

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
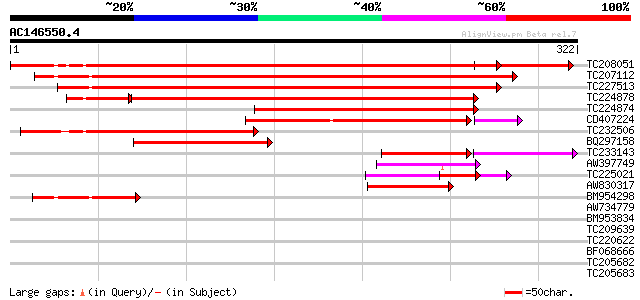
Query= AC146550.4 - phase: 0 /pseudo
(322 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC208051 weakly similar to UP|Q8S341 (Q8S341) Purple acid phosph... 410 e-115
TC207112 similar to UP|Q6J5M7 (Q6J5M7) Purple acid phosphatase 1... 337 4e-93
TC227513 weakly similar to UP|Q6J5M7 (Q6J5M7) Purple acid phosph... 335 2e-92
TC224878 285 4e-83
TC224874 similar to UP|Q8LFV2 (Q8LFV2) Acid phosphatase type 5, ... 196 2e-50
CD407224 179 2e-45
TC232506 157 8e-39
BQ297158 similar to GP|20257477|gb| purple acid phosphatase {Ara... 142 2e-34
TC233143 similar to UP|Q8VYZ2 (Q8VYZ2) Purple acid phosphatase, ... 85 1e-19
AW397749 87 1e-17
TC225021 similar to UP|Q8LFV2 (Q8LFV2) Acid phosphatase type 5, ... 70 1e-12
AW830317 67 1e-11
BM954298 48 7e-06
AW734779 32 0.39
BM953834 32 0.51
TC209639 30 1.1
TC220622 similar to UP|Q8S2H5 (Q8S2H5) Diphosphonucleotide phosp... 30 1.1
BF068666 30 1.9
TC205682 similar to GB|AAM67420.1|21689597|AY122887 AT5g55940/MY... 30 1.9
TC205683 similar to GB|AAM67420.1|21689597|AY122887 AT5g55940/MY... 30 1.9
>TC208051 weakly similar to UP|Q8S341 (Q8S341) Purple acid phosphatase,
partial (84%)
Length = 1249
Score = 410 bits (1053), Expect = e-115
Identities = 199/279 (71%), Positives = 237/279 (84%)
Frame = +2
Query: 1 MAWCLKQRMLSPVIFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKG 60
MA L QRM+ PV+ + L++ SIAE LPRF+H K +QQSLN LV+GDWGRKG
Sbjct: 26 MASSLNQRMVFPVMVVVGMFCLLVT--PSIAE-LPRFKHPPK-KQQSLNILVLGDWGRKG 193
Query: 61 NYNQSLVAHQMGIVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIW 120
YNQSLVA+QMGIVG+ L+IDFVISTGDNFY+DGL+GVDDP FY+SF+++YTAPSLQK W
Sbjct: 194 TYNQSLVANQMGIVGEKLDIDFVISTGDNFYEDGLKGVDDPAFYQSFIHMYTAPSLQKTW 373
Query: 121 YSVLGNHDYRGDVDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPK 180
Y+VLGNHDYRGDV+AQLS IL+QKDSRW+C+RSFILDG IVEFFFVDTTPF+E+YFTDP
Sbjct: 374 YTVLGNHDYRGDVEAQLSPILKQKDSRWLCMRSFILDGEIVEFFFVDTTPFVEEYFTDPG 553
Query: 181 EHTYDWNGVLPRESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLL 240
EHTYDW GVLPR +Y +ELLK+VDLAL +SKAKWK+VVGHHTI S GHHG+T++L+Q L+
Sbjct: 554 EHTYDWEGVLPRLAYLSELLKDVDLALAQSKAKWKMVVGHHTINSAGHHGSTEDLKQLLV 733
Query: 241 PILKSNNIDAYINGHDHCLQHIIDNERHGEAISSHGTRK 279
PIL++NN+DAYINGHDHCLQHIIDN I+S G K
Sbjct: 734 PILEANNVDAYINGHDHCLQHIIDNNNGIHFITSGGGSK 850
Score = 58.9 bits (141), Expect = 3e-09
Identities = 31/57 (54%), Positives = 39/57 (68%), Gaps = 1/57 (1%)
Frame = +3
Query: 265 NERHGEAISSHGTRK-*SFTMTDKDSCLCRSQK*M*MLYSMTFWARFCIDGAYRKNL 320
++RHG+A+SSHG K *S M KDSCLCRS K + + Y M A+FCI GAY K +
