

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.8 + phase: 0
(402 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
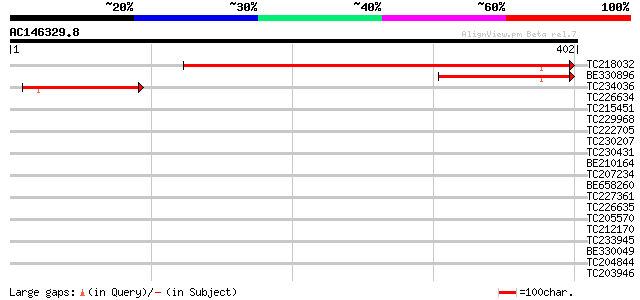
Score E
Sequences producing significant alignments: (bits) Value
TC218032 weakly similar to UP|Q9Y7Y4 (Q9Y7Y4) SPBC365.02c protei... 487 e-138
BE330896 105 5e-23
TC234036 87 1e-17
TC226634 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting... 34 0.13
TC215451 weakly similar to UP|Q94K07 (Q94K07) Nucleosome assembl... 33 0.23
TC229968 similar to UP|O64797 (O64797) T1F15.5 protein, partial ... 32 0.39
TC222705 similar to UP|Q6UK18 (Q6UK18) CaCBF1B, partial (54%) 31 1.1
TC230207 similar to UP|Q9LTR6 (Q9LTR6) Gb|AAF36750.1, partial (28%) 30 1.9
TC230431 weakly similar to UP|Q9FHQ4 (Q9FHQ4) Similarity to secr... 30 2.5
BE210164 homologue to GP|18086376|gb AT4g23460/F16G20_160 {Arabi... 30 2.5
TC207234 weakly similar to UP|Q9FH01 (Q9FH01) Similarity to CHP-... 29 3.3
BE658260 similar to GP|9757958|dbj mitochondrial carrier protein... 29 3.3
TC227361 similar to UP|Q9SD67 (Q9SD67) FtsH metalloprotease-like... 29 4.3
TC226635 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting... 29 4.3
TC205570 homologue to UP|Q93VJ8 (Q93VJ8) AT5g11200/F2I11_90 (AT5... 28 5.7
TC212170 28 5.7
TC233945 weakly similar to UP|Q8L5R4 (Q8L5R4) Tubulin folding co... 28 5.7
BE330049 homologue to GP|2252828|gb| A_IG005I10.5 gene product {... 28 7.4
TC204844 similar to UP|Q9LT85 (Q9LT85) Similarity to receptor pr... 28 7.4
TC203946 homologue to UP|Q9LN13 (Q9LN13) T6D22.2, partial (47%) 28 7.4
>TC218032 weakly similar to UP|Q9Y7Y4 (Q9Y7Y4) SPBC365.02c protein (Cox10
protein), partial (16%)
Length = 996
Score = 487 bits (1254), Expect = e-138
Identities = 235/279 (84%), Positives = 262/279 (93%), Gaps = 2/279 (0%)
Frame = +1
Query: 124 TSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLVLYSFVYTPLKQI 183
TS+RPLPSGRIT+PHAVGWASSVGLAGTALLATQTNMLAAGLAASNL+LY+FVYTPLKQI
Sbjct: 1 TSRRPLPSGRITIPHAVGWASSVGLAGTALLATQTNMLAAGLAASNLILYAFVYTPLKQI 180
Query: 184 HPVNTWVGAVVGAIPPLLGWAAASGDISLNSLILPAALYFWQIPHFMALAYMCREDYAAG 243
HP+NTWVGAVVGAIPPLLGWAAAS DISLN +ILPAALYFWQIPHFMALAY+CR+DYAAG
Sbjct: 181 HPINTWVGAVVGAIPPLLGWAAASNDISLNGMILPAALYFWQIPHFMALAYLCRDDYAAG 360
Query: 244 GFKMYSLADPSGRKTAMVAMRNSIYLIPLGFLAYDWGVTSGWFCLESTALTLAISAAAFS 303
