

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146341.8 + phase: 0
(268 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
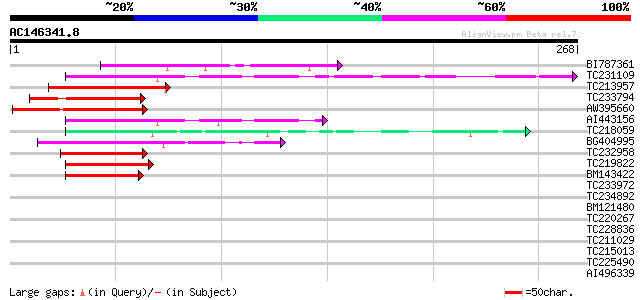
Score E
Sequences producing significant alignments: (bits) Value
BI787361 64 9e-11
TC231109 weakly similar to UP|Q9LUP7 (Q9LUP7) Gb|AAD25583.1, par... 57 7e-09
TC213957 similar to GB|AAP37713.1|30725382|BT008354 At3g23880 {A... 51 6e-07
TC233794 similar to GB|AAP37713.1|30725382|BT008354 At3g23880 {A... 51 6e-07
AW395660 49 2e-06
AI443156 47 1e-05
TC218059 weakly similar to UP|Q84KQ9 (Q84KQ9) F-box, partial (7%) 44 8e-05
BG404995 similar to GP|22831068|dbj OJ1131_E05.32 {Oryza sativa ... 43 1e-04
TC232958 42 4e-04
TC219822 homologue to UP|CENB_HUMAN (P07199) Major centromere au... 40 8e-04
BM143422 40 8e-04
TC233972 weakly similar to GB|AAP37713.1|30725382|BT008354 At3g2... 40 0.001
TC234892 weakly similar to UP|Q7QIA5 (Q7QIA5) AgCP3966 (Fragment... 36 0.016
BM121480 36 0.016
TC220267 similar to UP|Q6K235 (Q6K235) F-box-like protein, parti... 36 0.016
TC228836 similar to UP|Q6SH18 (Q6SH18) Lipase/esterase, Lip3/Bch... 35 0.046
TC211029 similar to UP|Q96VI4 (Q96VI4) Protease 1, partial (3%) 34 0.060
TC215013 homologue to UP|Q7XAU4 (Q7XAU4) Ethylene responsive pro... 34 0.079
TC225490 homologue to UP|Q7XAU4 (Q7XAU4) Ethylene responsive pro... 34 0.079
AI496339 weakly similar to GP|4995980|dbj| 68.2% identical to DR... 33 0.13
>BI787361
Length = 423
Score = 63.5 bits (153), Expect = 9e-11
Identities = 43/122 (35%), Positives = 65/122 (53%), Gaps = 8/122 (6%)
Frame = +3
Query: 44 ILRLICLSKSWNTLISDPTFVQKQLKRPPR--LILKPPCWVYPMRTIQSL-----PITRL 96
++R +S++WN+LI DPTFV+ L+R P+ +L +Y Q + I RL
Sbjct: 6 LMRFRYVSETWNSLIFDPTFVKLHLERSPKNTHVLLEFQAIYDRDVGQQVGVAPCSIRRL 185
Query: 97 LKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSDNKEY-HNYWFRIWNPATGTR 155
++NPS T+ D C + SCNGL+C C D +E+ +R+WNPATG
Sbjct: 186 VENPSFTI--DDCLT--LFKHTNSIFGSCNGLVCMTKCFDVREFEEECQYRLWNPATGIM 353
Query: 156 SK 157
S+
Sbjct: 354 SE 359
>TC231109 weakly similar to UP|Q9LUP7 (Q9LUP7) Gb|AAD25583.1, partial (8%)
Length = 1061
Score = 57.4 bits (137), Expect = 7e-09
Identities = 64/243 (26%), Positives = 103/243 (42%), Gaps = 1/243 (0%)
Frame = +3
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQL-KRPPRLILKPPCWVYPM 85
LP E++ +ILS L VK+++RL K W ++I FV L K LIL+ +Y
Sbjct: 93 LPVEVVTEILSRLPVKSVIRLRSTCKWWRSIIDSRHFVLFHLNKSHSSLILRHRSHLY-- 266
