

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146551.3 - phase: 1 /pseudo
(379 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
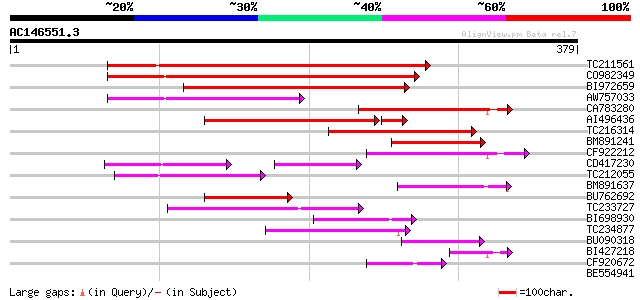
Score E
Sequences producing significant alignments: (bits) Value
TC211561 weakly similar to UP|O22675 (O22675) Reverse transcript... 186 1e-47
CO982349 177 7e-45
BI972659 134 5e-32
AW757033 114 7e-26
CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polym... 109 2e-24
AI496436 100 2e-23
TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {A... 103 1e-22
BM891241 84 1e-16
CF922212 78 8e-15
CD417230 weakly similar to GP|2244915|emb| reverse transcriptase... 50 9e-13
TC212055 70 2e-12
BM891637 60 2e-09
BU762692 59 4e-09
TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR ret... 57 2e-08
BI698930 52 3e-07
TC234877 49 3e-06
BU090318 weakly similar to GP|9743345|gb| F21J9.30 {Arabidopsis ... 47 2e-05
BI427218 42 6e-04
CF920672 41 8e-04
BE554941 36 0.033
>TC211561 weakly similar to UP|O22675 (O22675) Reverse transcriptase
(Fragment) , partial (59%)
Length = 1077
Score = 186 bits (473), Expect = 1e-47
Identities = 86/216 (39%), Positives = 133/216 (60%)
Frame = +1
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PL+PFLF + AEG LM V ++Y + VG + V ++ LQ+ADDT+ GE NV
Sbjct: 265 PLTPFLFNIVAEGLNGLMRRAVEENLYKPYLVG-ANGVPISILQYADDTIFFGEAAKENV 441
Query: 126 RTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASALMNCCRGNFTFVYLGVPIGG 185
+ +L FE +S L++NF KS ++D W EA+ ++C F F+YLG+PIG
Sbjct: 442 EAIKVILRSFELVSNLRINFAKSCFGVFGVTDQWKQEAANYLHCSLLAFPFIYLGIPIGA 621
Query: 186 DPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIISS 245
+ R+ W +I + +LS W + +S GGR+ L+K VL+S+P+YF SFF+ P ++
Sbjct: 622 NSRRCHMWDSIIRKCERKLSKWK*RHISFGGRVTLIKSVLTSIPIYFFSFFRVPQSVVDK 801
Query: 246 IESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGL 281
+ I ++F WGGG + +KI+WI+W+ +CL K GG+
Sbjct: 802 LVKI*RTFLWGGGPDQKKISWIRWEKLCLPKERGGI 909
>CO982349
Length = 795
Score = 177 bits (449), Expect = 7e-45
Identities = 84/209 (40%), Positives = 127/209 (60%)
Frame = +3
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PL+P LF + AEG T LM V ++SF VG E V+ LQ+ADDT+ E N+
Sbjct: 168 PLAPLLFNIVAEGLTGLMREAVDRKRFNSFLVGKNKEP-VSILQYADDTIFFEEATMENM 344
Query: 126 RTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASALMNCCRGNFTFVYLGVPIGG 185
+ + +L FE SGLK+NF +S + D W EA+ +NC + F YLG+PI
Sbjct: 345 KVIKTILRCFELASGLKINFAESRFGAIWKPDHWCKEAAEFLNCSMLSMPFSYLGIPIEA 524
Query: 186 DPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIISS 245
