

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146553.2 + phase: 0 /pseudo
(654 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
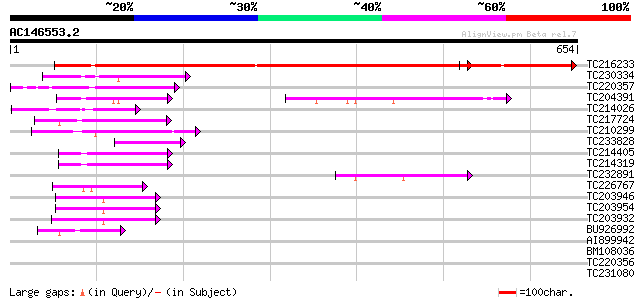
TC216233 GB|CAA50573.1|402753|GMFUSA translation elongation fact... 382 e-106
TC230334 similar to UP|Q8RWP3 (Q8RWP3) GTP-binding protein LepA-... 91 2e-18
TC220357 homologue to UP|Q6SYB1 (Q6SYB1) GTP-binding protein Typ... 91 2e-18
TC204391 homologue to UP|Q9ASR1 (Q9ASR1) At1g56070/T6H22_13 (Elo... 76 4e-14
TC214026 similar to UP|Q6G0R7 (Q6G0R7) GTP-binding protein typA,... 74 2e-13
TC217724 homologue to UP|EFTM_ARATH (Q9ZT91) Elongation factor T... 63 4e-10
TC210299 homologue to UP|EFT2_SOYBN (P46280) Elongation factor T... 57 3e-08
TC233828 similar to PIR|T01917|T01917 translation initiation fac... 55 8e-08
TC214405 homologue to UP|EFT1_SOYBN (Q43467) Elongation factor T... 54 3e-07
TC214319 homologue to UP|EFT1_SOYBN (Q43467) Elongation factor T... 54 3e-07
TC232891 similar to UP|Q9LNC5 (Q9LNC5) F9P14.8 protein, partial ... 49 9e-06
TC226767 similar to GB|AAP68328.1|31711944|BT008889 At1g18070 {A... 44 2e-04
TC203946 homologue to UP|Q9LN13 (Q9LN13) T6D22.2, partial (47%) 44 2e-04
TC203954 homologue to UP|EF1A_LYCES (P17786) Elongation factor 1... 44 2e-04
TC203932 UP|EF1A_SOYBN (P25698) Elongation factor 1-alpha (EF-1-... 43 5e-04
BU926992 homologue to GP|11181616|gb| translational elongation f... 42 7e-04
AI899942 similar to PIR|S17434|S17 translation elongation factor... 35 0.14
BM108036 34 0.18
TC220356 similar to UP|Q6SYB1 (Q6SYB1) GTP-binding protein TypA,... 34 0.18
TC231080 UP|Q9FQD5 (Q9FQD5) Glutathione S-transferase GST 23 , ... 34 0.18
>TC216233 GB|CAA50573.1|402753|GMFUSA translation elongation factor EF-G
{Glycine max;} , complete
Length = 2424
Score = 382 bits (980), Expect = e-106
Identities = 212/487 (43%), Positives = 302/487 (61%), Gaps = 4/487 (0%)
Frame = +1
Query: 52 MEKVRNIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITI 111
++ RNIGI AHID+GKTT TE IL+YTG+ + + EV MD+ E GITI
Sbjct: 37 LKDYRNIGIMAHIDAGKTTTTERILYYTGRNYKIGEVHEGTAT---MDWMEQEQERGITI 207
Query: 112 KSAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSITVDRQM 171
SAAT W +I IIDTPGHVDFT+EVERALRVLDGA+ + SV GV+ QS TV RQ
Sbjct: 208 TSAATTTFWNKHRINIIDTPGHVDFTLEVERALRVLDGAICLFDSVAGVEPQSETVWRQA 387
Query: 172 RRYQVPRIAFINKLDRPGADPWKVITQARSKLRHHCAALQVPIGLESDFKGVVDLVKLKA 231