GFKMYSLAD SG +TA+VA+RNSIYLIPLGFLAYDWG+TSGWFCLES ALTLAISAAAFS
Sbjct: 361 GFKMYSLADASGHRTALVALRNSIYLIPLGFLAYDWGLTSGWFCLESAALTLAISAAAFS 540
Query: 304 FYRDRTKERARRMFHASLLYLPVFMAGLLIHRRTDNQHFLEVNAEGFMKSPSSVETSEMD 363
FYR+RTKE+ARRMFHASLLYLPVFM+GLLIHRR++N+ FLE A+GF+KS SS E+SE+D
Sbjct: 541 FYRNRTKEKARRMFHASLLYLPVFMSGLLIHRRSENEQFLEDKAKGFVKSASSTESSELD 720
Query: 364 DKNGNQKIKGRQ--GTRARPPVAYASIAPFPFLPAPSYD 400
D NG+Q IK R+ T ARPPV+YAS+APFPFLPAPSY+
Sbjct: 721 DDNGDQNIKARRPYRTEARPPVSYASVAPFPFLPAPSYN 837
>BE330896
Length = 301
Score = 105 bits (261), Expect = 5e-23
Identities = 55/98 (56%), Positives = 69/98 (70%), Gaps = 2/98 (2%)
Frame = +2
Query: 305 YRDRTKERARRMFHASLLYLPVFMAGLLIHRRTDNQHFLEVNAEGFMKSPSSVETSEMDD 364
Y +RT +ARRMFHASLLYLPVFM+GLLI+RR++N+ FLE A+G +KS SS E +DD
Sbjct: 2 YINRTI*KARRMFHASLLYLPVFMSGLLIYRRSENEQFLEDKAKGLVKSASSTEFLHVDD 181
Query: 365 KNGNQKIKGRQ--GTRARPPVAYASIAPFPFLPAPSYD 400
NGN IK + T PPV A +AP F+ APSY+
Sbjct: 182 DNGNHNIKAMRPYRTETLPPVIDACVAPLSFVSAPSYN 295
>TC234036
Length = 422
Score = 87.0 bits (214), Expect = 1e-17
Identities = 48/92 (52%), Positives = 61/92 (66%), Gaps = 6/92 (6%)
Frame = +2
Query: 10 TLYSRLVNSS------SSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYGNCYSQLS 63
+ YS+ ++S SSS+ H ++F S + ++ + D SL RHYG CY +LS
Sbjct: 11 SFYSKFIDSHNLNCGRSSSNPHRVAASATSFDISSASASKATTDFESLTRHYGRCYWELS 190
Query: 64 KARLSLLVVATSGTGFVLGSGGAVDLSMLSYT 95
KARLS+LVVATSGTGFVLGSG AVDLS LS T
Sbjct: 191 KARLSMLVVATSGTGFVLGSGSAVDLSALSCT 286
>TC226634 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting protein
(Fragment), partial (51%)
Length = 1024
Score = 33.9 bits (76), Expect = 0.13
Identities = 31/109 (28%), Positives = 52/109 (47%), Gaps = 3/109 (2%)
Frame = -2
Query: 5 NWNCATLYSRLVNSSSSSSLHNFTHIRSTFSSSPSVSAS-KSNDLISLARHYGNCYSQLS 63
NWN + + SSSS+ +F S FSS S++ S S S A+ S
Sbjct: 483 NWNISASAFLSLTILSSSSICSFFFCLSIFSSPASLALSANSRSQASKAKRAFLNILIFS 304
Query: 64 KARLSLLVVATSGTGFVL--GSGGAVDLSMLSYTCLGTMMVAASASTLN 110
A +S+ ++A+ T +L S GA+ L + +++ L ++S T+N
Sbjct: 303 SATISIFLIASWTTSAILLHASWGAISLPLSAFSRLFGESSSSSWKTIN 157
>TC215451 weakly similar to UP|Q94K07 (Q94K07) Nucleosome assembly protein,
partial (20%)
Length = 777
Score = 33.1 bits (74), Expect = 0.23
Identities = 18/40 (45%), Positives = 27/40 (67%)
Frame = -1
Query: 7 NCATLYSRLVNSSSSSSLHNFTHIRSTFSSSPSVSASKSN 46
