

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146557.1 + phase: 0 /pseudo
(409 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
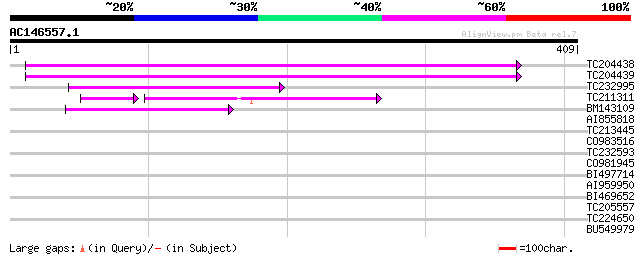
Score E
Sequences producing significant alignments: (bits) Value
TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, co... 95 6e-20
TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete 76 3e-14
TC232995 69 5e-12
TC211311 weakly similar to UP|O24587 (O24587) Pol protein, parti... 54 1e-09
BM143109 51 1e-06
AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza ... 39 0.006
TC213445 37 0.016
CO983516 34 0.14
TC232593 weakly similar to UP|Q9XG91 (Q9XG91) Tpv2-1c protein (F... 32 0.67
CO981945 31 1.2
BI497714 homologue to SP|P13890|POLS Structural polyprotein (P13... 30 1.5
AI959950 30 1.5
BI469652 weakly similar to GP|18149115|dbj| reverse transcriptas... 29 4.4
TC205557 similar to UP|PMM_ARATH (O80840) Probable phosphomannom... 28 5.7
TC224650 homologue to UP|O49081 (O49081) Clp-like energy-depende... 28 9.7
BU549979 28 9.7
>TC204438 homologue to UP|Q84VH6 (Q84VH6) Gag-pol polyprotein, complete
Length = 4734
Score = 94.7 bits (234), Expect = 6e-20
Identities = 99/358 (27%), Positives = 174/358 (47%)
Frame = +3
Query: 12 NGCQKCLS*WCH*RRSRCQTTSRV*GS*AS*PCL*T*EITIWLETSSQSLV**TK*FLN* 71
+GC++ +S W RS C + S +S C+ E ++W+E SS+SLV*
Sbjct: 3525 DGCEERVSEWIPE*RSLCGAAKGICRSNSSRSCIQAQEGSLWIEASSKSLV*KANRVPYS 3704
Query: 72 K*F*KRTS*HNTLQKDS*ERHFDCANIC**YNIWFY*CISLQRIF*VNAG*I*NEYDGRI 131
*+ +* ++L + + D +IC**+ +W +A *I*+E R
Sbjct: 3705 ARV*EGRN*QDSLCQTRC*KLDDSTDIC**HCVWRDVE*DASTFCPTDAI*I*DESCWRA 3884
Query: 132 EVLSWNSNQPK*RRSICSSNKIYKGASEEVQARRL*SDEHSNASNLHLKQRRYWNSSRPE 191
++ S ++ R I + ++ K +EV + +++ +L +R W+ +
Sbjct: 3885 DLFSGTPSEADGRLHIPLTKQVCKEHCQEVWDGKCQP*KNTCTYSLEAVKR*SWHQC*SK 4064
Query: 192 AIQRYDWFSVIPHCI*T*YLIQCMLVCKISIRS*RISFNCC*ENLQVFERNN*SWTPL*E 251
++Q++DW I + T*+ + +CKIS +S* S EN ++ + + * W +
Sbjct: 4065 SVQKHDWELTIFNS*QT*HHLCSRCLCKISSQS*DKSLESSKENSEICKWHQ*LWDYVLS 4244
Query: 252 IPRLQVD*IL*C*LCW*QD*KKINQWKLSIPGRKSDILGKQKTSNNCYVYSRSRIHFSCK 311
+ R +L*C*L W *+K + W + + G +S + +Q+ +Y SR++ S K
Sbjct: 4245 LFRFNAGWVL*C*LGWKCR*QKKHFWWMFLFGNQSYFMVQQEAELCVPIYC*SRVYCSRK 4424