+Y VPRI F+NK+DR GA+ ++ + L +Q+PIG E +FKGV+DLV+ KA
Sbjct: 388 DKYGVPRICFVNKMDRLGANFYRTRDMIVTNLGAKPLVIQLPIGSEDNFKGVIDLVRNKA 567
Query: 232 YCFDG-QYGQNVVVGEVPADMEALVAEKRRELIETVSEVDDVLAEAFLSDDENISAADLE 290
+ G + G + ++P D++ E R ++IE + E DD E +L E ++
Sbjct: 568 IVWSGEELGAKFDIVDIPEDLQEQAQEYRAQMIENIVEFDDQAMENYLEGIEP-DEETIK 744
Query: 291 GAIRRATIARKFIPVFMGSAVKNTGVQPLLDGVVSYLPCPIEV-SNYALDQSKNEEKVQL 349
IR+ TI+ F+PV GSA KN GVQPLLD VV YLP P+++ + D EE ++
Sbjct: 745 KLIRKGTISASFVPVMCGSAFKNKGVQPLLDAVVDYLPSPLDLPAMKGSDPENPEETIER 924
Query: 350 TGSPDGPLVALAFKLEQTKF-GQLTYLRVYEGVIRKGDFIVNVSTGKKIKVPRLVQMHSN 408
S D P LAFK+ F G LT++RVY G + G +++N + GKK ++ RL++MH+N
Sbjct: 925 VASDDEPFAGLAFKIMSDPFVGSLTFVRVYAGKLSAGSYVLNANKGKKERIGRLLEMHAN 1104
Query: 409 EMNDIEEAHAGQIVAVFGV-DCASSDTFTDGSVKYTMTSMNVPEPVMSLAVQPVSKDSGG 467
D++ A AG I+A+ G+ D + +T D + M+ P+PV+ +A++P +K
Sbjct: 1105SREDVKVALAGDIIALAGLKDTITGETLCDPDNPIVLERMDFPDPVIKVAIEPKTKADVD 1284
Query: 468 KFSKALNRFQREDPTFRVSLDPESGQTIISGMGELHLDIYVKRIKMEYGVDATVGKPRVN 527
K + L + +EDP+F S D E QT+I GMGELHL+I V R+K E+ V+A VG P+VN
Sbjct: 1285KMATGLIKLAQEDPSFHFSRDEEINQTVIEGMGELHLEIIVDRLKREFKVEANVGAPQVN 1464
Query: 528 FRETVTQ 534
+RE++++
Sbjct: 1465YRESISK 1485
Score = 114 bits (285), Expect = 1e-25
Identities = 59/134 (44%), Positives = 85/134 (63%)
Frame = +2
Query: 520 TVGKPRVNFRETVTQRADFDYLHKKQSGGQGQYGRVIGYIEPLPAGSGTKFEFDNMLVGQ 579
T+ P+ + + A+ Y+HKKQSGGQGQ+ + EP+ GSG +EF + + G
Sbjct: 1442 TLVPPK*TTGKAFPKSAEVKYVHKKQSGGQGQFADITVRFEPMDPGSG--YEFKSEIKGG 1615
Query: 580 AIPSNFFPAIEKGFKEAANSGALIGHPVQNLRVVLTDGAAHDVDSSELAFKLASIYAFRE 639
A+P + P + KG +E ++G L G PV ++R VLTDG+ HDVDSS LAF+LA+ AFRE
Sbjct: 1616 AVPKEYIPGVMKGLEECMSNGVLAGFPVVDVRAVLTDGSYHDVDSSVLAFQLAARGAFRE 1795
Query: 640 CYTASRPVILEPVM 653
+ P +LEP+M
Sbjct: 1796 GIRKAGPRMLEPIM 1837
>TC230334 similar to UP|Q8RWP3 (Q8RWP3) GTP-binding protein LepA-like
protein, partial (27%)
Length = 820
Score = 90.5 bits (223), Expect = 2e-18
Identities = 62/175 (35%), Positives = 91/175 (51%), Gaps = 4/175 (2%)
Frame = +2
Query: 38 TAATIDKDPWWKESMEKVRNIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPK 97
+AA +D K + +RN I AHID GK+TL + +L TG +H + KD
Sbjct: 263 SAAKAGEDRLSKVPVRNIRNFCIIAHIDHGKSTLADKLLQVTGTVH---QREMKDQFLDN 433
Query: 98 MDFKPLEIIMGITIKSAATYCNWKGSK----ITIIDTPGHVDFTIEVERALRVLDGAVLV 153
