

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146559.3 + phase: 0
(130 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
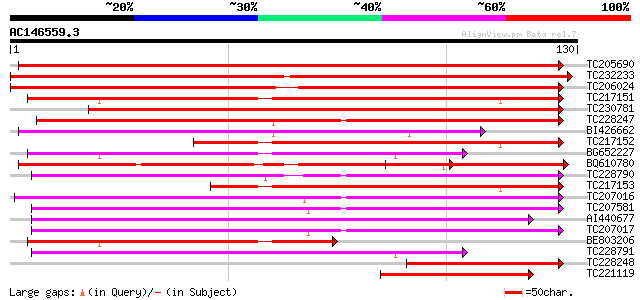
Score E
Sequences producing significant alignments: (bits) Value
TC205690 weakly similar to UP|Q6K3E9 (Q6K3E9) F-box family prote... 181 9e-47
TC232233 weakly similar to GB|AAT41865.1|48310676|BT014882 At1g8... 159 5e-40
TC206024 weakly similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific le... 149 5e-37
TC217151 similar to UP|Q6NPT8 (Q6NPT8) At2g02230, partial (6%) 137 1e-33
TC230781 weakly similar to UP|Q6K3E9 (Q6K3E9) F-box family prote... 132 6e-32
TC228247 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific le... 110 1e-25
BI426662 103 2e-23
TC217152 similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific lectin-li... 100 3e-22
BG652227 98 1e-21
BQ610780 94 1e-20
TC228790 94 2e-20
TC217153 similar to GB|AAS76702.1|45773796|BT012215 At1g56240 {A... 94 2e-20
TC207016 82 5e-17
TC207581 79 6e-16
AI440677 77 2e-15
TC207017 77 2e-15
BE803206 71 1e-13
TC228791 61 2e-10
TC228248 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific le... 49 5e-07
TC221119 similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific lectin-li... 48 1e-06
>TC205690 weakly similar to UP|Q6K3E9 (Q6K3E9) F-box family protein-like,
partial (33%)
Length = 1360
Score = 181 bits (459), Expect = 9e-47
Identities = 86/125 (68%), Positives = 102/125 (80%)
Frame = +2
Query: 3 EIQDLPEGCIAAIVSRTTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSPSL 62
E Q LPEGCIA+I+SRTTP D R SV SK FRSA++SDAVW +FLPSD SIIS SPS
Sbjct: 203 EFQGLPEGCIASILSRTTPADVCRFSVVSKIFRSAAESDAVWKRFLPSDYHSIISQSPSP 382
Query: 63 TNIPTKKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSARSLAIVWGDDRRYWNWIS 122
N P+KK+LYLALSDRP+IID KKS QLE+KSGKKCYML+AR+L+I+WGD +YWNW +
Sbjct: 383 LNYPSKKELYLALSDRPIIIDQGKKSFQLEKKSGKKCYMLAARALSIIWGDTEQYWNWTT 562
Query: 123 MPDSR 127
+SR
Sbjct: 563 DTNSR 577
>TC232233 weakly similar to GB|AAT41865.1|48310676|BT014882 At1g80110
{Arabidopsis thaliana;} , partial (47%)
Length = 687
Score = 159 bits (401), Expect = 5e-40
Identities = 76/129 (58%), Positives = 99/129 (75%)
Frame = +1
Query: 1 MTEIQDLPEGCIAAIVSRTTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSP 60
+ + +LPEGCIA I+S T+P D RLS+ S TFRSA+ SDAVWN+FLPSD +I+S S
Sbjct: 7 VVDFNNLPEGCIANILSFTSPRDVCRLSLLSSTFRSAAQSDAVWNKFLPSDFHTILSQSS 186
Query: 61 SLTNIPTKKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSARSLAIVWGDDRRYWNW 120
SL+ +P+KKDL+L L +P++ID KKS QL++ GKKCYMLSAR+L IVWGD RYW W
Sbjct: 187 SLS-LPSKKDLFLYLCQKPLLIDDGKKSYQLDKVYGKKCYMLSARNLFIVWGDTPRYWRW 363
Query: 121 ISMPDSRLA 129
S+PD+R +
Sbjct: 364 TSLPDARFS 390
>TC206024 weakly similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific lectin-like