NC T + L++SSSSSS + + S+ SSS S+S+S S+
Sbjct: 213 NCYTFFLVLLSSSSSSSSSSSSSSSSSSSSSSSISSSSSS 94
>TC229968 similar to UP|O64797 (O64797) T1F15.5 protein, partial (13%)
Length = 902
Score = 32.3 bits (72), Expect = 0.39
Identities = 16/27 (59%), Positives = 19/27 (70%)
Frame = +3
Query: 17 NSSSSSSLHNFTHIRSTFSSSPSVSAS 43
+SSSSSS+H+ H FSSS S SAS
Sbjct: 84 SSSSSSSIHSLLHTLHNFSSSESASAS 164
>TC222705 similar to UP|Q6UK18 (Q6UK18) CaCBF1B, partial (54%)
Length = 745
Score = 30.8 bits (68), Expect = 1.1
Identities = 36/140 (25%), Positives = 59/140 (41%), Gaps = 3/140 (2%)
Frame = -1
Query: 9 ATLYSRLVNSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYGNCYSQLSKARLS 68
++L + V S+ L I S+ SSS SV+ + + + + G S S A
Sbjct: 658 SSLPKQCVGDMSTMFLRYSGTIPSSCSSSRSVAVAATLSATASSSFPGRNASAASAAAF* 479
Query: 69 L-LVVATSGTGFVLGSGGAVDLSMLSYTCLG-TMMVAASASTLNQV-FEIKNDAIMRRTS 125
+ L A +GTG + + L + + T A+ S + +V +I + + RTS
Sbjct: 478 MSLASAVAGTGSRDAESAKLRQADLPLSAIAATSCARAAISAVGKVPSQILDFLLGSRTS 299
Query: 126 QRPLPSGRITVPHAVGWASS 145
LP R+ P GW S
Sbjct: 298 HTHLPESRLLTPRYTGWRVS 239
>TC230207 similar to UP|Q9LTR6 (Q9LTR6) Gb|AAF36750.1, partial (28%)
Length = 823
Score = 30.0 bits (66), Expect = 1.9
Identities = 21/62 (33%), Positives = 35/62 (55%), Gaps = 3/62 (4%)
Frame = -2
Query: 19 SSSSSLHNFTHI---RSTFSSSPSVSASKSNDLISLARHYGNCYSQLSKARLSLLVVATS 75
S+S+SLH+F H+ S+ SSSPS S+ + ++ N +S L L+ LV+ +S
Sbjct: 594 SASTSLHSFEHVSPASSSSSSSPSSGQESSSGINTIKLFLLNSFSLL----LTSLVIKSS 427
Query: 76 GT 77
+
Sbjct: 426 SS 421
>TC230431 weakly similar to UP|Q9FHQ4 (Q9FHQ4) Similarity to secretory
protein, partial (29%)
Length = 396
Score = 29.6 bits (65), Expect = 2.5
Identities = 13/28 (46%), Positives = 20/28 (71%)
Frame = +3
Query: 6 WNCATLYSRLVNSSSSSSLHNFTHIRST 33
WN +L + LVNS++ S+ +NFT + ST
Sbjct: 219 WNINSLLTSLVNSATYSAYNNFTVVGST 302
>BE210164 homologue to GP|18086376|gb AT4g23460/F16G20_160 {Arabidopsis
thaliana}, partial (14%)
Length = 427
Score = 29.6 bits (65), Expect = 2.5
Identities = 18/49 (36%), Positives = 28/49 (56%), Gaps = 1/49 (2%)
Frame = -2
Query: 7 NCATLYSRLVNSSSSSSLHNFTHI-RSTFSSSPSVSASKSNDLISLARH 54
N + +YS V+ + SLHN + I + TFS+ S+ A S L ++ RH
Sbjct: 333 NPSQMYSLQVSQDQAGSLHN*SDISKPTFSAPNSLFACNSPHL*TMTRH 187
>TC207234 weakly similar to UP|Q9FH01 (Q9FH01) Similarity to CHP-rich zinc
finger protein, partial (4%)