Sbjct: 843 DQRHGQAMSSHGNWKN*SCIMMGKDSCLCRSPKALHISYFMMLLAKFCIHGAYPKTV 1013
>TC207112 similar to UP|Q6J5M7 (Q6J5M7) Purple acid phosphatase 1, partial
(80%)
Length = 1200
Score = 337 bits (864), Expect = 4e-93
Identities = 167/274 (60%), Positives = 203/274 (73%)
Frame = +1
Query: 15 FPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIV 74
F V L LLS S++ L R EH +K SL+ +V+GDWGRKG YNQS VA QMG V
Sbjct: 25 FKRFVSLCLLS--VSVSGLLQRLEHPVKADG-SLSLMVIGDWGRKGTYNQSQVATQMGRV 195
Query: 75 GDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGDVD 134
LNIDFVISTGDNFY DGL G+DDP F SF IYTA SLQK WYSVLGNHDYRGDV+
Sbjct: 196 AAKLNIDFVISTGDNFYDDGLTGIDDPAFEISFSKIYTAKSLQKQWYSVLGNHDYRGDVE 375
Query: 135 AQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRES 194
AQL+ IL++ D RW+C RSFI+D I EFFF+D+TPF++KYF PK+H YDW GVLPRE
Sbjct: 376 AQLNPILQKIDPRWICQRSFIVDTEIAEFFFIDSTPFVDKYFLKPKDHKYDWRGVLPREK 555
Query: 195 YRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYING 254
Y ++LLK++++AL S AKWKIVVGHH ++S+GHHG+T+EL +QLLPIL+ NN+D YING
Sbjct: 556 YLSKLLKDLEIALKDSTAKWKIVVGHHPVRSIGHHGDTKELIRQLLPILEENNVDMYING 735
Query: 255 HDHCLQHIIDNERHGEAISSHGTRK*SFTMTDKD 288
HDHCL+HI + ++S G K DKD
Sbjct: 736 HDHCLEHISSRSSQIQFLTSGGGSKAWKGDMDKD 837
Score = 30.4 bits (67), Expect = 1.1
Identities = 19/55 (34%), Positives = 29/55 (52%), Gaps = 4/55 (7%)
Frame = +2
Query: 262 IIDNERHGEA----ISSHGTRK*SFTMTDKDSCLCRSQK*M*MLYSMTFWARFCI 312
+++ +RHG+ I G+ SFTM DKD C +K M L+ F +FC+
Sbjct: 794 VVEVQRHGKETWIKIRKMGS---SFTMMDKDLCPWSWKKPMPKLFISIFMGKFCM 949
>TC227513 weakly similar to UP|Q6J5M7 (Q6J5M7) Purple acid phosphatase 1,
partial (86%)
Length = 1229
Score = 335 bits (859), Expect = 2e-92
Identities = 157/252 (62%), Positives = 195/252 (77%)
Frame = +2
Query: 28 SSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGIVGDNLNIDFVISTG 87
SS KL +H K SL+FLVVGDWGRKG YNQSLVA QMG++G+ L++DFVISTG
Sbjct: 134 SSSKAKLESLQHAPKADG-SLSFLVVGDWGRKGAYNQSLVAFQMGVIGEKLDVDFVISTG 310
Query: 88 DNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYRGDVDAQLSSILRQKDSR 147
DNFY +GL GV DP+F ESF IYTAPSLQK WY+VLGNHDYRG+ AQ+S +LR +D+R
Sbjct: 311 DNFYDNGLTGVFDPSFEESFTKIYTAPSLQKKWYNVLGNHDYRGNAKAQISHVLRYRDNR 490
Query: 148 WVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAELLKNVDLAL 207
WVC RS+ L+ V+FFFVDTTPF++KYF + K H YDW G+LPR+ Y + LLK+VDLAL
Sbjct: 491 WVCFRSYTLNSENVDFFFVDTTPFVDKYFIEDKGHNYDWRGILPRKRYISNLLKDVDLAL 670
Query: 208 VKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHIIDNER 267
+S A WK+V+GHHTIKS+GHHG+TQEL LP+LK+NN+D YINGHDHCL+HI +
Sbjct: 671 RQSTATWKVVIGHHTIKSIGHHGDTQELLIHFLPLLKANNVDLYINGHDHCLEHISSLDS 850
Query: 268 HGEAISSHGTRK 279
