

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146341.2 - phase: 0 /pseudo
(875 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
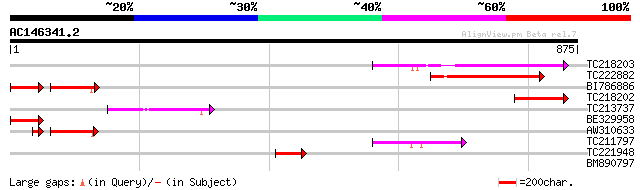
Score E
Sequences producing significant alignments: (bits) Value
TC218203 weakly similar to GB|AAB62841.1|2252842|IG005I10 A_IG00... 202 7e-52
TC222882 177 1e-44
BI786886 similar to GP|2252842|gb|A A_IG005I10.23 gene product {... 94 3e-39
TC218202 weakly similar to GB|AAB62841.1|2252842|IG005I10 A_IG00... 87 4e-17
TC213737 75 1e-13
BE329958 71 2e-12
AW310633 61 5e-12
TC211797 67 3e-11
TC221948 43 5e-04
BM890797 37 0.038
>TC218203 weakly similar to GB|AAB62841.1|2252842|IG005I10 A_IG005I10.23
{Arabidopsis thaliana;} , partial (9%)
Length = 1324
Score = 202 bits (513), Expect = 7e-52
Identities = 148/316 (46%), Positives = 175/316 (54%), Gaps = 13/316 (4%)
Frame = +3
Query: 560 LLSC*R*DSY*RGCRICKRDCYCRIFLTISWPQCRKC*PKP*YLRKF*LCRIFSMFGAGG 619
LLSC R*DSY*R CRI KR+ I W RK PK *YLR * CR+F MF +G
Sbjct: 192 LLSCQR*DSY*R*CRIYKREK*FGIVFGSKWFHHRKG-PKY*YLRNS*WCRMF*MFESGH 368
Query: 620 I--------SVCTTILH-----KLMIYVRVKYNFSPYKITISQTILRG*RLLWKLQMSTC 666
+V T+I H + + R + I I R * W LQ+ST
Sbjct: 369 TRREPIIISTVITSIFHH*EN*RARKWYRCIWATESC-ICS*YIIFR**LWPWALQIST* 545
Query: 667 *STSTTTACSIRRTRFFSCESNQKRKILYRRKRIDI*LHQCSSASLWIDKRSIVDEMPFL 726
+ +KIL+ RK IDI*LHQ A LW++ RS+VDEMPFL
Sbjct: 546 --------------------TI**KKILF*RK*IDI*LHQSCLACLWLNYRSVVDEMPFL 665
Query: 727 RQDARPFLV*SGRVITKPAMP*SETPIRMYQ*NSHRSLLRLLRRLTFCIICKS*HKANSK 786
RQD FLV*SGR + +PA+ SE PI +YQ*+SH L LL LT +CKS*HKANSK
Sbjct: 666 RQDTGSFLV*SGRALLEPALQQSEAPI*LYQ*SSHGDLSALLWGLTLGFVCKS*HKANSK 845
Query: 787 YENGYSHGLGRSVLVFTSHAPAAHVGKNC*ERHGKMWIMDGPSI*SRNRWF*TRRCHSIR 846
YE GY GLGRSVL + S AP H N *ERHGK MDGP * RN WF* HS
Sbjct: 846 YEKGYFKGLGRSVLAYASLAPTTHFRANR*ERHGKKGDMDGPWT*HRNNWF*DG*GHSC* 1025
Query: 847 SDGRYNTWLCK*ESRR 862
+DGR++ C *+SR+
Sbjct: 1026TDGRHHFEPCD*KSRK 1073
>TC222882
Length = 656
Score = 177 bits (450), Expect = 1e-44
Identities = 109/177 (61%), Positives = 124/177 (69%), Gaps = 1/177 (0%)
Frame = +2
Query: 650 TILRG*RLLWKLQMSTC*STSTTTACSIRRTRFFSCESNQKRKILYRRKRIDI*LHQCSS 709
T+LRG* W LQ+ST *STST A +I RT+ FS ESNQ R+IL RK +DI*LHQ S
Sbjct: 131 TVLRG*YQPWLLQISTG*STST--AAAI*RTKLFSFESNQ*RQILSERK*MDI*LHQSSP 304
Query: 710 ASLWIDKRSIVDEMPFLRQDARPFLV*SG-RVITKPAMP*SETPIRMYQ*NSHRSLLRLL 768
ASLWID RSI DEM FLRQD R FLV*SG RV+TKPA+ *SE ++Q* SH L LL
Sbjct: 305 ASLWIDCRSIADEMSFLRQDTRSFLV*SGNRVLTKPALQ*SEAHQ*LHQ*RSHGGLSELL 484
Query: 769 RRLTFCIICKS*HKANSKYENGYSHGLGRSVLVFTSHAPAAHVGKNC*ERHGKMWIM 825
R LT C+ICKS H A SKYE S L RSVL F S AP ++G+N *+RHG W M