+PR P+I + +L+ W + +S+GGR+ L+ +L++LP+YF SFF+AP+ ++
Sbjct: 525 NPRCREI*DPIIRKCETKLARWKQRHISLGGRVTLINAILTALPIYFFSFFRAPSKVVQK 704
Query: 246 IESIFKSFFWGGGEENRKIAWIKWDTICL 274
+ +I + F WGGG R+IAW+KW+T+CL
Sbjct: 705 LVNIQRKFLWGGGSXQRRIAWVKWETVCL 791
>BI972659
Length = 453
Score = 134 bits (338), Expect = 5e-32
Identities = 59/151 (39%), Positives = 96/151 (63%)
Frame = +1
Query: 117 IGEKCWLNVRTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASALMNCCRGNFTF 176
+GE W N+ + ++L FE +SGL++N+ KS V +W +A+ L+NC + + F
Sbjct: 1 VGEASWDNIIVLKSMLRGFEMVSGLRINYAKSQFGIVGFQPNWAHDAAQLLNCRQLDIPF 180
Query: 177 VYLGVPIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFF 236
YLG+PI W+P+I++ +LS WN K LS+GGR+ L+K VL++LP+Y LSFF
Sbjct: 181 HYLGMPIAVKASSRMVWEPLINKFKAKLSKWNQKYLSMGGRVTLIKSVLNALPIYLLSFF 360
Query: 237 KAPAGIISSIESIFKSFFWGGGEENRKIAWI 267
K P I+ + S+ ++F WGG + + +I+W+
Sbjct: 361 KIPQRIVDKLVSLQRTFMWGGNQHHNRISWV 453
>AW757033
Length = 441
Score = 114 bits (285), Expect = 7e-26
Identities = 60/132 (45%), Positives = 79/132 (59%)
Frame = -2
Query: 66 PLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNV 125
PL+P LF +AAEG T LM V + Y SF VG + V+ LQ+ADDT+ GE NV
Sbjct: 398 PLAPLLFNIAAEGLTGLMREAVARNHYKSFAVGKYKKP-VSILQYADDTIFFGEATMENV 222
Query: 126 RTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASALMNCCRGNFTFVYLGVPIGG 185
R + +L FE SGLK+NF KS +++ D W EA+ MNC + F YLG+PIG
Sbjct: 221 RVIKTILRGFELASGLKINFAKSRFGAISVPD*WRKEAAEFMNCSLLSLPFSYLGIPIGA 42
Query: 186 DPRKLSFWKPVI 197
+PR+ W P+I
Sbjct: 41 NPRRRETWDPII 6
>CA783280 weakly similar to PIR|T01871|T01 RNA-directed DNA polymerase
homolog T24M8.8 - Arabidopsis thaliana, partial (4%)
Length = 456
Score = 109 bits (273), Expect = 2e-24
Identities = 51/108 (47%), Positives = 69/108 (63%), Gaps = 5/108 (4%)
Frame = +2
Query: 234 SFFKAPAGIISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSL 293
SFF P II+ +ES+ + F WGG ++RKIAW+ W T+CL KA+GGLG++ + FN +L
Sbjct: 5 SFFSLP*KIIAKLESLQRRFLWGGEADSRKIAWVNWKTVCLPKAKGGLGIKDLRTFNTTL 184
Query: 294 LGKWCWRMLTERDELWYRVLKAKYG-----EEGGRIREGGRLSSAWWK 336
LGKW W + + E W +VL++KYG EEG G SAWWK
Sbjct: 185 LGKWRWDLFYIQQEPWAKVLQSKYGGWRALEEG----SSGSKDSAWWK 316
>AI496436
Length = 414
Score = 100 bits (249), Expect(2) = 2e-23
Identities = 48/117 (41%), Positives = 71/117 (60%)
Frame = +1
Query: 131 VLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASALMNCCRGNFTFVYLGVPIGGDPRKL 190
+L FE SGLK+NF KS + SD W EA+ +N + FVYLG+PIG + R
Sbjct: 4 ILRCFELSSGLKINFAKSQFGTICKSDIWRKEAADYLNSSVLSMPFVYLGIPIGANSRHS 183