+ ++S G K
Sbjct: 851 SVQFLTSGGGSK 886
Score = 28.9 bits (63), Expect = 3.3
Identities = 14/32 (43%), Positives = 19/32 (58%), Gaps = 1/32 (3%)
Frame = +3
Query: 262 IIDNERHGEAISSHGTR-K*SFTMTDKDSCLC 292
++ + RHGE + +*SFT T K SCLC
Sbjct: 870 VVVDRRHGEVTRNRAKEMR*SFTTTAKASCLC 965
>TC224878
Length = 1228
Score = 285 bits (729), Expect(2) = 4e-83
Identities = 130/197 (65%), Positives = 160/197 (80%)
Frame = +3
Query: 70 QMGIVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDY 129
QMG VG+ L+IDFV+STGDNFY +GL D F ESF IYTA SLQ WYSVLGNHDY
Sbjct: 255 QMGKVGEKLDIDFVVSTGDNFYDNGLTSDHDNAFQESFTQIYTAKSLQNQWYSVLGNHDY 434
Query: 130 RGDVDAQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGV 189
RGD +AQLS +LR+ DSRW+CLRSFI+D +VE FFVDTTPF+++YFT+P EH YDW G+
Sbjct: 435 RGDAEAQLSPVLREIDSRWLCLRSFIVDSELVEIFFVDTTPFVDEYFTEPPEHKYDWRGI 614
Query: 190 LPRESYRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNID 249
P++SY + LLK+++LAL S AKW+IVVGHH I+SVGHHG+TQEL +LLPIL++NN+
Sbjct: 615 GPQKSYISNLLKDLELALRGSTAKWRIVVGHHAIRSVGHHGDTQELINRLLPILQANNVH 794
Query: 250 AYINGHDHCLQHIIDNE 266
Y+NGHDHCL+HI D E
Sbjct: 795 FYMNGHDHCLEHISDTE 845
Score = 41.2 bits (95), Expect(2) = 4e-83
Identities = 20/38 (52%), Positives = 26/38 (67%)
Frame = +1
Query: 33 KLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQ 70
+L R H K +L+FLV+GDWGR+G YNQS V+ Q
Sbjct: 127 ELQRLSHSSK-HDGALSFLVLGDWGRRGAYNQSQVSFQ 237
>TC224874 similar to UP|Q8LFV2 (Q8LFV2) Acid phosphatase type 5, partial
(51%)
Length = 806
Score = 196 bits (497), Expect = 2e-50
Identities = 86/127 (67%), Positives = 108/127 (84%)
Frame = +3
Query: 140 ILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAEL 199
+LR+ DSRW+CLRSFI+D +VE FFVDTTPF+E+YFT+P+EH YDW G+ P++ Y L
Sbjct: 6 VLREIDSRWLCLRSFIVDSELVEIFFVDTTPFVEEYFTEPQEHKYDWRGIGPQKPYITNL 185
Query: 200 LKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCL 259
LK+++LAL +S AKWKIVVGHH I+SVGHHG+TQEL QLLPIL++NNID Y+NGHDHCL
Sbjct: 186 LKDLELALRESTAKWKIVVGHHAIRSVGHHGDTQELINQLLPILQANNIDFYMNGHDHCL 365
Query: 260 QHIIDNE 266
+HI D E
Sbjct: 366 EHISDTE 386
>CD407224
Length = 692
Score = 179 bits (454), Expect(2) = 2e-45
Identities = 79/128 (61%), Positives = 100/128 (77%)
Frame = -2
Query: 135 AQLSSILRQKDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRES 194
AQLS +LR+ D RW C RSFIL+ G+ EFFF+DTTPF+ KYF + H YDW GVLPR+
Sbjct: 685 AQLSPLLRKIDRRWFCQRSFILNAGVAEFFFIDTTPFMRKYFNNSNRH-YDWRGVLPRQK 509
Query: 195 YRAELLKNVDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYING 254
Y LLK+++ L KS A+WKI VGHH I+S+GHHG++ EL + L+P+LK+NN+D YING
Sbjct: 508 YLKTLLKDLEEELRKSTARWKIAVGHHAIRSIGHHGDSPELVKHLVPVLKANNVDMYING 329