Sbjct: 485 RGLTLCVICKSWH*AYSKYEEDDSQSLRRSVLAFPSLAPTTYLGQNY*KRHG*KWGM 655
>BI786886 similar to GP|2252842|gb|A A_IG005I10.23 gene product
{Arabidopsis thaliana}, partial (1%)
Length = 421
Score = 93.6 bits (231), Expect(2) = 3e-39
Identities = 41/51 (80%), Positives = 47/51 (91%)
Frame = +3
Query: 1 MAKKSQRRAVRYEKDKAGCMWGFISMLDFRHGHSTRKLIADKRRSSKHSEG 51
M KKSQRR V+YEKDK+GCMWGFI+M DFRHGHSTRK+IADKR+SSKH+ G
Sbjct: 21 MTKKSQRRPVQYEKDKSGCMWGFINMFDFRHGHSTRKMIADKRQSSKHAVG 173
Score = 87.8 bits (216), Expect(2) = 3e-39
Identities = 55/83 (66%), Positives = 60/83 (72%), Gaps = 8/83 (9%)
Frame = +2
Query: 64 CCAF*E*V*GVERYG*SLSWHF*RWRE*KTNSENYC**A*CEETHRRRDVH*SKCNEGYR 123
CCAF*E V* V ++G*SLS F +WRE*KTNS N C**A*CEETHRRRDVH* + NEGY
Sbjct: 173 CCAF*EQV*DVGQFG*SLSRQFWQWRE*KTNSSNCC**A*CEETHRRRDVH*PEYNEGYI 352
Query: 124 *F--------*KA*SLPEVRFQK 138
* KA*S EVR+QK
Sbjct: 353 *CPNRIKGIQIKA*SPFEVRYQK 421
>TC218202 weakly similar to GB|AAB62841.1|2252842|IG005I10 A_IG005I10.23
{Arabidopsis thaliana;} , partial (8%)
Length = 828
Score = 86.7 bits (213), Expect = 4e-17
Identities = 47/84 (55%), Positives = 56/84 (65%)
Frame = +2
Query: 779 S*HKANSKYENGYSHGLGRSVLVFTSHAPAAHVGKNC*ERHGKMWIMDGPSI*SRNRWF* 838
S*H+ANSKYE GY GLGRSVL + S AP AH NC*ERHGK MDGP * N WF*
Sbjct: 2 S*HEANSKYEKGYFKGLGRSVLAYPSLAPTAHFRANC*ERHGKKGDMDGPWT*CGNNWF* 181
Query: 839 TRRCHSIRSDGRYNTWLCK*ESRR 862
H+ +DG ++ C+*+SR+
Sbjct: 182 NG*GHTW*TDGGHHFEPCE*KSRK 253
>TC213737
Length = 512
Score = 75.1 bits (183), Expect = 1e-13
Identities = 72/170 (42%), Positives = 92/170 (53%), Gaps = 6/170 (3%)
Frame = +2
Query: 152 *FKFGCSIEI*NFASSALKEAI*R*S*CGYNN*RVL*SKRCLFHDA*R*WRSRKAWPTKP 211
*F+F C++EI* F +A K + R*S NN L CLFHD * W++ KP
Sbjct: 17 *FEFRCNLEI*IFT*TAFKTTVKR*SRFEQNNG*FLSC*SCLFHDE**SWQN**T--VKP 190
Query: 212 ETEACCVRKTFKRCDS*ICESDDIERQRSGGS*KVSLL**NNGSTSAYKFR*GIVSCIST 271
+ AC + K C+ *IC+SD+IE +RS ++ L * +GSTS YKFR* IVS ST
Sbjct: 191 K--ACYL*KP-S*CNP*ICKSDEIEWERSA*RWTIA*LT*THGSTSGYKFR*AIVS*TST 361
Query: 272 KSKSTCIEMCSRV*EFSSSERQR------FQLLGTRSWK*RTNQ*DC*PQ 315
+ T IE+ S V + S QR QLL T + TN DC PQ
Sbjct: 362 RP*FTLIEIHSGVGKCSRERWQRMQFSYQLQLL*T*TC*TETN*GDCQPQ 511
>BE329958
Length = 471
Score = 71.2 bits (173), Expect = 2e-12
Identities = 32/51 (62%), Positives = 38/51 (73%)
Frame = +3
Query: 1 MAKKSQRRAVRYEKDKAGCMWGFISMLDFRHGHSTRKLIADKRRSSKHSEG 51
MAK+SQR V YEKD++GCMWGFIS+ DFRH TRKLI D+ +KH G
Sbjct: 267 MAKRSQRFPVNYEKDQSGCMWGFISIFDFRHACFTRKLIVDRTHGNKHVVG 419
>AW310633
Length = 302
Score = 61.2 bits (147), Expect(2) = 5e-12
Identities = 42/82 (51%), Positives = 53/82 (64%), Gaps = 8/82 (9%)
Frame = -2