Query: 191 SFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIISSIE 247
W+PV+ + +L+SW K +S GGR+ L+ VL++LP+Y LSFF+ P ++ +
Sbjct: 184 DVWEPVVRKCERKLASWKQKYVSFGGRVTLINSVLTALPIYLLSFFRIPDKVVDKAD 354
Score = 26.6 bits (57), Expect(2) = 2e-23
Identities = 11/18 (61%), Positives = 12/18 (66%)
Frame = +2
Query: 249 IFKSFFWGGGEENRKIAW 266
I + F WGG E RKIAW
Sbjct: 359 IQRRFLWGGDLE*RKIAW 412
>TC216314 similar to GB|AAP68269.1|31711826|BT008830 At1g70320 {Arabidopsis
thaliana;} , partial (18%)
Length = 759
Score = 103 bits (258), Expect = 1e-22
Identities = 41/99 (41%), Positives = 66/99 (66%)
Frame = +2
Query: 214 IGGRLVLLKVVLSSLPVYFLSFFKAPAGIISSIESIFKSFFWGGGEENRKIAWIKWDTIC 273
+GGR+ L+K VLS+LP+ LSFFK P I+ + ++ + F WGG + + +I W+KW IC
Sbjct: 2 MGGRVTLIKSVLSALPIXXLSFFKIPQRIVDKLVTLQRQFLWGGTQHHNRIPWVKWADIC 181
Query: 274 LSKAEGGLGVRRMGAFNVSLLGKWCWRMLTERDELWYRV 312
K +GGLG++ + FN +L G+W W + + ++LW R+
Sbjct: 182 NPKIDGGLGIKDLSNFNAALRGRWIWGLASNHNQLWARL 298
>BM891241
Length = 407
Score = 84.0 bits (206), Expect = 1e-16
Identities = 28/63 (44%), Positives = 48/63 (75%)
Frame = -2
Query: 256 GGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTERDELWYRVLKA 315
GG +++KI W+KW+ +CL K +GGLG++ + FN++LLGKW W + +++ +LW R++ +
Sbjct: 406 GGDHDHKKIPWVKWEVVCLPKTKGGLGIKDISKFNIALLGKWIWALASDQQQLWARIINS 227
Query: 316 KYG 318
KYG
Sbjct: 226 KYG 218
>CF922212
Length = 445
Score = 77.8 bits (190), Expect = 8e-15
Identities = 40/114 (35%), Positives = 60/114 (52%), Gaps = 5/114 (4%)
Frame = -2
Query: 239 PAGIISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWC 298
P ++ + + + F WGGG +KIAW+ WD++CL K + GLG R + FN +LLGK
Sbjct: 438 PNKVLDKLVRLQQWFLWGGGLGQKKIAWVNWDSVCLPKEKEGLGNRDLRKFNYALLGKRR 259
Query: 299 WRMLTERDELWYRVLKAKYG-----EEGGRIREGGRLSSAWWKMVCHIREGLVD 347
W + + EL RVL +KY +E R++ S WW+ + I D
Sbjct: 258 WNLFHHQGELGARVLDSKYKRWRNLDEERRVKS----ESFWWQEISFITHSTED 109
>CD417230 weakly similar to GP|2244915|emb| reverse transcriptase like
protein {Arabidopsis thaliana}, partial (7%)
Length = 688
Score = 49.7 bits (117), Expect(2) = 9e-13
Identities = 29/85 (34%), Positives = 45/85 (52%)
Frame = -3
Query: 64 RGPLSPFLFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWL 123
R P++P+LF+L E + L+N V+ + S V G + ++HL F DD ++ E
Sbjct: 587 RDPIAPYLFVLCLERLSHLINLVMNKKL*RSIQVNRGG-LLISHLAFVDDLILFVEANMD 411
Query: 124 NVRTMMAVLLLFEEISGLKVNFHKS 148
V + L LF + SG KVN K+
Sbjct: 410 QVDVINLTLELFSDSSGAKVNVDKT 336
Score = 41.2 bits (95), Expect(2) = 9e-13
Identities = 23/58 (39%), Positives = 33/58 (56%)
Frame = -2