Length = 449
Score = 29.3 bits (64), Expect = 3.3
Identities = 25/71 (35%), Positives = 33/71 (46%), Gaps = 2/71 (2%)
Frame = -1
Query: 19 SSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLA--RHYGNCYSQLSKARLSLLVVATSG 76
SSSSS + S+ SSSPS S+S + S R +G+C A+ V T
Sbjct: 278 SSSSSSST*SSSSSS*SSSPSSSSSLWSSFFSEEK*RRWGHC------AKTEARTVKTGE 117
Query: 77 TGFVLGSGGAV 87
LG+GG V
Sbjct: 116 EALALGNGGTV 84
>BE658260 similar to GP|9757958|dbj mitochondrial carrier protein-like
{Arabidopsis thaliana}, partial (46%)
Length = 775
Score = 29.3 bits (64), Expect = 3.3
Identities = 27/99 (27%), Positives = 43/99 (43%), Gaps = 4/99 (4%)
Frame = -3
Query: 113 FEIKNDAIMRRTSQRPLPSGRITVPHAVGWASSVGLAGTALLATQTNMLAA----GLAAS 168
FE A++++T Q + + + A+ A S L T L +T ++ G++
Sbjct: 560 FEYLKAAVLQKTKQSYMEPVQSVLCGALAGAISASLT-TPLDVVKTRLMTQVRGEGVSKV 384
Query: 169 NLVLYSFVYTPLKQIHPVNTWVGAVVGAIPPLLGWAAAS 207
V+Y V +KQI WVG G P +L A S
Sbjct: 383 AAVMYDGVSATVKQILKEEGWVGLTRGMGPRVLHSACFS 267
>TC227361 similar to UP|Q9SD67 (Q9SD67) FtsH metalloprotease-like protein,
partial (26%)
Length = 1111
Score = 28.9 bits (63), Expect = 4.3
Identities = 24/87 (27%), Positives = 34/87 (38%), Gaps = 12/87 (13%)
Frame = +1
Query: 203 WAAASGDISLNSLILPAALYFWQIPHFMALAYMCREDYA-------AGGFKMYSLADPSG 255
WA AS DIS ++ H L Y C+ D+ A SL G
Sbjct: 712 WAKASADISQT-----------ELAHPYNLVYSCKVDFGIIFNT*PASYLTQNSLIQSHG 858
Query: 256 RKT-----AMVAMRNSIYLIPLGFLAY 277
RK +V + N IY++ + F+ Y
Sbjct: 859 RKKDKEKYIVVPLTNYIYILVIEFVVY 939
>TC226635 similar to UP|Q84N37 (Q84N37) Potyvirus VPg interacting protein
(Fragment), complete
Length = 2039
Score = 28.9 bits (63), Expect = 4.3
Identities = 27/95 (28%), Positives = 46/95 (48%), Gaps = 4/95 (4%)
Frame = -2
Query: 5 NWNCATLYSRLVNSSSSSSLHNFTHIRSTFSS--SPSVSASKSNDLISLARHYGNCYSQL 62
NWN + + SSSS+ +F S SS S ++SAS + R + N
Sbjct: 1540 NWNISASAFFSLTIFSSSSICSFFFCLSILSSVTSLALSASSRSQASRDKRAFLNILI-F 1364
Query: 63 SKARLSLLVVATSGTGFVL--GSGGAVDLSMLSYT 95
S A +S+ ++A+ T +L S GA++L + ++
Sbjct: 1363 SSATISIFLIASCTTSAILLHASSGAINLPLSPFS 1259
>TC205570 homologue to UP|Q93VJ8 (Q93VJ8) AT5g11200/F2I11_90
(AT5g11170/F2I11_60), partial (98%)
Length = 1745
Score = 28.5 bits (62), Expect = 5.7
Identities = 25/73 (34%), Positives = 41/73 (55%), Gaps = 9/73 (12%)
Frame = -1
Query: 10 TLYSRL---VNSSSSSSLHNFTHIRSTFS-----SSP-SVSASKSNDLISLARHYGNCYS 60
