

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146343.3 - phase: 0
(670 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
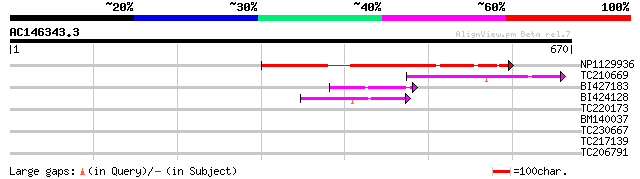
Score E
Sequences producing significant alignments: (bits) Value
NP1129936 CHX3 [Glycine max] 253 2e-67
TC210669 similar to UP|Q9FFR9 (Q9FFR9) Na+/H+ antiporter-like pr... 92 7e-19
BI427183 72 8e-13
BI424128 similar to GP|10178015|db Na+/H+ antiporter-like protei... 61 2e-09
TC220173 33 0.53
BM140037 32 0.91
TC230667 weakly similar to UP|AGA1_YEAST (P32323) A-agglutinin a... 30 4.5
TC217139 weakly similar to UP|Q6K6I0 (Q6K6I0) Nucellin-like aspa... 29 7.7
TC206791 similar to UP|Q8L4Y5 (Q8L4Y5) Actin-related protein 7, ... 29 7.7
>NP1129936 CHX3 [Glycine max]
Length = 1125
Score = 253 bits (646), Expect = 2e-67
Identities = 137/301 (45%), Positives = 187/301 (61%)
Frame = +1
Query: 301 LSSEALSVLSINVLVIGTLARIGVKYLYDPSRKYAGYQKRNILSLKPNSELKIVSCILKP 360
+S V+ IN++VI ++ + VK +YDPSRKYAGY KRNI SLKP+SEL++V+C+ K
Sbjct: 301 ISGPTYGVMMINIMVIASIVKWSVKLIYDPSRKYAGYPKRNIASLKPDSELRVVACLHKT 480
Query: 361 SHIIPIKNVLDICSPTSSNPLVIHILHLLELVGRSSPVFISHRLQERVGSSSHTFSEAVI 420
H +K+ LD+C PT+ +P S+ ++S+ +I
Sbjct: 481 HHASAVKDCLDLCCPTTEDP--------------------------NCSSNHKSYSDDII 582
Query: 421 VTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLASIIILPFHLRWSEDGSVES 480
+ FDL+EHDN G + YTAISP MH+D+C+LALDK+ASIIILPFHLRWS DG++ES
Sbjct: 583 LAFDLYEHDNMGAVTAHVYTAISPPSLMHEDVCHLALDKVASIIILPFHLRWSGDGAIES 762
Query: 481 ADETTRSLNTKVLERAPCSVAILVNRGHSSPFNHNENSKQIAMIFLGGSDDREALCLAKR 540
+ R+LN K+LE APCSV ILV R H ++ Q+AMIFLGG+DDREALCLAKR
Sbjct: 763 DYKNARALNCKLLEIAPCSVGILVGRSAI----HCDSFIQVAMIFLGGNDDREALCLAKR 930
Query: 541 TIKEDTYHLVVYHLVSTIKNDEFTSWDVMLDDELLKGVKGVYGSVDNVTYEKVEVENTSD 600
+ +LVVYHLV ++ + + D E LK V + + NV+Y+KV V +
Sbjct: 931 VTRNPRVNLVVYHLV---PKEQTPDVEYIQDKEALKHVMKPH--LGNVSYQKVMVNGGPE 1095
Query: 601 T 601
T
Sbjct: 1096T 1098
>TC210669 similar to UP|Q9FFR9 (Q9FFR9) Na+/H+ antiporter-like protein,
partial (16%)
Length = 952
Score = 92.0 bits (227), Expect = 7e-19
Identities = 66/201 (32%), Positives = 101/201 (49%), Gaps = 12/201 (5%)
Frame = +3
Query: 475 DGSVESADETTRSLNTKVLERAPCSVAILVNR--GHSSPFNHNENSKQIAMIFLGGSDDR 532
DGS+ R +N +VLE APCSV I V+R G +S + + S ++ ++F GG DDR
Sbjct: 12 DGSLNITRNDXRWVNKRVLEHAPCSVGIFVDRGLGGTSHVSASNVSYRVTVLFFGGGDDR 191
Query: 533 EALCLAKRTIKEDTYHLVVYHLVSTIKND-EFTSWDV---------MLDDELLKGVKGVY 582
EAL R + L+V V N+ E DV D+E L K
Sbjct: 192 EALAYGARMAEHPGIRLLVIRFVGEPMNEGEIVRVDVGDSTGTKLISQDEEFLDEFKAKI 371
Query: 583 GSVDNVTYEKVEVENTSDTTEFISDIAIQHDFIIVGRRNGIKSPQTQALASWTEYPELGV 642
+ D++ YE+ V++ ++T I ++ + F++ R + + + +E PELG
Sbjct: 372 ANDDSIIYEEKVVKDGAETVAIICELNSCNLFLVGSR----PASEVASAMKRSECPELGP 539
Query: 643 LGDLLASPDTNTKASILVVQQ 663
+G LLAS D T AS+LV+QQ
Sbjct: 540 VGGLLASQDYPTTASVLVMQQ 602
>BI427183
Length = 421
Score = 72.0 bits (175), Expect = 8e-13
Identities = 40/105 (38%), Positives = 59/105 (56%)
Frame = +2
Query: 383 IHILHLLELVGRSSPVFISHRLQERVGSSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAI 442
+ I+HL+EL RSS + ++ + GSS + E + F H G SV + T I
Sbjct: 44 LFIMHLVELTERSSSIILAQNTDNKSGSSHVEWLEQLYRAFQA--HSQLGQVSVQSKTTI 217
Query: 443 SPVRFMHDDICYLALDKLASIIILPFHLRWSEDGSVESADETTRS 487
S + MHDDIC++A +K+ ++IILPFH RW + VE +E S
Sbjct: 218 SSLSTMHDDICHVADEKMVTMIILPFHKRWKK---VEMENEEENS 343
>BI424128 similar to GP|10178015|db Na+/H+ antiporter-like protein
{Arabidopsis thaliana}, partial (16%)