protein, partial (47%)
Length = 954
Score = 149 bits (375), Expect = 5e-37
Identities = 72/127 (56%), Positives = 95/127 (74%)
Frame = +3
Query: 1 MTEIQDLPEGCIAAIVSRTTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSP 60
+ E+QDLPEGCIA I+S TTP+D RLS+ SK FRSA++SD VW+ FL SD +SII S
Sbjct: 18 LMELQDLPEGCIAKILSYTTPVDVCRLSLVSKAFRSAAESDTVWDCFLLSDFTSIIPISS 197
Query: 61 SLTNIPTKKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSARSLAIVWGDDRRYWNW 120
+ +KKDLY LSD P II +KS+QL++++GKKC MLSAR+L I+WGD ++W W
Sbjct: 198 T-----SKKDLYFTLSDHPTIIHQGRKSVQLDKRTGKKCCMLSARNLTIIWGDTVQHWEW 362
Query: 121 ISMPDSR 127
S+P+SR
Sbjct: 363 TSLPESR 383
>TC217151 similar to UP|Q6NPT8 (Q6NPT8) At2g02230, partial (6%)
Length = 514
Score = 137 bits (345), Expect = 1e-33
Identities = 71/128 (55%), Positives = 91/128 (70%), Gaps = 5/128 (3%)
Frame = +3
Query: 5 QDLPEGCIAAIVSRT-TPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSPSLT 63
QDLPEGC+A I+S TP D RLS+ SK F SA+D D VW++F+PSD SS IS L+
Sbjct: 30 QDLPEGCVAHILSYICTPEDIVRLSLVSKAFYSAADYDTVWDRFIPSDFSSTIS---PLS 200
Query: 64 NIPTKKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSARSLAIVW----GDDRRYWN 119
+ +KKDLY LSDRP IID +KS QLE+++ KKCYMLSAR ++I W G+ +YW
Sbjct: 201 SSNSKKDLYFTLSDRPTIIDQGRKSFQLEKRTAKKCYMLSARDISITWAPTQGEASQYWE 380
Query: 120 WISMPDSR 127
W S+P+SR
Sbjct: 381 WKSLPESR 404
>TC230781 weakly similar to UP|Q6K3E9 (Q6K3E9) F-box family protein-like,
partial (29%)
Length = 971
Score = 132 bits (331), Expect = 6e-32
Identities = 63/109 (57%), Positives = 80/109 (72%)
Frame = +2
Query: 19 TTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSPSLTNIPTKKDLYLALSDR 78
TT D RLS+ SK F SA++++ VW+ FLPSD+SSIIS S S+ + KDLYL LSDR
Sbjct: 2 TTRADVCRLSLVSKAFHSAAEANTVWDCFLPSDLSSIISPSSSVPPFRSIKDLYLYLSDR 181
Query: 79 PVIIDHDKKSIQLERKSGKKCYMLSARSLAIVWGDDRRYWNWISMPDSR 127
P IID KS QLE+++G KCYMLSAR L+I+WGD YW W ++P+SR
Sbjct: 182 PTIIDQGIKSFQLEKRTGNKCYMLSARDLSIIWGDTTHYWEWTTLPESR 328
>TC228247 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific lectin
PP2-like protein, partial (46%)
Length = 1124
Score = 110 bits (276), Expect = 1e-25
Identities = 56/124 (45%), Positives = 81/124 (65%), Gaps = 3/124 (2%)
Frame = +3
Query: 7 LPEGCIAAIVSRTTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHS---PSLT 63
LPE C++ I+S T+P DA R S+ S T RS++DSD +W F PSD S I+S + SL
Sbjct: 81 LPEDCVSKILSYTSPPDACRFSMVSSTLRSSADSDLLWRTFFPSDYSDIVSRALNPLSLN 260
Query: 64 NIPTKKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSARSLAIVWGDDRRYWNWISM 123
+ + K L+ AL P+++D S +L++ SGKK Y+LSAR L+I W +D YW+W +
Sbjct: 261 SSSSYKHLFYALC-HPLLLDGGNMSFKLDKSSGKKSYILSARQLSITWSNDPLYWSWRPV 437
Query: 124 PDSR 127
P+SR
Sbjct: 438 PESR 449
>BI426662
Length = 421
Score = 103 bits (258), Expect = 2e-23
Identities = 65/137 (47%), Positives = 80/137 (57%), Gaps = 30/137 (21%)