MD LE GITIK A + + +IDTPGHVDF+ EV R+L +GA+LV
Sbjct: 434 MD---LERERGITIKLQAARMRYVFENEPYCLNLIDTPGHVDFSYEVSRSLAACEGALLV 604
Query: 154 LCSVGGVQCQSITVDRQMRRYQVPRIAFINKLDRPGADPWKVITQARSKLRHHCA 208
+ + GV+ Q++ + I +NK+D PGA+P +VI + + C+
Sbjct: 605 VDASQGVEAQTLANVYLALENNLEIIPVLNKIDLPGAEPDRVIKEIEEIVGLDCS 769
>TC220357 homologue to UP|Q6SYB1 (Q6SYB1) GTP-binding protein TypA, partial
(25%)
Length = 668
Score = 90.5 bits (223), Expect = 2e-18
Identities = 72/197 (36%), Positives = 101/197 (50%), Gaps = 2/197 (1%)
Frame = +2
Query: 2 QRFGYALCSSTLTTEGSLIAGTFHIRHFSAGNVARATAATIDKDPWWKESMEK--VRNIG 59
Q G L SS+ T +A F R +V+ AT +K ++ + + VRNI
Sbjct: 89 QLLGLPLSSSSSLT----VAKRFSCRPIKC-SVSEATEPKTEKK---RQLLRRGDVRNIA 244
Query: 60 ISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITIKSAATYCN 119
I AH+D GKTTL + +L T V+ + MD LE GITI S T
Sbjct: 245 IVAHVDHGKTTLVDAMLKQTKVFRDNQFVQERI-----MDSNDLERERGITILSKNTSVT 409
Query: 120 WKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSITVDRQMRRYQVPRI 179
+K +KI IIDTPGH DF EVER L +++G +LV+ SV G Q+ V ++ + +
Sbjct: 410 YKDAKINIIDTPGHSDFGGEVERILNMVEGILLVVDSVEGPMPQTRFVLKKALEFGHSVV 589
Query: 180 AFINKLDRPGADPWKVI 196
+NK+DRP A P V+
Sbjct: 590 VVVNKIDRPSARPEYVV 640
>TC204391 homologue to UP|Q9ASR1 (Q9ASR1) At1g56070/T6H22_13 (Elongation
factor EF-2), complete
Length = 2941
Score = 76.3 bits (186), Expect = 4e-14
Identities = 53/149 (35%), Positives = 79/149 (52%), Gaps = 16/149 (10%)
Frame = +1
Query: 55 VRNIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITIKSA 114
+RN+ + AH+D GK+TLT+ ++ G I + G D + E GITIKS
Sbjct: 175 IRNMSVIAHVDHGKSTLTDSLVAAAGII-----AQEVAGDVRMTDTRADEAERGITIKST 339
Query: 115 ATYC----------NWKGSK------ITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVG 158
++KG + I +ID+PGHVDF+ EV ALR+ DGA++V+ V
Sbjct: 340 GISLYYEMTDEALKSFKGERNGNEYLINLIDSPGHVDFSSEVTAALRITDGALVVVDCVE 519
Query: 159 GVQCQSITVDRQMRRYQVPRIAFINKLDR 187
GV Q+ TV RQ ++ + +NK+DR
Sbjct: 520 GVCVQTETVLRQALGERIRPVLTVNKMDR 606
Score = 62.4 bits (150), Expect = 6e-10
Identities = 71/284 (25%), Positives = 122/284 (42%), Gaps = 24/284 (8%)
Frame = +1
Query: 319 LLDGVVSYLPCPIEVSNYALDQSKNEEKVQLTGS------PDGPLVALAFKL--EQTKFG 370
LL+ ++ +LP P Y ++ S P+GPL+ K+ K
Sbjct: 1114 LLEMMIFHLPSPSTAQKYRVENLYEGPLDDQYASAIRNCDPEGPLMLYVSKMIPASDKGR 1293
Query: 371 QLTYLRVYEGVIRKGDFI----VNVSTGKKI-----KVPRLVQMHSNEMNDIEEAHAGQI 421
