTLADASNERQSLSAQQ 274
PFPY++FNYCAVATNVKY Y++GSLVGMVPEIFV+IYTGILI+ LA+AS++ +LS Q
Sbjct: 687 PFPYIIFNYCAVATNVKYXXYLLGSLVGMVPEIFVSIYTGILIEALANASHQNHTLSTPQ 508
Query: 275 IILNALGFCLTVVTTVIITVYAKRRLKELQEEDKMLL 311
I+LN +GFC+TV T + T Y+KR+LKELQ+++ L+
Sbjct: 507 IVLNVVGFCVTVATIIFFTAYSKRQLKELQQKENDLV 397
>BE822452
Length = 758
Score = 80.5 bits (197), Expect = 9e-16
Identities = 39/59 (66%), Positives = 48/59 (81%)
Frame = -2
Query: 253 TGILIKTLADASNERQSLSAQQIILNALGFCLTVVTTVIITVYAKRRLKELQEEDKMLL 311
+GILI+TLADAS+E+ SLSA QIILN GFC+TV TT+ T YA+RRL ELQ E++ LL
Sbjct: 469 SGILIRTLADASHEKHSLSAPQIILNVAGFCITVATTIFFTAYARRRLDELQREEEPLL 293
>BU765133 similar to GP|13605571|gb| At2g02370/T16F16.16 {Arabidopsis
thaliana}, partial (20%)
Length = 430
Score = 77.4 bits (189), Expect = 8e-15
Identities = 36/94 (38%), Positives = 57/94 (60%)
Frame = +2
Query: 56 EMVEECSLPSRRSVVWYWVKMVLLFLSLGFLAVAVLKWVGPYLIDKEVIPIINWETETFS 115
EM++ + +S +W +K+ L + L + +LKW P+ +K + PI+ WE TF
Sbjct: 152 EMLQPLADSRMKSFMWC-IKVFLWCFIIVILGLVILKWGVPFTFEKVLYPIMEWEATTFG 328
Query: 116 PPVLTILLFASVAIFPTILLPSTPSMWVAGVTLG 149
PVL ++L AS+A+FP +PS PSMW+AG+ G
Sbjct: 329 RPVLALVLVASLALFPVFFIPSGPSMWLAGMIFG 430
>TC212929
Length = 456
Score = 72.4 bits (176), Expect = 2e-13
Identities = 37/90 (41%), Positives = 52/90 (57%), Gaps = 3/90 (3%)
Frame = +2
Query: 29 IEADDDNKGDYIKLIPGSDECLPLTAVEMVEECSLPSRR---SVVWYWVKMVLLFLSLGF 85
+E + D G+Y+KL+ + ++P+ R S +WYWVK+VL FL LG
Sbjct: 218 VERNHDGGGEYVKLVWDP------------QPEAVPTHRGGSSRLWYWVKLVLCFLCLGL 361
Query: 86 LAVAVLKWVGPYLIDKEVIPIINWETETFS 115
LA+ +WV P I+K +IPII WET TFS
Sbjct: 362 LALVAFEWVAPLFIEKVIIPIIKWETNTFS 451
>TC207706
Length = 1197
Score = 72.0 bits (175), Expect = 3e-13
Identities = 49/187 (26%), Positives = 89/187 (47%), Gaps = 4/187 (2%)
Frame = +2
Query: 122 LLFASVAIFPTILLPSTPSMWVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSIF-HHKIE 180
L A A + +P+ P AG+ G G +++ + + S+ F+I F +I
Sbjct: 44 LFVAVYAGLEILAIPAIPLTMSAGLLFGSVVGTIIVSISGTVAASVAFLIARYFARERIV 223
Query: 181 GWLEKYPKKASILKSAGAGNWFHQFRAVALIRISPF-PYMVFNYCAVATNVKYGPYIIGS 239
+E K +I K+ G FR V L+R+SP P+ + NY T+VK+ PY++GS
Sbjct: 224 KLVEGNKKFVAIDKAIGENG----FRVVTLLRLSPLLPFSLGNYLYGLTSVKFIPYVLGS 391
Query: 240 LVGMVPEIFVAIYTGILIKTLADASNERQSL--SAQQIILNALGFCLTVVTTVIITVYAK 297