Frame = +3
Query: 3 EIQDLPEGCIAAIVSRTTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHS--- 59
E + LPEGCIA IVS TTP DA LS+ S +FRSAS +D VW +FLPSD +IIS S
Sbjct: 9 EFEHLPEGCIANIVSFTTPPDACVLSLVSSSFRSASVTDFVWERFLPSDYQAIISQSSKP 188
Query: 60 PSLTNIPTKKDLYLALSDRPVIIDHDKKSIQ---------------------------LE 92
+LTN +KKDLYL L P++ID KK +Q L+
Sbjct: 189 STLTNYSSKKDLYLHLCHNPLLIDAGKKVLQIKSFFLSIIIHVYLSLIGKLINHQSFALD 368
Query: 93 RKSGKKCYMLSARSLAI 109
+ +GK CYMLSARSL+I
Sbjct: 369 KLNGKICYMLSARSLSI 419
>TC217152 similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific lectin-like protein,
partial (8%)
Length = 909
Score = 100 bits (248), Expect = 3e-22
Identities = 48/89 (53%), Positives = 64/89 (70%), Gaps = 4/89 (4%)
Frame = +3
Query: 43 VWNQFLPSDISSIISHSPSLTNIPTKKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYML 102
VW++F+PSD SS IS L++ +KKDLY LSDRP IID +KS QLE+++ KKCYML
Sbjct: 3 VWDRFIPSDFSSTIS---PLSSSNSKKDLYFTLSDRPTIIDQGRKSFQLEKRTAKKCYML 173
Query: 103 SARSLAIVW----GDDRRYWNWISMPDSR 127
SAR ++I W G+ +YW W S+P+SR
Sbjct: 174 SARDISITWAPTQGEASQYWEWKSLPESR 260
>BG652227
Length = 416
Score = 98.2 bits (243), Expect = 1e-21
Identities = 61/134 (45%), Positives = 75/134 (55%), Gaps = 33/134 (24%)
Frame = +3
Query: 5 QDLPEGCIAAIVSRT-TPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSPSLT 63
QDLPEGC+A I+S TP D RLS+ SK F SA+D D VW++F+PSD SS IS L+
Sbjct: 24 QDLPEGCVAHILSYICTPEDIVRLSLVSKAFYSAADYDTVWDRFIPSDFSSTIS---PLS 194
Query: 64 NIPTKKDLYLALSDRPVIIDHDKK--------------------------------SIQL 91
+ +KKDLY LSDRP IID +K S QL
Sbjct: 195 SSNSKKDLYFTLSDRPTIIDQGRKVRTLFLLACSDVFYGIPEMKCVVYVASLKFLQSFQL 374
Query: 92 ERKSGKKCYMLSAR 105
E+++ KKCYMLSAR
Sbjct: 375 EKRTAKKCYMLSAR 416
>BQ610780
Length = 447
Score = 94.4 bits (233), Expect = 1e-20
Identities = 56/102 (54%), Positives = 70/102 (67%), Gaps = 2/102 (1%)
Frame = +3
Query: 3 EIQDLPEGCIAAIVSRTTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSPSL 62
E+QDLPEGCIA I+S TTP+DA RLSV S FRSA++SD VW+ FL SD +S I PS
Sbjct: 3 ELQDLPEGCIAKILSYTTPVDACRLSV-SIAFRSAAESDTVWDCFLLSDFTSFI--PPSS 173
Query: 63 TNIPTKKDLYLALSDRPVIIDHDKKSIQLERKSGKK--CYML 102
T +K DLY LSD P I+D +KS+QL + G+ C+ L
Sbjct: 174 T---SKNDLYFTLSDLPTIMDQGRKSVQLAKGPGRSVTCFPL 290
Score = 62.8 bits (151), Expect = 4e-11
Identities = 24/42 (57%), Positives = 34/42 (80%)
Frame = +2
Query: 87 KSIQLERKSGKKCYMLSARSLAIVWGDDRRYWNWISMPDSRL 128
K + +++GKKCYMLSAR+LAI+WGD R+W W S+P+S+L
Sbjct: 236 KECSIGKRTGKKCYMLSARNLAIIWGDTVRHWEWTSLPESKL 361
>TC228790
Length = 1215
Score = 93.6 bits (231), Expect = 2e-20
Identities = 49/129 (37%), Positives = 73/129 (55%), Gaps = 7/129 (5%)
Frame = +2
Query: 6 DLPEGCIAAIVSRTTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIIS-------H 58
D+PE C+A + TP + L+ ++ FR A+ SD+VW LP + ++ H