+ RV+ G + G + N G+K V R V +E+ G
Sbjct: 1294 FFAFGRVFSGRVSTGLKVRIMGPNYVPGEKKDLYVKSVQRTVIWMGKRQETVEDVPCGNT 1473
Query: 422 VAVFGVDCASSDTFTDGSVK----YTMTSMNVP-EPVMSLAVQPVSKDSGGKFSKALNRF 476
VA+ G+D + T + K + + +M PV+ +AVQ K + L R
Sbjct: 1474 VAMVGLDQFITKNATLTNEKEVDAHPIRAMKFSVSPVVRVAVQCKVASDLPKLVEGLKRL 1653
Query: 477 QREDPTFRVSLDPESGQTIISGMGELHLDIYVKRIKMEY--GVDATVGKPRVNFRETVTQ 534
+ DP +++ ESG+ I++G GELHL+I +K ++ ++ G + P V+FRETV +
Sbjct: 1654 AKSDPMVVCTIE-ESGEHIVAGAGELHLEICLKDLQDDFMGGAEIIKSDPVVSFRETVLE 1830
Query: 535 RADFDYLHKKQSGGQGQYGRVIGYIEPLPAGSGTKFEFDNMLVG 578
R+ + K + ++ R+ Y+E P G D+ +G
Sbjct: 1831 RSCRTVMSKSPN----KHNRL--YMEARPLEEGLAEAIDDGKIG 1944
>TC214026 similar to UP|Q6G0R7 (Q6G0R7) GTP-binding protein typA, partial
(14%)
Length = 501
Score = 74.3 bits (181), Expect = 2e-13
Identities = 52/151 (34%), Positives = 76/151 (49%), Gaps = 2/151 (1%)
Frame = +1
Query: 3 RFGYALCSSTLTTEGSLIAGTFHIRHFSAGNVARATAATIDK--DPWWKESMEKVRNIGI 60
R ++ SS+ + L F R F+A AAT DP ++RN+ +
Sbjct: 94 RKSFSSSSSSFSPSPPLSHSRFFSRAFAAAPATAPAAATTASSLDP------SRLRNLAV 255
Query: 61 SAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITIKSAATYCNW 120
AH+D GKTTL + +L G L +E MD LE GITI S T +W
Sbjct: 256 IAHVDHGKTTLMDRLLRQCGA-DLPHE--------RAMDSISLERERGITISSKVTSVSW 408
Query: 121 KGSKITIIDTPGHVDFTIEVERALRVLDGAV 151
K +++ ++DTPGH DF EVER + +++GA+
Sbjct: 409 KENELNMVDTPGHADFGGEVERVVGMVEGAI 501
>TC217724 homologue to UP|EFTM_ARATH (Q9ZT91) Elongation factor Tu,
mitochondrial precursor, partial (87%)
Length = 1679
Score = 63.2 bits (152), Expect = 4e-10
Identities = 46/164 (28%), Positives = 75/164 (45%), Gaps = 6/164 (3%)
Frame = +2
Query: 29 FSAGNVARATAATIDKDPWWKESMEKVR-----NIGISAHIDSGKTTLTEWILFYTGKIH 83
F+ + + +++++ +PWW+ R N+G H+D GKTTLT I
Sbjct: 209 FTNDSTSSSSSSSSSSNPWWRSMATFTRTKPHVNVGTIGHVDHGKTTLTAAITRV----- 373
Query: 84 LMYEVRSKDGMGPKMDFKPLEIIMGITIKSAATYCNWKGSKITIIDTPGHVDFTIEVERA 143
L EV++K ++D P E GITI +A +D PGH D+ +
Sbjct: 374 LADEVKAKAVAFDEIDKAPEEKKRGITIATAHVEYETAKRHYAHVDCPGHADYVKNMITG 553
Query: 144 LRVLDGAVLVLCSVGGVQCQSITVDRQMRRYQVPR-IAFINKLD 186
+DG +LV+ + G Q+ R+ VP + F+NK+D
Sbjct: 554 AAQMDGGILVVSAPDGPMPQTKEHILLARQVGVPSLVCFLNKVD 685
>TC210299 homologue to UP|EFT2_SOYBN (P46280) Elongation factor Tu,
chloroplast precursor (EF-Tu), complete
Length = 1615
Score = 57.0 bits (136), Expect = 3e-08
Identities = 55/201 (27%), Positives = 83/201 (40%), Gaps = 6/201 (2%)