Query: 312 LLYTTTLDETSVGRLSD*C*QYSHFL**Y*CYLFVKESNSTFKSQAY*DQTPFY*RLC 369
L+TT+LDE + + L* + CY + +S ST ++QA+* T Y*R C
Sbjct: 4425 QLFTTSLDEADAEGVQCRTRCHDIVL*QHECY*YF*KSCSTQQNQAH*H*TSLY*RSC 4598
>TC204439 UP|Q84VI4 (Q84VI4) Gag-pol polyprotein, complete
Length = 4731
Score = 75.9 bits (185), Expect = 3e-14
Identities = 93/358 (25%), Positives = 168/358 (45%)
Frame = +3
Query: 12 NGCQKCLS*WCH*RRSRCQTTSRV*GS*AS*PCL*T*EITIWLETSSQSLV**TK*FLN* 71
+GC++ +S W RS C + +S C+ E ++W+E SS+SLV*
Sbjct: 3522 DGCEERISEWIPE*RSLCGAAKGICRPDSSRSCIQAQEGSLWIEASSKSLV*KANRVPYS 3701
Query: 72 K*F*KRTS*HNTLQKDS*ERHFDCANIC**YNIWFY*CISLQRIF*VNAG*I*NEYDGRI 131
*+ +* + L + + DC +IC**+ +W +A *I*+E R
Sbjct: 3702 ARV*EGRN*QDPLCQTRC*KLDDCTDIC**HCVWRDVE*DASTFCSTDAI*I*DESCWRA 3881
Query: 132 EVLSWNSNQPK*RRSICSSNKIYKGASEEVQARRL*SDEHSNASNLHLKQRRYWNSSRPE 191
++ S S++ I + ++ K +EV S + + +L + + +
Sbjct: 3882 DLFSGTSSEADGGLHIPLTKQVCKEHCQEVWDGECQS*KDTCTYSLEAVKG*SRHQC*SK 4061
Query: 192 AIQRYDWFSVIPHCI*T*YLIQCMLVCKISIRS*RISFNCC*ENLQVFERNN*SWTPL*E 251
++Q++D I + T + + +CKIS +S S + EN ++ + + * W +
Sbjct: 4062 SVQKHDRELTIFNS*QTRHHLCSRCLCKISSQSQDKSLDSSKENSEICKWH**LWDYVLS 4241
Query: 252 IPRLQVD*IL*C*LCW*QD*KKINQWKLSIPGRKSDILGKQKTSNNCYVYSRSRIHFSCK 311
+ + +L*C*L W *+K + W + + G++ + +Q+ +YSRSR++ S K
Sbjct: 4242 LFKSNAGWVL*C*LGWKCR*QKKHFWWMLLFGKQPYFMVQQEAELCVPIYSRSRVYCSRK 4421
Query: 312 LLYTTTLDETSVGRLSD*C*QYSHFL**Y*CYLFVKESNSTFKSQAY*DQTPFY*RLC 369
L+T +LDE + + L* + CY + +S ST ++QA+* T Y R C
Sbjct: 4422 QLFTASLDEADAEGVQCRTRCHDIVL*QHECY*YF*KSCSTQQNQAH*H*TSLYQRSC 4595
>TC232995
Length = 1009
Score = 68.6 bits (166), Expect = 5e-12
Identities = 56/156 (35%), Positives = 85/156 (53%)
Frame = +1
Query: 43 PCL*T*EITIWLETSSQSLV**TK*FLN*K*F*KRTS*HNTLQKDS*ERHFDCANIC**Y 102
PCL* + ++W ETS +V* K*F +*K +R S ++ + K+ +F +NIC**Y
Sbjct: 43 PCL*ITKGSLWFETSP*GMV*TIK*FSS*KRILQR*SGYHIIHKEKA**YFVGSNIC**Y 222
Query: 103 NIWFY*CISLQRIF*VNAG*I*NEYDGRIEVLSWNSNQPK*RRSICSSNKIYKGASEEVQ 162
N W +* +Q +F A *I*N DGR +VLS +NQ R I S +I +G +++
Sbjct: 223 NFWIH**FIVQGVFP*YAK*I*NVNDGRTKVLSGITNQANSIRYIHQSIQILQGIDQKIW 402
Query: 163 ARRL*SDEHSNASNLHLKQRRYWNSSRPEAIQRYDW 198
+ +++ L L+ R W+ R + I R W
Sbjct: 403 DG*CKTHVYTDEH*LLLR*R*IWSVYRHKTISRCYW 510