Sbjct: 182 DIPESCVACVFLHLTPPEICNLARLNRAFRGAASSDSVWEAKLPRNYQDLLDLVPPPERH 361
Query: 59 SPSLTNIPTKKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSARSLAIVWGDDRRYW 118
SL+ KKD++ LS RP+ DH K + L+R +GK C +SA+++ I DDRRYW
Sbjct: 362 HRSLS----KKDIFALLS-RPLPFDHGHKEVWLDRVTGKVCMSISAKAMVITGIDDRRYW 526
Query: 119 NWISMPDSR 127
NWI +SR
Sbjct: 527 NWIPTEESR 553
>TC217153 similar to GB|AAS76702.1|45773796|BT012215 At1g56240 {Arabidopsis
thaliana;} , partial (13%)
Length = 635
Score = 93.6 bits (231), Expect = 2e-20
Identities = 46/85 (54%), Positives = 60/85 (70%), Gaps = 4/85 (4%)
Frame = +3
Query: 47 FLPSDISSIISHSPSLTNIPTKKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSARS 106
F+PSD SS IS L++ +KKDLY LSDRP IID +KS QLE+++ KKCYMLSAR
Sbjct: 3 FIPSDFSSTIS---PLSSSNSKKDLYFTLSDRPTIIDQGRKSFQLEKRTAKKCYMLSARD 173
Query: 107 LAIVW----GDDRRYWNWISMPDSR 127
++I W G+ +YW W S+P+SR
Sbjct: 174 ISITWAPTQGEASQYWEWKSLPESR 248
>TC207016
Length = 1310
Score = 82.4 bits (202), Expect = 5e-17
Identities = 44/128 (34%), Positives = 71/128 (55%), Gaps = 2/128 (1%)
Frame = +1
Query: 2 TEIQDLPEGCIAAIVSRTTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSPS 61
T + D+PE CI++++ P + L+ +KTF AS ++ VW LP + +++
Sbjct: 301 TSLDDIPENCISSMMMNFDPQEICSLARVNKTFHRASSANFVWESKLPQNYKFLLNKVLG 480
Query: 62 LTNIP--TKKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSARSLAIVWGDDRRYWN 119
N+ TKK++Y L +P D K + L+R SG+ C +S++S I DDRRYWN
Sbjct: 481 EQNLGSMTKKEIYAKLC-QPNFFDGGTKEVWLDRSSGQVCMFISSKSFKITGIDDRRYWN 657
Query: 120 WISMPDSR 127
+I +SR
Sbjct: 658 YIPTEESR 681
>TC207581
Length = 1562
Score = 79.0 bits (193), Expect = 6e-16
Identities = 42/124 (33%), Positives = 70/124 (55%), Gaps = 2/124 (1%)
Frame = +2
Query: 6 DLPEGCIAAIVSRTTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSPSLTNI 65
D+PE C+A ++ P D +L+ ++ FR AS +D +W LP + I+ + ++
Sbjct: 461 DIPESCVALVLMYLDPPDICKLARLNRAFRDASSADFIWESKLPLNYKFIVEKALKDVSV 640
Query: 66 PT--KKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSARSLAIVWGDDRRYWNWISM 123
K+D+Y L RP D+ K I L++++G C +S+++L I DDRRYW+ IS
Sbjct: 641 EQLGKRDIYARLC-RPNSFDNGTKEIWLDKRTGGVCLAISSQALRITGIDDRRYWSRIST 817
Query: 124 PDSR 127
+SR
Sbjct: 818 EESR 829
>AI440677
Length = 421
Score = 77.0 bits (188), Expect = 2e-15
Identities = 37/115 (32%), Positives = 60/115 (52%)
Frame = +1
Query: 6 DLPEGCIAAIVSRTTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSPSLTNI 65
D+PE C+A + TP + L+ ++ FR A+ +D+VW LP + ++ P +
Sbjct: 52 DIPENCVARVFLHLTPPEICNLARLNRAFRGAASADSVWQTKLPRNYQDLLDLMPPERHR 231
Query: 66 PTKKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSARSLAIVWGDDRRYWNW 120
K AL R V D K + L+R + + C +SA++++I DDRRYW W
Sbjct: 232 NLSKKDIFALLSRAVPFDDGNKGVWLDRVTRRVCMSISAKAMSITGIDDRRYWTW 396
>TC207017
Length = 905
Score = 77.0 bits (188), Expect = 2e-15
Identities = 44/124 (35%), Positives = 66/124 (52%), Gaps = 2/124 (1%)