Query: 86 RTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSDNKEYHNYWF 145
SL + +NP CY +++ KV+ S NGLLC +D+
Sbjct: 267 ----SLDLKSPEQNPVELSHPLMCY-----SNSIKVLGSSNGLLCISNVADD-------I 398
Query: 146 RIWNPATGTRSKALGTNHDYNLQLRSLRFNFGSEAFGSCYYDYRRRYLLGCLKFTFGCGI 205
+WNP R + ++ SL F + +G ++ Y L L T+ +
Sbjct: 399 ALWNPF--LRKHRILPADRFHRPQSSL---FAARVYGFGHHSPSNDYKL--LSITYFVDL 557
Query: 206 LSGTYKTVEFRAKGDEENKYGPWRSQVRILSLSDDCWRNIDSFPVIPLISPFNNGVYLSG 265
T+ SQV++ +L D W+N+ S P L GV++SG
Sbjct: 558 QKRTFD------------------SQVQLYTLKSDSWKNLPSMP-YALCCARTMGVFVSG 680
Query: 266 TIY 268
+++
Sbjct: 681 SLH 689
>TC213957 similar to GB|AAP37713.1|30725382|BT008354 At3g23880 {Arabidopsis
thaliana;} , partial (9%)
Length = 696
Score = 50.8 bits (120), Expect = 6e-07
Identities = 30/60 (50%), Positives = 38/60 (63%), Gaps = 2/60 (3%)
Frame = +3
Query: 19 KKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRPP--RLIL 76
+K TLS+ LP ELI +IL V+++LR C+ KSW +LISDP F L P RLIL
Sbjct: 9 EKHTLSVTLPLELIREILLRSPVRSVLRFKCVCKSWLSLISDPQFTHFDLAASPTHRLIL 188
>TC233794 similar to GB|AAP37713.1|30725382|BT008354 At3g23880 {Arabidopsis
thaliana;} , partial (9%)
Length = 449
Score = 50.8 bits (120), Expect = 6e-07
Identities = 28/55 (50%), Positives = 38/55 (68%)
Frame = +1
Query: 10 KKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFV 64
K+ K K+N+ LT LP+ELI +IL L VK++LR C+ KS+ +LISDP FV
Sbjct: 67 KRSMKKKQNQSLTT---LPQELIREILLRLPVKSLLRFKCVCKSFLSLISDPQFV 222
>AW395660
Length = 381
Score = 49.3 bits (116), Expect = 2e-06
Identities = 27/64 (42%), Positives = 41/64 (63%)
Frame = +3
Query: 2 RYSKLMSIKKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDP 61
R S L +K+ ++ L L FLP+EL+ +ILS L VK++L+ C+ KSW +LI DP
Sbjct: 192 RTSPLPPSSVQKQQGMSESLPLP-FLPDELVVEILSRLPVKSLLQFRCVCKSWMSLIYDP 368
Query: 62 TFVQ 65
F++
Sbjct: 369 YFMK 380
>AI443156
Length = 414
Score = 46.6 bits (109), Expect = 1e-05
Identities = 41/126 (32%), Positives = 59/126 (46%), Gaps = 2/126 (1%)
Frame = +3
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQL-KRPPRLILKPPCWVYPM 85
LP E++ +ILS L VK+++RL K W ++I F+ L K LIL+ +Y
Sbjct: 90 LPVEVVTEILSRLPVKSVIRLRSTCKWWRSIIDSRHFILFHLNKSHTSLILRHRSQLY-- 263
Query: 86 RTIQSLPITRLL-KNPSITVSGDFCYDGGGLNDNCKVVISCNGLLCFIFCSDNKEYHNYW 144
SL + LL NP CY +++ KV+ S NGLLC D+
Sbjct: 264 ----SLDLKSLLDPNPFELSHPLMCY-----SNSIKVLGSSNGLLCISNVXDD------- 395
Query: 145 FRIWNP 150
+WNP
Sbjct: 396 IALWNP 413
>TC218059 weakly similar to UP|Q84KQ9 (Q84KQ9) F-box, partial (7%)