Query: 178 YLGVPIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSF 235
YLGVPI + +ID++ R S W + LLS+ GRL L K V+ +LP + + F
Sbjct: 243 YLGVPIFHKNPTSHTFDFIIDKVNKRFSRWKSNLLSMVGRLTLTK*VV*ALPSHIMQF 70
>TC212055
Length = 776
Score = 69.7 bits (169), Expect = 2e-12
Identities = 40/101 (39%), Positives = 56/101 (54%)
Frame = +1
Query: 71 LFLLAAEGFTVLMNAVVGTDMYHSFGVGNGSEVRVTHLQFADDTLIIGEKCWLNVRTMMA 130
LF + AEG T LM + +Y S VG +++ V LQ+ADDT+ +GE NV T+ +
Sbjct: 1 LFNIVAEGLTGLMREALDKSLYSSLMVGK-NKIPVNILQYADDTIFLGEATMQNVMTIKS 177
Query: 131 VLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASALMNCCR 171
+L +FE SGLK++F KS + D W EA CR
Sbjct: 178 MLRVFELASGLKIHFAKSSFGAIGKPDHWQKEAGLNTFNCR 300
>BM891637
Length = 427
Score = 60.1 bits (144), Expect = 2e-09
Identities = 32/79 (40%), Positives = 43/79 (53%), Gaps = 3/79 (3%)
Frame = -1
Query: 260 ENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTERDELWYRVLKAKYGE 319
E +KIAWI W C S GGLG++ + N +LL KW W M + +LW R+L +KY
Sbjct: 424 EGKKIAWISWRQCCASGDVGGLGIKDIKILNNALLIKWKWLMFHQPHQLWNRILISKY-- 251
Query: 320 EGGRIREGGRLS---SAWW 335
+G R + G S WW
Sbjct: 250 KGWRGLDQGPQKYYFSPWW 194
>BU762692
Length = 423
Score = 58.9 bits (141), Expect = 4e-09
Identities = 26/59 (44%), Positives = 37/59 (62%)
Frame = -3
Query: 131 VLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASALMNCCRGNFTFVYLGVPIGGDPRK 189
+L FE +SGLK+NF KS ++D W EA++ +NC FVY G+PIG +PR+
Sbjct: 181 ILRTFELVSGLKINFAKSCFGAFGMTDRWKTEAASCLNCSLLAIPFVYQGIPIGANPRR 5
>TC233727 similar to UP|Q9FKH8 (Q9FKH8) Similarity to non-LTR retroelement
reverse transcriptase, partial (7%)
Length = 956
Score = 56.6 bits (135), Expect = 2e-08
Identities = 39/132 (29%), Positives = 66/132 (49%), Gaps = 1/132 (0%)
Frame = -2
Query: 106 THLQFADDTLIIGEKCWLNVRTMMAVLLLFEEISGLKVNFHKSMLTGVNISDSWLVEASA 165
+H+ ADD +I + NV ++ L+++ G ++ HKS + +I L S
Sbjct: 451 SHVMHADDIMIFCKGTSRNVHNLINFFQLYKDSFGQVISKHKSKIFIGSIPGQRLRVISD 272
Query: 166 LMNCCRGNFTFVYLGVPI-GGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVV 224
L+ + F Y GVPI G P + + V D+I +L+SW +LSI GR+ L+ V
Sbjct: 271 LLQYFHSDIPFNYFGVPIFKGKPTRRHLY-CVADKIKCKLASWKGSILSIMGRIQLVNSV 95
Query: 225 LSSLPVYFLSFF 236
+ + +Y S +
Sbjct: 94 IHGMLLYSFSVY 59
>BI698930
Length = 384
Score = 52.4 bits (124), Expect = 3e-07
Identities = 26/69 (37%), Positives = 40/69 (57%)
Frame = +3
Query: 204 LSSWNNKLLSIGGRLVLLKVVLSSLPVYFLSFFKAPAGIISSIESIFKSFFWGGGEENRK 263
L+ W +++LS GR L+ +LSSLP++ SFFK P + SI ++F G E+
Sbjct: 81 LALWKHEVLSFAGRTCLINSMLSSLPLFSFSFFKVPLNVGMRFISI*RNFV-GEWRES*M 257