Query: 255 HDHCLQHI 262
HDHCLQHI
Sbjct: 328 HDHCLQHI 305
Score = 21.2 bits (43), Expect(2) = 2e-45
Identities = 11/27 (40%), Positives = 15/27 (54%)
Frame = -3
Query: 265 NERHGEAISSHGTRK*SFTMTDKDSCL 291
++RHGE + T *+ M K SCL
Sbjct: 261 DQRHGEVMLKKPTLM*NSFMMVKVSCL 181
>TC232506
Length = 437
Score = 157 bits (396), Expect = 8e-39
Identities = 82/135 (60%), Positives = 99/135 (72%)
Frame = +3
Query: 7 QRMLSPVIFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSL 66
Q M ++F ++ L L+ + + L RFE LK Q SL+FLV+GDWGRKG YNQS
Sbjct: 45 QHMGLQLVFIGTIALCLVVSSAV----LERFEQALK-QDGSLSFLVIGDWGRKGAYNQSK 209
Query: 67 VAHQMGIVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGN 126
VA QMG++G L+IDFVISTGDNFY GL G+DDP F +SF IYTA SLQK WYSVLGN
Sbjct: 210 VAFQMGVIGQLLDIDFVISTGDNFYDSGLTGIDDPDFDDSFTKIYTASSLQKQWYSVLGN 389
Query: 127 HDYRGDVDAQLSSIL 141
HDYRG+V+AQLS +L
Sbjct: 390 HDYRGNVEAQLSPVL 434
>BQ297158 similar to GP|20257477|gb| purple acid phosphatase {Arabidopsis
thaliana}, partial (23%)
Length = 422
Score = 142 bits (359), Expect = 2e-34
Identities = 63/79 (79%), Positives = 75/79 (94%)
Frame = +1
Query: 71 MGIVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQKIWYSVLGNHDYR 130
MGIVG+ L+IDFVISTGDNFY+DGL+GVDDP FY+SF+++YTAPSLQK WY+VLGNHDYR
Sbjct: 184 MGIVGEKLDIDFVISTGDNFYEDGLKGVDDPAFYQSFIHMYTAPSLQKTWYTVLGNHDYR 363
Query: 131 GDVDAQLSSILRQKDSRWV 149
GDV+AQLS IL+QKDSRW+
Sbjct: 364 GDVEAQLSPILKQKDSRWL 420
>TC233143 similar to UP|Q8VYZ2 (Q8VYZ2) Purple acid phosphatase, partial
(36%)
Length = 765
Score = 85.1 bits (209), Expect(2) = 1e-19
Identities = 38/51 (74%), Positives = 42/51 (81%)
Frame = +3
Query: 212 AKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHI 262
AKWKIVVGHHTI+ G H NT EL +QLLPIL++NNID YINGHDH QHI
Sbjct: 3 AKWKIVVGHHTIRXAGVHXNTDELVKQLLPILEANNIDLYINGHDHXXQHI 155
Score = 28.9 bits (63), Expect(2) = 1e-19
Identities = 21/60 (35%), Positives = 31/60 (51%), Gaps = 1/60 (1%)
Frame = +1
Query: 264 DNERHGEAISSHGTRK-*SFTMTDKDSCLCRSQK*M*MLYSMTFWARFCIDGAYRKNLKQ 322
+ +R G + + G++K *S T K+SCL RS K YSM A F +G + + Q
Sbjct: 196 EGQRRGGVL*TGGSQKR*SSTTMAKESCL*RSLKLKLT*YSMMSMAMFYTNGTHPNSFTQ 375
>AW397749
Length = 272
Score = 87.0 bits (214), Expect = 1e-17
Identities = 45/89 (50%), Positives = 52/89 (57%), Gaps = 30/89 (33%)
Frame = -1
Query: 209 KSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILK------------------------ 244
+S AKWKIVVGHH I+SVGHHG+TQEL QLLPIL+
Sbjct: 269 ESTAKWKIVVGHHAIRSVGHHGDTQELINQLLPILQVTKS*G*RTSPLFPSSFT*LF*FY 90
Query: 245 ------SNNIDAYINGHDHCLQHIIDNER 267
+NNID Y+NGHDHCL+HI D ER
Sbjct: 89 FTFKFQANNIDFYMNGHDHCLEHISDTER 3