Length = 1185
Score = 43.9 bits (102), Expect = 8e-05
Identities = 60/235 (25%), Positives = 95/235 (39%), Gaps = 15/235 (6%)
Frame = +3
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQK--------QLKRPPRLILKP 78
LP EL++ +LS L K +L C+ KSW LI+DP FV Q + L+++
Sbjct: 66 LPGELVSNVLSRLPSKVLLLCKCVCKSWFDLITDPHFVSNYYVVYNSLQSQEEHLLVIRR 245
Query: 79 PCWVYPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCK----VVISCNGLLCFIFC 134
P + ++T S+ ++ +P VS D N + K ++ CNG+ F
Sbjct: 246 P-FFSGLKTYISV-LSWNTNDPKKHVSSDVLNPPYEYNSDHKYWTEILGPCNGI---YFL 410
Query: 135 SDNKEYHNYWFRIWNPATGTRSKALGTNHDYNLQLRSLRFNFGSEAFGSCYYDYRRRYLL 194
N + NP+ G KAL +H F S + DY
Sbjct: 411 EGNPNV------LMNPSLG-EFKALPKSH------------FTSPHGTYTFTDYA----- 518
Query: 195 GCLKFTFGCGILSGTYKTVEFR---AKGDEENKYGPWRSQVRILSLSDDCWRNID 246
FG + YK V + K +E + G W ++ + SL+ + WR +D
Sbjct: 519 -----GFGFDPKTNDYKVVVLKDLWLKETDEREIGYWSAE--LYSLNSNSWRKLD 662
>BG404995 similar to GP|22831068|dbj OJ1131_E05.32 {Oryza sativa (japonica
cultivar-group)}, partial (8%)
Length = 394
Score = 43.1 bits (100), Expect = 1e-04
Identities = 38/123 (30%), Positives = 58/123 (46%), Gaps = 6/123 (4%)
Frame = +1
Query: 14 KDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLKRP-- 71
++ K LT+S LP E++ +ILS L V+++LR SKSW +LI L R
Sbjct: 1 EETKPVLLTMSDHLPREVLTEILSRLPVRSLLRFRSTSKSWKSLIDSQHLNWLHLTRSLT 180
Query: 72 ----PRLILKPPCWVYPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNG 127
LIL+ +Y +L L +P + CY +++ ++ SCNG
Sbjct: 181 LASNTSLILRVDSDLY-QTNFPTLDPPVSLNHPLM------CY-----SNSITLLGSCNG 324
Query: 128 LLC 130
LLC
Sbjct: 325 LLC 333
>TC232958
Length = 758
Score = 41.6 bits (96), Expect = 4e-04
Identities = 20/41 (48%), Positives = 28/41 (67%)
Frame = +1
Query: 25 MFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQ 65
M LP++LI +IL L VK+++R + KSW LISDP F +
Sbjct: 55 MVLPQDLITEILLRLPVKSLVRFKSVCKSWLFLISDPRFAK 177
>TC219822 homologue to UP|CENB_HUMAN (P07199) Major centromere autoantigen B
(Centromere protein B) (CENP-B), partial (5%)
Length = 710
Score = 40.4 bits (93), Expect = 8e-04
Identities = 19/42 (45%), Positives = 27/42 (64%)
Frame = +1
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQL 68
LPE L+ +IL + V L+L C+ K W +L+ DP FV+K L
Sbjct: 196 LPEGLMIEILVWIRVSNPLQLRCVCKRWKSLVVDPQFVKKHL 321
>BM143422
Length = 424
Score = 40.4 bits (93), Expect = 8e-04
Identities = 20/37 (54%), Positives = 25/37 (67%)
Frame = +1
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTF 63