Frame = +1
Query: 26 IRHFSAGNVA-RATAATIDKDPWWKESMEKVRNIGISAHIDSGKTTLTEWILFYTGKIHL 84
I H +A N R + T+ E + NIG H+D GKTTLT L
Sbjct: 154 ILHLAAANTTTRRRSFTVRAARGKFERKKPHVNIGTIGHVDHGKTTLTA---------AL 306
Query: 85 MYEVRSKDGMGPK----MDFKPLEIIMGITIKSAATYCNWKGSKITIIDTPGHVDFTIEV 140
+ S PK +D P E GITI +A + +D PGH D+ +
Sbjct: 307 TMALASLGNSAPKKYDEIDAAPEERARGITINTATVEYETENRHYAHVDCPGHADYVKNM 486
Query: 141 ERALRVLDGAVLVLCSVGGVQCQSITVDRQMRRYQVPRI-AFINKLDRPGADPWKVITQA 199
+DGA+LV+ G Q+ ++ VP I F+NK D+ D +++
Sbjct: 487 ITGAAQMDGAILVVSGADGPMPQTKEHILLAKQVGVPNIVVFLNKQDQ--VDDEELLQLV 660
Query: 200 RSKLRHHCAALQVPIGLESDF 220
++R ++VP SD+
Sbjct: 661 ELEVRESSFQVRVPRRRCSDY 723
>TC233828 similar to PIR|T01917|T01917 translation initiation factor IF-2
homolog F2P3.9 - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (23%)
Length = 459
Score = 55.5 bits (132), Expect = 8e-08
Identities = 31/81 (38%), Positives = 43/81 (52%)
Frame = +2
Query: 122 GSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSITVDRQMRRYQVPRIAF 181
G+ IT +DTPGH F+ R V D VLV+ + GV Q++ + VP +
Sbjct: 23 GASITFLDTPGHAAFSAMRARGAAVTDIVVLVVAADDGVMPQTLEAMSHAKAANVPIVVA 202
Query: 182 INKLDRPGADPWKVITQARSK 202
INK D+PGA+ KV Q S+
Sbjct: 203 INKCDKPGANSEKVKMQLASE 265
>TC214405 homologue to UP|EFT1_SOYBN (Q43467) Elongation factor Tu,
chloroplast precursor (EF-Tu), complete
Length = 2050
Score = 53.5 bits (127), Expect = 3e-07
Identities = 38/132 (28%), Positives = 57/132 (42%), Gaps = 1/132 (0%)
Frame = +3
Query: 57 NIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITIKSAAT 116
NIG H+D GKTTLT + + S ++D P E GITI +A
Sbjct: 273 NIGTIGHVDHGKTTLTAALTMALAALG-----NSAPKKYDEIDAAPEERARGITINTATV 437
Query: 117 YCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSITVDRQMRRYQV 176
+ +D PGH D+ + +DGA+LV+ G Q+ ++ V
Sbjct: 438 EYETENRHYAHVDCPGHADYVKNMITGAAQMDGAILVVSGADGPMPQTKEHILLAKQVGV 617
Query: 177 PR-IAFINKLDR 187
P + F+NK D+
Sbjct: 618 PNMVVFLNKQDQ 653
>TC214319 homologue to UP|EFT1_SOYBN (Q43467) Elongation factor Tu,
chloroplast precursor (EF-Tu), complete
Length = 1583
Score = 53.5 bits (127), Expect = 3e-07
Identities = 38/132 (28%), Positives = 57/132 (42%), Gaps = 1/132 (0%)
Frame = +1
Query: 57 NIGISAHIDSGKTTLTEWILFYTGKIHLMYEVRSKDGMGPKMDFKPLEIIMGITIKSAAT 116
NIG H+D GKTTLT + + S ++D P E GITI +A
Sbjct: 250 NIGTIGHVDHGKTTLTAALTMALAALG-----NSAPKKYDEIDAAPEERARGITINTATV 414