>TC225021 similar to UP|Q8LFV2 (Q8LFV2) Acid phosphatase type 5, partial
(35%)
Length = 817
Score = 70.1 bits (170), Expect = 1e-12
Identities = 38/83 (45%), Positives = 50/83 (59%)
Frame = +3
Query: 203 VDLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYINGHDHCLQHI 262
++LAL S AKW+IVVGHH I+SVGHHG+TQEL +LLPIL+ G DH L +
Sbjct: 3 LELALRGSTAKWRIVVGHHAIRSVGHHGDTQELINRLLPILQVTKG*G*RTGDDHGLSFL 182
Query: 263 IDNERHGEAISSHGTRK*SFTMT 285
+ + I R+ +FT T
Sbjct: 183 VPLPNSSDFILLSNFRQITFTFT 251
Score = 41.6 bits (96), Expect = 5e-04
Identities = 15/23 (65%), Positives = 19/23 (82%)
Frame = +2
Query: 245 SNNIDAYINGHDHCLQHIIDNER 267
+NN+ Y+NGHDHCL+HI D ER
Sbjct: 230 ANNVHFYMNGHDHCLEHISDTER 298
>AW830317
Length = 410
Score = 66.6 bits (161), Expect = 1e-11
Identities = 31/49 (63%), Positives = 39/49 (79%)
Frame = +1
Query: 204 DLALVKSKAKWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDAYI 252
DLAL +S AKWKIVVGHHTI+S G HGNT EL +QLLPIL+ ++ ++
Sbjct: 1 DLALQQSNAKWKIVVGHHTIRSAGLHGNTDELVKQLLPILEVTLLNFFL 147
Score = 39.3 bits (90), Expect = 0.002
Identities = 15/19 (78%), Positives = 17/19 (88%)
Frame = +2
Query: 244 KSNNIDAYINGHDHCLQHI 262
++NNID YING DHCLQHI
Sbjct: 233 QANNIDLYINGQDHCLQHI 289
>BM954298
Length = 397
Score = 47.8 bits (112), Expect = 7e-06
Identities = 30/61 (49%), Positives = 39/61 (63%)
Frame = +3
Query: 14 IFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKGNYNQSLVAHQMGI 73
I+ V L LLS S++ L R EH +K SL+ +V+GDWGRKG YNQS VA Q+ I
Sbjct: 12 IWMGFVSLCLLS--VSVSGLLQRLEHPVKADG-SLSLMVIGDWGRKGTYNQSQVATQVFI 182
Query: 74 V 74
+
Sbjct: 183L 185
>AW734779
Length = 450
Score = 32.0 bits (71), Expect = 0.39
Identities = 23/70 (32%), Positives = 33/70 (46%), Gaps = 3/70 (4%)
Frame = +1
Query: 79 NIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPSLQK--IWYSVLGNHDYRGDVD-A 135
N DF+ TGDN + G P ES + P ++ W +VLGNHD +D
Sbjct: 256 NPDFLAFTGDNIF-----GSSSPDAAESLFRAF-GPVMESGLPWAAVLGNHDQESTMDRE 417
Query: 136 QLSSILRQKD 145
+L S++ D
Sbjct: 418 ELMSLISLMD 447
>BM953834
Length = 421
Score = 31.6 bits (70), Expect = 0.51
Identities = 23/73 (31%), Positives = 34/73 (46%), Gaps = 8/73 (10%)
Frame = +2
Query: 201 KNVDLALVKSKAKWKIV-VGHHT-----IKSVGHH--GNTQELEQQLLPILKSNNIDAYI 252
++ DL + S WK +GH+T ++ G GN EL QQLL L NN+
Sbjct: 86 QDADLHALMSHLTWKGGNLGHYTWDGQLLRKKGKLVVGNDAELRQQLLEWLDINNLSFCW 265
Query: 253 NGHDHCLQHIIDN 265
H H H +++
Sbjct: 266 EHHQHLFSHFLND 304
>TC209639
Length = 1451
Score = 30.4 bits (67), Expect = 1.1
Identities = 24/75 (32%), Positives = 34/75 (45%), Gaps = 2/75 (2%)
Frame = +1
Query: 56 WGRKGNYNQSLVAHQMGIVGDNLNIDFVISTGDNFYKDGLEGVDDPTFYESFVNIYTAPS 115
WG + + N V M V N N DFVI GD + + + +++ TAP+