Query: 64 CCAF*E*V*GVERYG*SLSWHF*RWRE*KTNSENYC**A*CEETHRRRDVH*SKCNEGYR 123
CC + E V* VE++G* + F*+ RE*+TN++ C *A*CEETHRRRD H* N+G R
Sbjct: 247 CCTYQEQV*SVEQFG*RI*RQF*QRRE*ETNTDK*CR*A*CEETHRRRDDH*PG*NKGPR 68
Query: 124 *F*--------KA*SLPEVRFQ 137
* * +A* P+ RFQ
Sbjct: 67 *C*SRIKAIQIRA*RPPKDRFQ 2
Score = 28.5 bits (62), Expect(2) = 5e-12
Identities = 12/17 (70%), Positives = 14/17 (81%)
Frame = -3
Query: 35 TRKLIADKRRSSKHSEG 51
TRKLIAD+R SKH+ G
Sbjct: 297 TRKLIADRRHGSKHAVG 247
>TC211797
Length = 531
Score = 67.0 bits (162), Expect = 3e-11
Identities = 66/156 (42%), Positives = 83/156 (52%), Gaps = 12/156 (7%)
Frame = +1
Query: 561 LSC*R*DSY*RGCRICKRDCYCRIFLTISWPQCRKC*PKP*YLRKF*LCRIFSMFGAG-- 618
LSC*R*DSY*R RIC+R+ I L W + RK PK *YL * CR+F F +G
Sbjct: 67 LSC*R*DSY*RLYRICEREK*FGIVLESEWLRHRKG-PKY*YLGNS*WCRMF*AFESGHN 243
Query: 619 --GISVCTTILHKLMIY-----VRVKYNF--SPYKITISQTILRG*RLL-WKLQMSTC*S 668
++ +T + L + R Y + S I+ G* LL W LQM C*
Sbjct: 244 RREPAIISTAISSLFCH*EN*RARKWYRCI*ATESCICS*YIILG**LLFWALQM*AC*I 423
Query: 669 TSTTTACSIRRTRFFSCESNQKRKILYRRKRIDI*L 704
TST+ SI RT F+ + +RKIL+ RK IDI*L
Sbjct: 424 TSTSPTNSI*RT*LFTSRTI*QRKILF*RK*IDI*L 531
>TC221948
Length = 209
Score = 43.1 bits (100), Expect = 5e-04
Identities = 26/47 (55%), Positives = 30/47 (63%)
Frame = +1
Query: 411 RQCRNEIS**RSFLY*KNRKAFSRCY*RRQDWY**RFGSDCGA*KGY 457
R C NEIS**R FL+*KN K + C R+QD Y R +CG *K Y
Sbjct: 55 R*CWNEIS**RPFLH*KNYKTYV*CSERKQDRYNERL*IECGT*KWY 195
>BM890797
Length = 421
Score = 37.0 bits (84), Expect = 0.038
Identities = 20/52 (38%), Positives = 32/52 (61%), Gaps = 1/52 (1%)
Frame = +1
Query: 1 MAKKSQRRAVR-YEKDKAGCMWGFISMLDFRHGHSTRKLIADKRRSSKHSEG 51
MA++S R + YE GC W + + +FR GHS R+L+++ RR + H+ G
Sbjct: 136 MARRSPPRVNQTYE---TGCTWRMLRIFNFREGHSDRRLVSN-RRLNAHANG 279
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.374 0.166 0.667
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,503,134
Number of Sequences: 63676
Number of extensions: 658047
Number of successful extensions: 10364
Number of sequences better than 10.0: 20
Number of HSP's better than 10.0 without gapping: 3060
Number of HSP's successfully gapped in prelim test: 342
Number of HSP's that attempted gapping in prelim test: 7138
Number of HSP's gapped (non-prelim): 3775
length of query: 875
length of database: 12,639,632
effective HSP length: 106
effective length of query: 769
effective length of database: 5,889,976
effective search space: 4529391544
effective search space used: 4529391544
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 13 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 35 (21.5 bits)
S2: 63 (28.9 bits)
Medicago: description of AC146341.2