Query: 117 YCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQCQSITVDRQMRRYQV 176
+ +D PGH D+ + +DGA+LV+ G Q+ ++ V
Sbjct: 415 EYETENRHYAHVDCPGHADYVKNMITGAAQMDGAILVVSGADGPMPQTKEHILLAKQVGV 594
Query: 177 PR-IAFINKLDR 187
P + F+NK D+
Sbjct: 595 PNMVVFLNKQDQ 630
>TC232891 similar to UP|Q9LNC5 (Q9LNC5) F9P14.8 protein, partial (26%)
Length = 785
Score = 48.5 bits (114), Expect = 9e-06
Identities = 48/176 (27%), Positives = 77/176 (43%), Gaps = 17/176 (9%)
Frame = +3
Query: 376 RVYEGVIRKGDFIVNVSTGKKI---------KVPRLVQMHSNEMNDIEEAHAGQIVAVFG 426
RVY G I+ G + + G +V +L + + + EA G V + G
Sbjct: 213 RVYSGKIQTGQTVRVLGEGYSPDDEEDMTVKEVTKLWVYQARDRMPVAEAPPGSWVLIEG 392
Query: 427 VDCASSDTFTDGSVKYTMTSMNVPEP-------VMSLAVQPVSKDSGGKFSKALNRFQRE 479
VD + T T +V Y + + P V+ A +P++ K + L + +
Sbjct: 393 VDASIMKTATLCNVDYD-EDVYIFRPLQFNTLSVVKTATEPLNPSELPKMVEGLRKISKS 569
Query: 480 DPTFRVSLDPESGQTIISGMGELHLDIYVKRIKMEYG-VDATVGKPRVNFRETVTQ 534
P V+ ESG+ I G GEL+LD +K ++ Y V+ V P V+F ETV +
Sbjct: 570 YP-LAVTKVEESGEHTILGTGELYLDSIMKDLRELYSEVEVKVADPVVSFCETVVE 734
>TC226767 similar to GB|AAP68328.1|31711944|BT008889 At1g18070 {Arabidopsis
thaliana;} , partial (80%)
Length = 2011
Score = 44.3 bits (103), Expect = 2e-04
Identities = 32/120 (26%), Positives = 54/120 (44%), Gaps = 10/120 (8%)
Frame = +2
Query: 50 ESMEKVRNIGISAHIDSGKTTLTEWILFYTGKIH----LMYEVRSKDG------MGPKMD 99
E ++ N+ H+D+GK+T ILF +G++ YE +KD M MD
Sbjct: 230 EMTKRHLNVVFIGHVDAGKSTTGGQILFLSGQVDERTIQKYEKEAKDKSRESWYMAYIMD 409
Query: 100 FKPLEIIMGITIKSAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGG 159
E + G T++ + + ++ TI+D PGH + + D VLV+ + G
Sbjct: 410 TNEEERVKGKTVEVGRAHFETETTRFTILDAPGHKSYVPNMISGASQADIGVLVISARKG 589
>TC203946 homologue to UP|Q9LN13 (Q9LN13) T6D22.2, partial (47%)
Length = 1858
Score = 43.9 bits (102), Expect = 2e-04
Identities = 40/133 (30%), Positives = 57/133 (42%), Gaps = 11/133 (8%)
Frame = +3
Query: 53 EKVR-NIGISAHIDSGKTTLTEWILFYTGKI--HLMYEVRSKDGMGPKMDFKPLEII--- 106
EKV NI + H+DSGK+T T +++ G I ++ + K FK ++
Sbjct: 51 EKVHINIVVIGHVDSGKSTTTGHLIYKLGGIDKRVIERFEKEAAEMNKRSFKYAWVLDKL 230
Query: 107 -----MGITIKSAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQ 161
GITI A T+ID PGH DF + D AVL++ S G
Sbjct: 231 KAERERGITIDIALWKFETTKYYCTVIDAPGHRDFIKNMITGTSQADCAVLIIDSTTGGF 410
Query: 162 CQSITVDRQMRRY 174
I+ D Q R +