Sbjct: 268 WGPRQDLNSIRV---MSTVLHNENPDFVIYLGDVITANNIMIANASLYWDQA----TAPA 426
Query: 116 LQK--IWYSVLGNHD 128
+ W SV GNHD
Sbjct: 427 RNRGIPWASVFGNHD 471
>TC220622 similar to UP|Q8S2H5 (Q8S2H5) Diphosphonucleotide phosphatase-like
protein, partial (29%)
Length = 966
Score = 30.4 bits (67), Expect = 1.1
Identities = 26/116 (22%), Positives = 47/116 (40%), Gaps = 2/116 (1%)
Frame = +2
Query: 144 KDSRWVCLRSFILDGGIVEFFFVDTTPFIEKYFTDPKEHTYDWNGVLPRESYRAELLKNV 203
KD W + ++ G V F + T EH DW+ S + E ++
Sbjct: 113 KDKPW-----YSIEQGSVHFTVIST------------EH--DWS----ENSEQYEWVQKD 223
Query: 204 DLALVKSKAKWKIVVGHHTIKSVGHH--GNTQELEQQLLPILKSNNIDAYINGHDH 257
++ + K W I +GH + + H + + + + P+L N +D + GH H
Sbjct: 224 MASVNRQKTPWLIFMGHRPMYTTNHGFLPSENKFMEAVEPLLLENKVDLVLFGHVH 391
>BF068666
Length = 405
Score = 29.6 bits (65), Expect = 1.9
Identities = 19/57 (33%), Positives = 28/57 (48%), Gaps = 3/57 (5%)
Frame = +2
Query: 197 AELLKNVDLALVKSKA---KWKIVVGHHTIKSVGHHGNTQELEQQLLPILKSNNIDA 250
A LL ++ L+ + +A K K H T + H N QEL+ L+ I K +DA
Sbjct: 173 ANLLPDMKLSTLSKRAASIKIKSSSNHQTFSASTQHNNKQELDASLVVIRKVMKMDA 343
>TC205682 similar to GB|AAM67420.1|21689597|AY122887 AT5g55940/MYN21_5
{Arabidopsis thaliana;} , partial (62%)
Length = 992
Score = 29.6 bits (65), Expect = 1.9
Identities = 19/59 (32%), Positives = 30/59 (50%)
Frame = -2
Query: 2 AWCLKQRMLSPVIFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKG 60
AW ++M SP+ F + L LS+ S A K P + L + S+ + +WGR+G
Sbjct: 385 AWGK*RQMWSPIFF--ATLPSSLSSDLSFA*K*PTMFNPLAEKYSSIMMRDISNWGRRG 215
>TC205683 similar to GB|AAM67420.1|21689597|AY122887 AT5g55940/MYN21_5
{Arabidopsis thaliana;} , partial (96%)
Length = 1191
Score = 29.6 bits (65), Expect = 1.9
Identities = 19/59 (32%), Positives = 30/59 (50%)
Frame = -3
Query: 2 AWCLKQRMLSPVIFPASVLLRLLSNHSSIAEKLPRFEHHLKPQQQSLNFLVVGDWGRKG 60
AW ++M SP+ F + L LS+ S A K P + L + S+ + +WGR+G
Sbjct: 451 AWGK*RQMWSPIFF--ATLPSSLSSDLSFA*K*PTMFNPLAEKYSSIMMRDISNWGRRG 281
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.327 0.141 0.450
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,435,818
Number of Sequences: 63676
Number of extensions: 244875
Number of successful extensions: 1441
Number of sequences better than 10.0: 47
Number of HSP's better than 10.0 without gapping: 1432
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1436
length of query: 322
length of database: 12,639,632
effective HSP length: 97
effective length of query: 225
effective length of database: 6,463,060
effective search space: 1454188500
effective search space used: 1454188500
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146550.4