>TC211311 weakly similar to UP|O24587 (O24587) Pol protein, partial (15%)
Length = 1213
Score = 53.9 bits (128), Expect(2) = 1e-09
Identities = 61/173 (35%), Positives = 89/173 (51%), Gaps = 2/173 (1%)
Frame = +2
Query: 98 IC**YNIWFY*CISLQRIF*VNAG*I*NEYDGRIEVLSWNSNQPK*RRSICSSNKIYKGA 157
+C**+N+W +Q +F*VN G I*NEY+ +V SN + S +IYK
Sbjct: 503 LC**HNLWCNLKKDVQGVF*VNEGWI*NEYER*AKVPPRTSNHSESLWDFYPSREIYKVP 682
Query: 158 SEEVQARRL*SDEHSN--ASNLHLKQRRYWNSSRPEAIQRYDWFSVIPHCI*T*YLIQCM 215
S++VQ *S + N AS + Q S + + YD F +I + *T Y + +
Sbjct: 683 SKKVQNG--*SQTYGNPYASFHNH*QG*ER*SYFIKGV*WYD*FFIIFNF**TRYCVCRL 856
Query: 216 LVCKISIRS*RISFNCC*ENLQVFERNN*SWTPL*EIPRLQVD*IL*C*LCW* 268
+CKIS+ S S C *++L++ N *S + +*E + IL*C CW*
Sbjct: 857 PLCKISVLSKNFSCYCS*KDLKISCWNY*SLSMV*EKV*V*SFRIL*CLFCW* 1015
Score = 26.6 bits (57), Expect(2) = 1e-09
Identities = 17/42 (40%), Positives = 24/42 (56%)
Frame = +1
Query: 52 IWLETSSQSLV**TK*FLN*K*F*KRTS*HNTLQKDS*ERHF 93
+W ETSS+SLV* K + K +R + T+QK S + F
Sbjct: 364 LWFETSSKSLV*KAKFISSFKWIHQRNNGPRTIQKGSKRKPF 489
>BM143109
Length = 415
Score = 50.8 bits (120), Expect = 1e-06
Identities = 45/121 (37%), Positives = 63/121 (51%)
Frame = +3
Query: 41 S*PCL*T*EITIWLETSSQSLV**TK*FLN*K*F*KRTS*HNTLQKDS*ERHFDCANIC* 100
+* CL*T + IW++TS LV* +* + F KR *+ E + +IC*
Sbjct: 33 A*SCL*TEKGFIWIKTSP*GLV*TFE*ISFRQGFFKR*G*Y*PFYLKEIE*YTLSTDIC* 212
Query: 101 *YNIWFY*CISLQRIF*VNAG*I*NEYDGRIEVLSWNSNQPK*RRSICSSNKIYKGASEE 160
*Y WF * SLQ+IF A *I*N D +++ SW +NQ +I S KI + +
Sbjct: 213 *YYFWFN**FSLQKIFSRYAK*I*NVNDA*VKLFSWTTNQANKEWNIYQSIKILQRPDSQ 392
Query: 161 V 161
+
Sbjct: 393 I 395
>AI855818 weakly similar to GP|21741393|e OSJNBb0051N19.6 {Oryza sativa
(japonica cultivar-group)}, partial (10%)
Length = 463
Score = 38.5 bits (88), Expect = 0.006
Identities = 25/57 (43%), Positives = 33/57 (57%)
Frame = -2
Query: 12 NGCQKCLS*WCH*RRSRCQTTSRV*GS*AS*PCL*T*EITIWLETSSQSLV**TK*F 68
NGC KC S W + RRS C TT R+* + CL* + ++W +TS +V* K F
Sbjct: 183 NGC*KCFSKWFNSRRSIC*TTPRL*NPG*TNSCL*IAKGSLWFKTSP*GVV*TYKQF 13
>TC213445
Length = 705
Score = 37.0 bits (84), Expect = 0.016
Identities = 25/64 (39%), Positives = 38/64 (59%)
Frame = +3