+GM+P + + G + + +E ++ Q++ LG T + +T AK
Sbjct: 392 WLGMLPGTWAYVSAGAFGRAIIQDESELSNIFGGNSQLLTLGLGLLATALAAAYVTRLAK 571
Query: 298 RRLKELQ 304
+K+++
Sbjct: 572 DAIKDIE 592
>TC216780 similar to UP|Q8H3D8 (Q8H3D8) Integral membrane protein-like,
partial (82%)
Length = 1381
Score = 71.6 bits (174), Expect = 4e-13
Identities = 61/252 (24%), Positives = 107/252 (42%), Gaps = 2/252 (0%)
Frame = +3
Query: 59 EECSLPSRRSVVWYWVKMVLLFLSLGFLAVAVLKWVGPYLIDKEVIPIINWETETFSPPV 118
EEC RR ++ + L L L AV+ I+K + + W P
Sbjct: 294 EECRPWRRRLIMSFSWASTLRITILLLLLAAVIVACFTLPIEKMMKDFLLWVDHDLGPWG 473
Query: 119 LTILLFASVAIFPTILLPSTPSMWVAGVTLGYGFGFLLIITAAAIGVSLPFIIG-SIFHH 177
+L A + + + +P++ G G GF+ A +G F++G +I
Sbjct: 474 PLVLAVAYIPL-TVLAVPASVLTLGGGYLFGLPVGFVADSIGATVGAGAAFLLGRTIGRS 650
Query: 178 KIEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISPF-PYMVFNYCAVATNVKYGPYI 236
+ L+ YP+ S+ + F+ V L+R+ P P+ + NY T V G Y+
Sbjct: 651 FVVSRLKDYPQFRSVAIAIRRSG----FKIVLLLRLVPLLPFNMLNYLLSVTPVSIGEYM 818
Query: 237 IGSLVGMVPEIFVAIYTGILIKTLADASNERQSLSAQQIILNALGFCLTVVTTVIITVYA 296
+ S +GM+P +Y G +K L+D ++ S + LG ++VV + +T A
Sbjct: 819 LASWLGMMPITLALVYVGTTLKDLSDVTHGWGEFSKTRWAFIILGLVISVVLMICVTKVA 998
Query: 297 KRRLKELQEEDK 308
K L + E++
Sbjct: 999 KAALDKALAENE 1034
>TC222954 similar to UP|Q9ZVQ5 (Q9ZVQ5) Expressed protein
(At2g02370/T16F16.16), partial (21%)
Length = 365
Score = 63.9 bits (154), Expect = 9e-11
Identities = 30/62 (48%), Positives = 43/62 (68%)
Frame = +2
Query: 243 MVPEIFVAIYTGILIKTLADASNERQSLSAQQIILNALGFCLTVVTTVIITVYAKRRLKE 302
MVPE+F+ +Y+G LI+TLADA + L+ +I+ N + F + VVTT+ VYAKR L E
Sbjct: 2 MVPEVFIYMYSGRLIRTLADAQYGKHQLTIVEIVYNIISFIVAVVTTIAFIVYAKRTLNE 181
Query: 303 LQ 304
L+
Sbjct: 182 LK 187
>TC228350 similar to UP|Q75IA4 (Q75IA4) Expressed protein, partial (65%)
Length = 1410
Score = 50.8 bits (120), Expect = 8e-07
Identities = 37/147 (25%), Positives = 66/147 (44%), Gaps = 3/147 (2%)
Frame = +1
Query: 117 PVLTILLFASVAIF-PTILLPSTPSM-WVAGVTLGYGFGFLLIITAAAIGVSLPFIIGSI 174
P IL + S IF T ++P T M +AG G G LL++ A G S F + +
Sbjct: 466 PAQFILGYCSTYIFMQTFMIPGTIFMSLLAGALFGVVRGILLVVFNATAGASSCFFLSKL 645
Query: 175 FHHKIEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISP-FPYMVFNYCAVATNVKYG 233