LP +LI IL L VK++ R C+ KSW +LISDP F
Sbjct: 229 LPLDLIELILLKLPVKSVTRFKCVCKSWLSLISDPQF 339
>TC233972 weakly similar to GB|AAP37713.1|30725382|BT008354 At3g23880
{Arabidopsis thaliana;} , partial (10%)
Length = 631
Score = 40.0 bits (92), Expect = 0.001
Identities = 33/110 (30%), Positives = 54/110 (49%), Gaps = 6/110 (5%)
Frame = -2
Query: 4 SKLMSIKKKKKD--KKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDP 61
+KLM+I + ++ + +FLP+E+I +IL L VK++ + KS +LIS+P
Sbjct: 615 AKLMNINYXXXSIGETRRQKMVXLFLPQEIIIEILLRLPVKSLPSFKFVCKS*LSLISNP 436
Query: 62 TFVQKQLKR----PPRLILKPPCWVYPMRTIQSLPITRLLKNPSITVSGD 107
F + +R +L+ P WV +QS RL+ P S D
Sbjct: 435 HFAKWHFERNAAHTEKLLFVNPSWV---PKVQSFMTIRLVLPPQTLFSLD 295
>TC234892 weakly similar to UP|Q7QIA5 (Q7QIA5) AgCP3966 (Fragment), partial
(4%)
Length = 370
Score = 36.2 bits (82), Expect = 0.016
Identities = 19/40 (47%), Positives = 28/40 (69%)
Frame = -2
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQK 66
LP+ELI I L + I+RL L+KS++ +ISD TFV++
Sbjct: 243 LPQELIQNIFLSLVLPEIIRLKLLNKSFSRIISDNTFVRQ 124
>BM121480
Length = 917
Score = 36.2 bits (82), Expect = 0.016
Identities = 55/210 (26%), Positives = 87/210 (41%), Gaps = 3/210 (1%)
Frame = +3
Query: 10 KKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQLK 69
K+ K K+N+ LT LP+ELI +IL L VK++LR C+ + T D +
Sbjct: 24 KRSMKKKQNQSLTT---LPQELIREILLRLPVKSLLRFKCV--LFKTCSRDVVY------ 170
Query: 70 RPPRLILKPPCWVYPMRTIQSLPITRLLKNPSITVSGDFCYDGGGLNDNCKVVISCNGLL 129
+P+ + S+P RL D G+ K++ SC GL+
Sbjct: 171 -------------FPL-PLPSIPCLRL--------------DDFGIRP--KILGSCRGLV 260
Query: 130 CFIFCSDNKEYHNYWFRIWNPATGTRSKALGTNHDYNLQLRSLRFNFGSEAFGSCYYDYR 189
++ D+ +WNP+ G R K L +Y + S + FG Y + +
Sbjct: 261 -LLYYDDSAN-----LILWNPSLG-RHKXL---PNYRDDITSFPYGFG-------YDESK 389
Query: 190 RRYLL---GCLKFTFGCGILSGTYKTVEFR 216
YLL G KF G ++KT ++
Sbjct: 390 DEYLLILIGLPKFGPETGADIYSFKTESWK 479
>TC220267 similar to UP|Q6K235 (Q6K235) F-box-like protein, partial (7%)
Length = 586
Score = 36.2 bits (82), Expect = 0.016
Identities = 14/38 (36%), Positives = 26/38 (67%)
Frame = +2
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFV 64
LP++L+ +IL+ L + +I R C+SK W+ +++ FV
Sbjct: 338 LPDDLLERILAYLPIASIFRAGCVSKRWHEIVNSERFV 451
>TC228836 similar to UP|Q6SH18 (Q6SH18) Lipase/esterase, Lip3/BchO family,
partial (9%)
Length = 702
Score = 34.7 bits (78), Expect = 0.046
Identities = 16/59 (27%), Positives = 35/59 (59%)
Frame = +3