Query: 298 CYVYSRSRIHFSCKLLYTTTLDETSVGRLSD*C*QYSHFL**Y*CYLFVKESNSTFKSQA 357
C + RSRI+F KLL T LDET+ L * Y++ +* Y C +++S S ++A
Sbjct: 486 CLINCRSRIYFC*KLLCTNLLDETTTF*LWFET*SYTYPM*QYKCN*SIQKSYSVL*NKA 665
Query: 358 Y*DQ 361
Y*++
Sbjct: 666 Y*NK 677
Score = 32.0 bits (71), Expect = 0.52
Identities = 16/32 (50%), Positives = 22/32 (68%)
Frame = +1
Query: 208 T*YLIQCMLVCKISIRS*RISFNCC*ENLQVF 239
T Y + C+ VCKIS +S RIS C *+N ++F
Sbjct: 232 TSYNV*CLYVCKISSKSQRISPKCH*KNNEIF 327
>CO983516
Length = 724
Score = 33.9 bits (76), Expect = 0.14
Identities = 28/94 (29%), Positives = 48/94 (50%)
Frame = +1
Query: 12 NGCQKCLS*WCH*RRSRCQTTSRV*GS*AS*PCL*T*EITIWLETSSQSLV**TK*FLN* 71
+GC++ +S W RS C + S +S C+ E ++W+E SS+SLV*
Sbjct: 442 DGCEERVSEWIPE*RSLCGAAKGIYRSNSSRSCIQAQEGSLWIEASSKSLV*KANRVTYS 621
Query: 72 K*F*KRTS*HNTLQKDS*ERHFDCANIC**YNIW 105
*+ +* ++L + + D +IC**+ +W
Sbjct: 622 ARV*EGRN*QDSLCQTRC*KLDDSTDIC**HCVW 723
>TC232593 weakly similar to UP|Q9XG91 (Q9XG91) Tpv2-1c protein (Fragment),
partial (16%)
Length = 562
Score = 31.6 bits (70), Expect = 0.67
Identities = 22/74 (29%), Positives = 35/74 (46%)
Frame = +3
Query: 132 EVLSWNSNQPK*RRSICSSNKIYKGASEEVQARRL*SDEHSNASNLHLKQRRYWNSSRPE 191
++ SWN +Q K +S+ S +I K EEV + H+N S ++Q R +
Sbjct: 309 DLFSWN*DQAKSEQSVNLSKEICKRNFEEVSNGGMQIC*HTNESKGEVQQGRRC**N**R 488
Query: 192 AIQRYDWFSVIPHC 205
+ DW S + HC
Sbjct: 489 IL*ELDWMSNVSHC 530
>CO981945
Length = 584
Score = 30.8 bits (68), Expect = 1.2
Identities = 13/34 (38%), Positives = 22/34 (64%)
Frame = -3
Query: 273 KINQWKLSIPGRKSDILGKQKTSNNCYVYSRSRI 306
KIN W + +P +SD++ ++ +C +Y RSRI
Sbjct: 441 KINIWGMHLP*PESDLMVVKEADLDCLLYCRSRI 340
>BI497714 homologue to SP|P13890|POLS Structural polyprotein (P130)
[Contains: Coat protein C (EC 3.4.21.-) (Capsid protein
C); Spike, partial (1%)
Length = 433
Score = 30.4 bits (67), Expect = 1.5
Identities = 14/38 (36%), Positives = 22/38 (57%)
Frame = +2
Query: 152 KIYKGASEEVQARRL*SDEHSNASNLHLKQRRYWNSSR 189
KI + A++EVQ + DE N NL ++ +RY N +
Sbjct: 98 KIQRRATDEVQENKKIIDERENHKNLSVQDKRYCNMEK 211
>AI959950
Length = 466
Score = 30.4 bits (67), Expect = 1.5
Identities = 16/36 (44%), Positives = 22/36 (60%)
Frame = -3
Query: 12 NGCQKCLS*WCH*RRSRCQTTSRV*GS*AS*PCL*T 47