Length = 421
Score = 60.8 bits (146), Expect = 2e-09
Identities = 41/136 (30%), Positives = 67/136 (49%), Gaps = 5/136 (3%)
Frame = +1
Query: 348 NSELKIVSCILKPSHIIPIKNVLDICSPT-SSNPLVIHILHLLELVGRSSPVFISHRLQE 406
NS+L+I++C +I + N+++ + L ++ +HL E RSS + + H+ +
Sbjct: 19 NSQLRILTCFHGARNIPSMINLIEASRGIRKGDALCVYAMHLKEFSERSSTILMVHKARR 198
Query: 407 RV----GSSSHTFSEAVIVTFDLFEHDNAGTASVSTYTAISPVRFMHDDICYLALDKLAS 462
H S VIV F+ + S+ AIS + +H+DIC A K A+
Sbjct: 199 NGLPFWNKGHHADSNHVIVAFEAYRQ--LSQVSIRPMIAISSMNNIHEDICATAERKGAA 372
Query: 463 IIILPFHLRWSEDGSV 478
+IILPFH DGS+
Sbjct: 373 VIILPFHKHQRLDGSL 420
>TC220173
Length = 567
Score = 32.7 bits (73), Expect = 0.53
Identities = 21/52 (40%), Positives = 31/52 (59%)
Frame = +1
Query: 612 HDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLLASPDTNTKASILVVQQ 663
+D IIVG+ G S A ++ ELG +GD+LAS + +S+LV+QQ
Sbjct: 253 YDLIIVGK--GQFSLSLVADLVDRQHEELGPIGDILASSTHDVVSSVLVIQQ 402
>BM140037
Length = 272
Score = 32.0 bits (71), Expect = 0.91
Identities = 20/71 (28%), Positives = 37/71 (51%), Gaps = 2/71 (2%)
Frame = -3
Query: 596 ENTSDTTEFISDI--AIQHDFIIVGRRNGIKSPQTQALASWTEYPELGVLGDLLASPDTN 653
+N ++ TE + + A HD +IVG++ +++ + ELG +GDL S
Sbjct: 264 KNATNITEEVLSVGKAKDHDLVIVGKQQ-LETTMLTNIDFRHGNEELGPIGDLFVSSGNG 88
Query: 654 TKASILVVQQQ 664
+S+LV+Q +
Sbjct: 87 ITSSLLVIQDR 55
>TC230667 weakly similar to UP|AGA1_YEAST (P32323) A-agglutinin attachment
subunit precursor, partial (5%)
Length = 865
Score = 29.6 bits (65), Expect = 4.5
Identities = 20/77 (25%), Positives = 35/77 (44%), Gaps = 9/77 (11%)
Frame = -2
Query: 202 PLGTEMIKQLELFCSWFLFPIFVTSCAMKIDLSVYVKSDYIYVWLGIIVAVHLFKMLV-- 259
P+ + QL+ F ++F P+ V KID+ V + I VW II + M++
Sbjct: 627 PILLPAMPQLKKFDTFFFIPMLV----YKIDMMVLMNRSKIRVWKIIITTIITMMMMIWA 460
Query: 260 -------TIGICWYCNM 269
T+G+ W ++
Sbjct: 459 VMIMMRMTMGMVWMMSL 409
>TC217139 weakly similar to UP|Q6K6I0 (Q6K6I0) Nucellin-like aspartic
protease-like, partial (61%)
Length = 967
Score = 28.9 bits (63), Expect = 7.7
Identities = 19/54 (35%), Positives = 26/54 (47%)
Frame = +1
Query: 93 SFNSIVTGIGTAFISSIKTDSHDNGDGKGTLKAFLNVFYYLCFMVVTPLVLRPI 146
S N +G AF+S + TD+ GD KGT+ YL + PLV + I
Sbjct: 595 SVNMTAVQVGHAFLS-LSTDTSTQGDRKGTIIDSGTTLAYLPEGIYEPLVYKII 753
>TC206791 similar to UP|Q8L4Y5 (Q8L4Y5) Actin-related protein 7, partial
(63%)
Length = 1117
Score = 28.9 bits (63), Expect = 7.7
Identities = 26/89 (29%), Positives = 37/89 (41%), Gaps = 8/89 (8%)
Frame = +1
Query: 217 WFLFPIFV--TSCAMKIDLSVYV--KSDYIYVWLGIIVAVHLFKMLVTIGICWYCNM--- 269
WF I + MK+DL ++ S+ IY H F + + I WYCNM
Sbjct: 622 WFSLKISI*LRQTMMKLDLPLFTGNASNSIYP--------HGFNNYLFLCIYWYCNML** 777
Query: 270 -PMADGLCLALMLSCKGVVDFCNNVFLHD 297
+ C + SC + CN + LHD
Sbjct: 778 LYIPSARCRVYLKSCSD-LH*CNLLTLHD 861
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.139 0.417
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 31,868,979
Number of Sequences: 63676
Number of extensions: 485079
Number of successful extensions: 3411
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 3374
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3403
length of query: 670
length of database: 12,639,632
effective HSP length: 104
effective length of query: 566
effective length of database: 6,017,328
effective search space: 3405807648
effective search space used: 3405807648
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.5 bits)
S2: 62 (28.5 bits)
Medicago: description of AC146343.3