+ WL +P+K ++ A + +RI+P P + N + +V +
Sbjct: 646 IGRPLVSWL--WPEKLRFFQAEIAKRRDRLLNYMLFLRITPTLPNLFINLASPIVDVPFH 819
Query: 234 PYIIGSLVGMVPEIFVAIYTGILIKTL 260
+ +L+G+VP ++ + G+ + L
Sbjct: 820 IFFSATLIGLVPASYITVRAGLALGDL 900
>TC222408
Length = 489
Score = 43.1 bits (100), Expect = 2e-04
Identities = 20/35 (57%), Positives = 28/35 (79%)
Frame = +2
Query: 14 GGGWKKEVVSDVTITIEADDDNKGDYIKLIPGSDE 48
GGG ++EVV DVT+TI++DD N GDY+KL +D+
Sbjct: 386 GGGRREEVVPDVTLTIQSDDGN-GDYVKLRANADD 487
>TC212468 similar to UP|Q8H3D8 (Q8H3D8) Integral membrane protein-like,
partial (23%)
Length = 617
Score = 34.3 bits (77), Expect = 0.075
Identities = 15/45 (33%), Positives = 22/45 (48%)
Frame = +1
Query: 220 VFNYCAVATNVKYGPYIIGSLVGMVPEIFVAIYTGILIKTLADAS 264
+ NY T V G Y + S +GM+P +Y G K L+D +
Sbjct: 7 ILNYLLSVTPVPLGEYTLASWLGMMPITLALVYVGTTFKDLSDVT 141
>TC231481 similar to UP|Q9MAL1 (Q9MAL1) F27F5.5, partial (44%)
Length = 677
Score = 34.3 bits (77), Expect = 0.075
Identities = 27/94 (28%), Positives = 46/94 (48%), Gaps = 2/94 (2%)
Frame = +3
Query: 205 FRAVALIRISPFPYMVFNYCAVATNVKY-GPYIIGSLVGMVPEIFVAIYTGILI-KTLAD 262
++ V L R SP P V NY AT+V++ +++ + +G +P I G L +A
Sbjct: 99 WKFVLLARFSPVPSYVINYTLAATDVRFLLDFLLPTAIGCLPMILQNTSIGSLAGAAVAT 278
Query: 263 ASNERQSLSAQQIILNALGFCLTVVTTVIITVYA 296
AS ++S I +G +VV ++ I Y+
Sbjct: 279 ASGSKKS-QIWSYIFPVVGILSSVVISLRIKKYS 377
>TC228351 similar to UP|Q75IA4 (Q75IA4) Expressed protein, partial (44%)
Length = 854
Score = 30.0 bits (66), Expect = 1.4
Identities = 18/93 (19%), Positives = 41/93 (43%), Gaps = 1/93 (1%)
Frame = +1
Query: 169 FIIGSIFHHKIEGWLEKYPKKASILKSAGAGNWFHQFRAVALIRISP-FPYMVFNYCAVA 227
F+ + + WL +P+K ++ A + +RI+P P + N +
Sbjct: 37 FLFVQLIGRPLVSWL--WPEKLWFFQAEIAKRRDRLLNYMLFLRITPTLPNLFINLASPI 210
Query: 228 TNVKYGPYIIGSLVGMVPEIFVAIYTGILIKTL 260
+V + + +L+G+VP ++ + G+ + L
Sbjct: 211 VDVPFHIFFSATLIGLVPASYITVRAGLALGDL 309
>TC216304 weakly similar to UP|Q8VWX5 (Q8VWX5) Ribosome-like protein
(Fragment), partial (59%)
Length = 900
Score = 29.3 bits (64), Expect = 2.4
Identities = 14/40 (35%), Positives = 24/40 (60%), Gaps = 6/40 (15%)
Frame = -2
Query: 7 DCGGGGGGGGW------KKEVVSDVTITIEADDDNKGDYI 40
+CGG GGGGG+ +K ++ D E+D+D+ G+ +
Sbjct: 320 NCGGEGGGGGFVRKKKERKLLMVDEAAMGESDEDDHGNNV 201
>BI892527
Length = 423