NGC+KC+S W + + S C TT+ +* S C *T
Sbjct: 116 NGCKKCISKWLNPKGSLC*TTAWI*K*NPSSTCF*T 9
>BI469652 weakly similar to GP|18149115|dbj| reverse transcriptase {Silene
noctiflora}, partial (60%)
Length = 427
Score = 28.9 bits (63), Expect = 4.4
Identities = 21/46 (45%), Positives = 27/46 (58%)
Frame = +2
Query: 23 H*RRSRCQTTSRV*GS*AS*PCL*T*EITIWLETSSQSLV**TK*F 68
+*R S CQTTS++ S P T + IW ET + LV**T+ F
Sbjct: 290 Y*RGSVCQTTSKLCRPYTSRPYFQTSKGFIWSETGTLCLV**TELF 427
>TC205557 similar to UP|PMM_ARATH (O80840) Probable phosphomannomutase (PMM) ,
complete
Length = 1699
Score = 28.5 bits (62), Expect = 5.7
Identities = 17/55 (30%), Positives = 31/55 (55%), Gaps = 1/55 (1%)
Frame = -1
Query: 147 ICSSNKIYKGASEEVQARRL*SDEHSNASNLHLKQ-RRYWNSSRPEAIQRYDWFS 200
+CSSNK+ G+ + + + + + + LHL+Q R W +SRP+ + +FS
Sbjct: 1132 LCSSNKVV-GSRDPIALQWRCTHHYFHLYLLHLQQPSRLWKNSRPQEFFFFFFFS 971
>TC224650 homologue to UP|O49081 (O49081) Clp-like energy-dependent protease,
partial (72%)
Length = 1067
Score = 27.7 bits (60), Expect = 9.7
Identities = 11/40 (27%), Positives = 23/40 (57%)
Frame = -3
Query: 280 SIPGRKSDILGKQKTSNNCYVYSRSRIHFSCKLLYTTTLD 319
+IP + + + S++C+ Y R + F+C + T+T+D
Sbjct: 333 TIPEQNRCEIKRSSRSHHCWSYRRIPLGFNCTITCTSTMD 214
>BU549979
Length = 615
Score = 27.7 bits (60), Expect = 9.7
Identities = 15/37 (40%), Positives = 19/37 (50%)
Frame = -3
Query: 265 LCW*QD*KKINQWKLSIPGRKSDILGKQKTSNNCYVY 301
LCW + KIN W R S IL K + + NCY +
Sbjct: 469 LCWLR*FTKINIWLYFYVSRWSCILEKFQANLNCYFH 359
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.382 0.171 0.721
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,968,020
Number of Sequences: 63676
Number of extensions: 260862
Number of successful extensions: 3854
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 1584
Number of HSP's successfully gapped in prelim test: 146
Number of HSP's that attempted gapping in prelim test: 2230
Number of HSP's gapped (non-prelim): 1797
length of query: 409
length of database: 12,639,632
effective HSP length: 100
effective length of query: 309
effective length of database: 6,272,032
effective search space: 1938057888
effective search space used: 1938057888
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146557.1