Query: 9 IKKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQKQ 67
+ K++ K ++ S+ LP++++ I S LD+K+++ + + +SWN +D +KQ
Sbjct: 435 VAHKRRCKAHQVEKGSLPLPQDILVLIFSFLDMKSLVSMGLVCRSWNIAANDNHLWEKQ 611
>TC211029 similar to UP|Q96VI4 (Q96VI4) Protease 1, partial (3%)
Length = 858
Score = 34.3 bits (77), Expect = 0.060
Identities = 20/61 (32%), Positives = 36/61 (58%)
Frame = +3
Query: 3 YSKLMSIKKKKKDKKNKKLTLSMFLPEELIAQILSLLDVKTILRLICLSKSWNTLISDPT 62
+S L S+ + + +K ++T L +LI ILSLL + T++R + K W+++IS +
Sbjct: 96 FSLLFSLFQLQHEKMMTEITN---LSTDLIELILSLLPIPTLIRASTVCKLWHSIISSSS 266
Query: 63 F 63
F
Sbjct: 267 F 269
>TC215013 homologue to UP|Q7XAU4 (Q7XAU4) Ethylene responsive protein,
complete
Length = 1999
Score = 33.9 bits (76), Expect = 0.079
Identities = 20/47 (42%), Positives = 25/47 (52%), Gaps = 4/47 (8%)
Frame = -3
Query: 150 PATGTRSKALGTN----HDYNLQLRSLRFNFGSEAFGSCYYDYRRRY 192
PA GT +G N HD++ QLRSL F E F +YRR+Y
Sbjct: 302 PAGGTCRYEVGDNRTTTHDHSTQLRSLLFRIKVETFFHPMENYRRKY 162
>TC225490 homologue to UP|Q7XAU4 (Q7XAU4) Ethylene responsive protein,
partial (64%)
Length = 865
Score = 33.9 bits (76), Expect = 0.079
Identities = 20/47 (42%), Positives = 25/47 (52%), Gaps = 4/47 (8%)
Frame = -3
Query: 150 PATGTRSKALGTN----HDYNLQLRSLRFNFGSEAFGSCYYDYRRRY 192
PA GT +G N HD++ QLRSL F E F +YRR+Y
Sbjct: 170 PAGGTCRYEVGDNRTTTHDHSTQLRSLLFRIKVETFFHPMENYRRKY 30
>AI496339 weakly similar to GP|4995980|dbj| 68.2% identical to DR2 gene of
strain U1102 of HHV-6~US22 gene family {Human
herpesvirus 6}, partial (3%)
Length = 365
Score = 33.1 bits (74), Expect = 0.13
Identities = 17/40 (42%), Positives = 27/40 (67%)
Frame = +3
Query: 27 LPEELIAQILSLLDVKTILRLICLSKSWNTLISDPTFVQK 66
LP+ELI I L + I+RL ++KS++ +ISD FV++
Sbjct: 93 LPQELIQNIFLSLVLPEIIRLKLVNKSFSRIISDHAFVRQ 212
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.142 0.455
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 16,030,818
Number of Sequences: 63676
Number of extensions: 281499
Number of successful extensions: 1845
Number of sequences better than 10.0: 79
Number of HSP's better than 10.0 without gapping: 1800
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1832
length of query: 268
length of database: 12,639,632
effective HSP length: 96
effective length of query: 172
effective length of database: 6,526,736
effective search space: 1122598592
effective search space used: 1122598592
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 58 (26.9 bits)
Medicago: description of AC146341.8