Sbjct: 411 EAGISKDGQTREH 449
>TC203954 homologue to UP|EF1A_LYCES (P17786) Elongation factor 1-alpha
(EF-1-alpha), partial (61%)
Length = 1423
Score = 43.9 bits (102), Expect = 2e-04
Identities = 40/133 (30%), Positives = 57/133 (42%), Gaps = 11/133 (8%)
Frame = +3
Query: 53 EKVR-NIGISAHIDSGKTTLTEWILFYTGKI--HLMYEVRSKDGMGPKMDFKPLEII--- 106
EKV NI + H+DSGK+T T +++ G I ++ + K FK ++
Sbjct: 441 EKVHINIVVIGHVDSGKSTTTGHLIYKLGGIDKRVIERFEKEAAEMNKRSFKYAWVLDKL 620
Query: 107 -----MGITIKSAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQ 161
GITI A T+ID PGH DF + D AVL++ S G
Sbjct: 621 KAERERGITIDIALWKFETTKYYCTVIDAPGHRDFIKNMITGTSQADCAVLIIDSTTGGF 800
Query: 162 CQSITVDRQMRRY 174
I+ D Q R +
Sbjct: 801 EAGISKDGQTREH 839
>TC203932 UP|EF1A_SOYBN (P25698) Elongation factor 1-alpha (EF-1-alpha),
complete
Length = 1766
Score = 42.7 bits (99), Expect = 5e-04
Identities = 40/137 (29%), Positives = 58/137 (42%), Gaps = 11/137 (8%)
Frame = +2
Query: 49 KESMEKVR-NIGISAHIDSGKTTLTEWILFYTGKI--HLMYEVRSKDGMGPKMDFKPLEI 105
K EKV +I + H+DSGK+T T +++ G I ++ + K FK +
Sbjct: 50 KMGKEKVHISIVVIGHVDSGKSTTTGHLIYKLGGIDKRVIERFEKEAAEMNKRSFKYAWV 229
Query: 106 I--------MGITIKSAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSV 157
+ GITI A T+ID PGH DF + D AVL++ S
Sbjct: 230 LDKLKAERERGITIDIALWKFETTKYYCTVIDAPGHRDFIKNMITGTSQADCAVLIIDST 409
Query: 158 GGVQCQSITVDRQMRRY 174
G I+ D Q R +
Sbjct: 410 TGGFEAGISKDGQTREH 460
>BU926992 homologue to GP|11181616|gb| translational elongation factor EF-TuM
{Zea mays}, partial (20%)
Length = 444
Score = 42.4 bits (98), Expect = 7e-04
Identities = 32/106 (30%), Positives = 49/106 (46%), Gaps = 5/106 (4%)
Frame = +1
Query: 33 NVARATAATIDKDPWWKESMEKVR-----NIGISAHIDSGKTTLTEWILFYTGKIHLMYE 87
N + +++++ +PWW+ R N+G H+D GKTTLT I K+ L E
Sbjct: 139 NDSASSSSSSTPNPWWRSMATFTRTKPHVNVGTIGHVDHGKTTLTAAIT----KV-LADE 303
Query: 88 VRSKDGMGPKMDFKPLEIIMGITIKSAATYCNWKGSKITIIDTPGH 133
++K ++D P E GITI +A +D PGH
Sbjct: 304 GKAKAVAFDEIDKAPEEKKRGITIATAHVEYETAKRHYAHVDCPGH 441
>AI899942 similar to PIR|S17434|S17 translation elongation factor eEF-1 alpha
chain (gene tefS1) - soybean, partial (37%)
Length = 529
Score = 34.7 bits (78), Expect = 0.14
Identities = 39/133 (29%), Positives = 56/133 (41%), Gaps = 11/133 (8%)
Frame = +2
Query: 53 EKVR-NIGISAHIDSGKTTLTEWILFYTG----KIHLMYEVRSKD--GMGPK----MDFK 101
EKV I + H+DSGK+T T +++ G ++ YE+ + + M K +D