Score = 29.3 bits (64), Expect = 2.4
Identities = 19/50 (38%), Positives = 24/50 (48%)
Frame = -1
Query: 9 GGGGGGGGWKKEVVSDVTITIEADDDNKGDYIKLIPGSDECLPLTAVEMV 58
GGGG GGG + EVV D T D D D + + G + L V +V
Sbjct: 174 GGGGEGGGGEGEVVEDKDAT-SGDADADADAVAVTGGDVALVELGIVWVV 28
>TC229256
Length = 867
Score = 28.5 bits (62), Expect = 4.1
Identities = 21/54 (38%), Positives = 27/54 (49%), Gaps = 1/54 (1%)
Frame = +1
Query: 101 KEVIPIINWETETFSPPVLTILLFASVAIFPTILLPSTPSMWVAG-VTLGYGFG 153
K+V+ + NW + TFS P ILL S A P V G V++GY FG
Sbjct: 307 KDVVAVCNWLSNTFSLP--RILLLGSSAGAPIAGSAVDQIEQVIGYVSIGYPFG 462
>TC212572
Length = 706
Score = 28.5 bits (62), Expect = 4.1
Identities = 22/65 (33%), Positives = 33/65 (49%), Gaps = 6/65 (9%)
Frame = -1
Query: 99 IDKEVIPIINWETETFSPPVLTILLFAS-----VAIFPTI-LLPSTPSMWVAGVTLGYGF 152
+ + V P + T T S L++L A+ V++F + L S P M A LG+GF
Sbjct: 295 LQEHVNPTSHDPTTTTSSSSLSLLTLATAPPSIVSLFRLVSFLKSEPKMPKALFGLGFGF 116
Query: 153 GFLLI 157
FLL+
Sbjct: 115 AFLLV 101
>TC225702 UP|Q9ZRD8 (Q9ZRD8) GMFP5 (Fragment), partial (98%)
Length = 1431
Score = 28.5 bits (62), Expect = 4.1
Identities = 12/18 (66%), Positives = 13/18 (71%)
Frame = +3
Query: 3 DNTDDCGGGGGGGGWKKE 20
D + GGGGGGGG KKE
Sbjct: 684 DKKEKEGGGGGGGGEKKE 737
>TC206910 weakly similar to UP|Q84ZX0 (Q84ZX0) HEN4, partial (9%)
Length = 1862
Score = 28.5 bits (62), Expect = 4.1
Identities = 10/47 (21%), Positives = 30/47 (63%)
Frame = +3
Query: 46 SDECLPLTAVEMVEECSLPSRRSVVWYWVKMVLLFLSLGFLAVAVLK 92
++ + ++A+E +E C+ P++ +V+ + +++ + GFL V+ ++
Sbjct: 612 AERIVTISAIESLESCNSPAQDAVILVFARIIEDHIGKGFLQVSSME 752
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.142 0.435
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,019,134
Number of Sequences: 63676
Number of extensions: 231228
Number of successful extensions: 3777
Number of sequences better than 10.0: 64
Number of HSP's better than 10.0 without gapping: 2190
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3209
length of query: 312
length of database: 12,639,632
effective HSP length: 97
effective length of query: 215
effective length of database: 6,463,060
effective search space: 1389557900
effective search space used: 1389557900
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146564.14