Query: 264 IAWIKWDTI 272
I W+ WD +
Sbjct: 258 IPWVNWDKV 284
>TC234877
Length = 534
Score = 49.3 bits (116), Expect = 3e-06
Identities = 25/99 (25%), Positives = 48/99 (48%), Gaps = 2/99 (2%)
Frame = +1
Query: 172 GNFTFVYLGVPIGGDPRKLSFWKPVIDRIVGRLSSWNNKLLSIGGRLVLLKVVLSSLPVY 231
G+ F+YL VP+ + + +P+ DRI L SW LLS G + L+ ++ + +
Sbjct: 31 GSLPFIYLSVPLFQEKPRPVHLRPIADRIKAELQSWKGNLLSFMGCVQLIISIIDGMLL* 210
Query: 232 FLSFFKAPAGIISSIESIFKSFFWGGG--EENRKIAWIK 268
+ P ++ +++ ++F W G +E W+K
Sbjct: 211 SFHLYSWPKALLIEMQTWTRNFIWTRGYFKEKTGYCWLK 327
>BU090318 weakly similar to GP|9743345|gb| F21J9.30 {Arabidopsis thaliana},
partial (1%)
Length = 422
Score = 46.6 bits (109), Expect = 2e-05
Identities = 22/55 (40%), Positives = 32/55 (58%)
Frame = +3
Query: 263 KIAWIKWDTICLSKAEGGLGVRRMGAFNVSLLGKWCWRMLTERDELWYRVLKAKY 317
K+ W W +C K GGLG+R + N SLL K M T+ ++L RV+++KY
Sbjct: 3 KVHWTSWKALCSPKKSGGLGLRLLSHMNKSLLMKST*NMCTKPNDL*VRVVRSKY 167
>BI427218
Length = 421
Score = 41.6 bits (96), Expect = 6e-04
Identities = 21/47 (44%), Positives = 25/47 (52%), Gaps = 5/47 (10%)
Frame = +1
Query: 295 GKWCWRMLTERDELWYRVLKAKYG-----EEGGRIREGGRLSSAWWK 336
GKW W + ELW R+L +KYG EE R G +SAWWK
Sbjct: 1 GKWKWSFFYNKGELWARILDSKYGDWRHLEEPTR----GNRASAWWK 129
>CF920672
Length = 792
Score = 41.2 bits (95), Expect = 8e-04
Identities = 19/54 (35%), Positives = 29/54 (53%)
Frame = -2
Query: 239 PAGIISSIESIFKSFFWGGGEENRKIAWIKWDTICLSKAEGGLGVRRMGAFNVS 292
P + + I + F WGGG + +KIAW+KWD C + L V+ +N+S
Sbjct: 791 PNRVAEKLTQIQRRFXWGGGLDQKKIAWVKWD--C*LMEQEPLRVKYPRLYNIS 636
>BE554941
Length = 383
Score = 35.8 bits (81), Expect = 0.033
Identities = 18/60 (30%), Positives = 27/60 (45%)
Frame = +1
Query: 282 GVRRMGAFNVSLLGKWCWRMLTERDELWYRVLKAKYGEEGGRIREGGRLSSAWWKMVCHI 341
G++ + FN+SLL K L E +W +LK YG +S WW+ + I
Sbjct: 7 GIKNLSIFNISLLVKRR*HFLVETKAIWAPILKFHYGSLDLSNTGSASRTSQWWRDILRI 186
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.331 0.146 0.494
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 20,750,621
Number of Sequences: 63676
Number of extensions: 335750
Number of successful extensions: 2126
Number of sequences better than 10.0: 59
Number of HSP's better than 10.0 without gapping: 2100
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2122
length of query: 379
length of database: 12,639,632
effective HSP length: 99
effective length of query: 280
effective length of database: 6,335,708
effective search space: 1773998240
effective search space used: 1773998240
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146551.3