Sbjct: 8 EKVHITIVVIGHVDSGKSTTTGHLIYKLGGIDKRVIERYEMEAAEMTTMSYKYAMVLDKL 187
Query: 102 PLEIIMGITIKSAATYCNWKGSKITIIDTPGHVDFTIEVERALRVLDGAVLVLCSVGGVQ 161
E GITI A T ID H DF + D AVL++ S G
Sbjct: 188 KAERERGITIDIALWKLETTKCYCTDIDAHRHRDFIKNMITETSQADCAVLIIDSTTGGS 367
Query: 162 CQSITVDRQMRRY 174
I+ D Q R +
Sbjct: 368 EAGISKDGQTREH 406
>BM108036
Length = 469
Score = 34.3 bits (77), Expect = 0.18
Identities = 27/76 (35%), Positives = 33/76 (42%)
Frame = +2
Query: 150 AVLVLCSVGGVQCQSITVDRQMRRYQVPRIAFINKLDRPGADPWKVITQARSKLRHHCAA 209
AVL+ C V G+ + I VD R+ P +N L + A I R KL A
Sbjct: 38 AVLIFCKVNGIDFEEIKVDLSKRQQLSPEFRAVNPLRKVPA-----IVDGRFKLFESHAI 202
Query: 210 LQVPIGLESDFKGVVD 225
L I L S F GV D
Sbjct: 203 L---IYLASAFPGVAD 241
>TC220356 similar to UP|Q6SYB1 (Q6SYB1) GTP-binding protein TypA, partial
(9%)
Length = 432
Score = 34.3 bits (77), Expect = 0.18
Identities = 27/80 (33%), Positives = 39/80 (48%), Gaps = 2/80 (2%)
Frame = +3
Query: 2 QRFGYALCSSTLTTEGSLIAGTFHIRHFSAGNVARATAATIDKDPWWKESMEK--VRNIG 59
Q G L SS+ + A F R +V+ AT +K ++ + + VRNI
Sbjct: 168 QLLGLPLSSSSSSASSLTAAKRFSCRPIKC-SVSEATEPKTEKK---RQLLRRGDVRNIA 335
Query: 60 ISAHIDSGKTTLTEWILFYT 79
I AH+D GKTTL + +L T
Sbjct: 336 IVAHVDHGKTTLVDAMLKQT 395
>TC231080 UP|Q9FQD5 (Q9FQD5) Glutathione S-transferase GST 23 , partial
(34%)
Length = 568
Score = 34.3 bits (77), Expect = 0.18
Identities = 27/76 (35%), Positives = 33/76 (42%)
Frame = +3
Query: 150 AVLVLCSVGGVQCQSITVDRQMRRYQVPRIAFINKLDRPGADPWKVITQARSKLRHHCAA 209
AVL+ C V G+ + I VD R+ P +N L + A I R KL A
Sbjct: 60 AVLIFCKVNGIDFEEIKVDLSKRQQLSPEFRAVNPLRKVPA-----IVDGRFKLFESHAI 224
Query: 210 LQVPIGLESDFKGVVD 225
L I L S F GV D
Sbjct: 225 L---IYLASAFPGVAD 263
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.319 0.136 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 23,296,082
Number of Sequences: 63676
Number of extensions: 266827
Number of successful extensions: 1215
Number of sequences better than 10.0: 50
Number of HSP's better than 10.0 without gapping: 1190
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1196
length of query: 654
length of database: 12,639,632
effective HSP length: 103
effective length of query: 551
effective length of database: 6,081,004
effective search space: 3350633204
effective search space used: 3350633204
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146553.2