Frame = +3
Query: 6 DLPEGCIAAIVSRTTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSPSLTNI 65
D+PE CI+++ P D +L+ ++ F AS +D VW LP + + NI
Sbjct: 186 DIPESCISSLFMNLDPPDICKLARVNRAFHRASSADFVWESKLPPSYKFLANKVLGEENI 365
Query: 66 PT--KKDLYLALSDRPVIIDHDKKSIQLERKSGKKCYMLSARSLAIVWGDDRRYWNWISM 123
T KK++Y L P D K + L++ SG+ C +S++SL I DDRRYWN+I
Sbjct: 366 ATMTKKEIYAKLC-LPNRFDGGTKEVWLDKCSGQVCLFMSSKSLKITGIDDRRYWNYIPT 542
Query: 124 PDSR 127
+SR
Sbjct: 543 EESR 554
>BE803206
Length = 404
Score = 71.2 bits (173), Expect = 1e-13
Identities = 40/72 (55%), Positives = 50/72 (68%), Gaps = 1/72 (1%)
Frame = +2
Query: 5 QDLPEGCIAAIVSRT-TPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSPSLT 63
QDLPEGC+A I+S TP D RLS+ SK F SA+D D VW++F+PSD SS IS L+
Sbjct: 23 QDLPEGCVAHILSYICTPEDIVRLSLVSKAFYSAADYDTVWDRFIPSDFSSTIS---PLS 193
Query: 64 NIPTKKDLYLAL 75
+ +KKDLY L
Sbjct: 194SSNSKKDLYFTL 229
>TC228791
Length = 584
Score = 60.8 bits (146), Expect = 2e-10
Identities = 33/106 (31%), Positives = 52/106 (48%), Gaps = 6/106 (5%)
Frame = +1
Query: 6 DLPEGCIAAIVSRTTPLDAGRLSVFSKTFRSASDSDAVWNQFLPSDISSIISHSPSLTNI 65
D+PE C+A + TP + L+ ++ FR A+ SD+VW LP + ++ P +
Sbjct: 265 DIPESCVACVFLHLTPPEICNLARLNRAFRGAASSDSVWEAKLPRNYQDLLDLVPPERHR 444
Query: 66 PTKKDLYLALSDRPVIIDHDKK------SIQLERKSGKKCYMLSAR 105
K AL RP+ DH K + L+R +GK C +SA+
Sbjct: 445 SLSKKDIFALLSRPLPFDHGHKVGAALQEVWLDRVTGKVCMSISAK 582
>TC228248 weakly similar to UP|Q8LBR0 (Q8LBR0) Phloem-specific lectin
PP2-like protein, partial (28%)
Length = 519
Score = 49.3 bits (116), Expect = 5e-07
Identities = 19/36 (52%), Positives = 27/36 (74%)
Frame = +3
Query: 92 ERKSGKKCYMLSARSLAIVWGDDRRYWNWISMPDSR 127
++ SGKK Y+LSAR L+I W +D YW+W +P+SR
Sbjct: 15 DKSSGKKSYILSARQLSITWSNDPLYWSWRPVPESR 122
>TC221119 similar to UP|Q9FLU7 (Q9FLU7) Phloem-specific lectin-like protein,
partial (6%)
Length = 971
Score = 48.1 bits (113), Expect = 1e-06
Identities = 17/35 (48%), Positives = 26/35 (73%)
Frame = +3
Query: 86 KKSIQLERKSGKKCYMLSARSLAIVWGDDRRYWNW 120
K+ + +E+ +GK CY+LSAR L+I WG+ YW+W
Sbjct: 153 KQMLSIEKTTGKICYLLSARQLSITWGNSTLYWSW 257
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.132 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 5,863,512
Number of Sequences: 63676
Number of extensions: 68658
Number of successful extensions: 450
Number of sequences better than 10.0: 56
Number of HSP's better than 10.0 without gapping: 428
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 430
length of query: 130
length of database: 12,639,632
effective HSP length: 86
effective length of query: 44
effective length of database: 7,163,496
effective search space: 315193824
effective search space used: 315193824
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 53 (25.0 bits)
Medicago: description of AC146559.3