TL++++ + SS+S NF S+F+ +SP SV+ + S+ IS+ G+C +
Sbjct: 1013 TLFTKITT*LKSSASRRSFNFRFFSSSFNLM*CCTSP*SVNLASSST*ISI----GSCIN 846
Query: 61 QLSKARLSLLVVA 73
L R+SLL VA
Sbjct: 845 FLQTGRISLLSVA 807
>TC212170
Length = 793
Score = 28.5 bits (62), Expect = 5.7
Identities = 25/90 (27%), Positives = 35/90 (38%)
Frame = +3
Query: 48 LISLARHYGNCYSQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYTCLGTMMVAASAS 107
L S+ H G+ Y+ LSK SL V + +++ D + L CLG V S
Sbjct: 252 LDSILMHMGSMYATLSKFEKSLDVYERA--VYIMERTYGKDSTFLVTPCLGMAKVIGSIG 425
Query: 108 TLNQVFEIKNDAIMRRTSQRPLPSGRITVP 137
+ E I S R S + VP
Sbjct: 426 KATKAIETYQRVITLLESSRGANSKDLVVP 515
>TC233945 weakly similar to UP|Q8L5R4 (Q8L5R4) Tubulin folding cofactor C,
partial (30%)
Length = 612
Score = 28.5 bits (62), Expect = 5.7
Identities = 16/50 (32%), Positives = 25/50 (50%)
Frame = +1
Query: 14 RLVNSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYGNCYSQLS 63
R N SSS L F+H++++ S + S S S+D +L H +S
Sbjct: 106 RSQNPESSSILSRFSHLKTSIESQLAQSQSISSDPSNLKLHINQISQSIS 255
>BE330049 homologue to GP|2252828|gb| A_IG005I10.5 gene product {Arabidopsis
thaliana}, partial (54%)
Length = 467
Score = 28.1 bits (61), Expect = 7.4
Identities = 23/75 (30%), Positives = 37/75 (48%)
Frame = +1
Query: 7 NCATLYSRLVNSSSSSSLHNFTHIRSTFSSSPSVSASKSNDLISLARHYGNCYSQLSKAR 66
+C++ +S++SSS F S SSSP++SAS+S+ + S S++
Sbjct: 178 SCSSPSKPAASSAASSSSSPFRSSSSPTSSSPNLSASRSS---------SSSPSPASRSA 330
Query: 67 LSLLVVATSGTGFVL 81
S A S GF+L
Sbjct: 331 TSNSSHAPSSLGFML 375
>TC204844 similar to UP|Q9LT85 (Q9LT85) Similarity to receptor protein
kinase, partial (19%)
Length = 1738
Score = 28.1 bits (61), Expect = 7.4
Identities = 17/46 (36%), Positives = 23/46 (49%), Gaps = 1/46 (2%)
Frame = -1
Query: 352 KSPSSVETSEMDDKNGNQKIKGRQGTRAR-PPVAYASIAPFPFLPA 396
K P+ + E+ KN N + R+ R + PP AS APF PA
Sbjct: 574 KPPTKEKLLELTWKNVNSGMLSRRPVRGKLPPFLVASFAPFRSSPA 437
>TC203946 homologue to UP|Q9LN13 (Q9LN13) T6D22.2, partial (47%)
Length = 1858
Score = 28.1 bits (61), Expect = 7.4
Identities = 22/90 (24%), Positives = 37/90 (40%), Gaps = 7/90 (7%)
Frame = +1
Query: 39 SVSASKSNDLISLARHY---GNCYSQLSKARLSLLVVATSGTGFVLGSGGAVDLSMLSYT 95
S+ + N + + H+ G C S L L TGF++ G A+ S+L +
Sbjct: 1417 SIIMTTINGFLFVVLHFFV*GTCLSDFVSGLLRHLATFVPRTGFLIDGGKAISFSLLCFV 1596
Query: 96 ----CLGTMMVAASASTLNQVFEIKNDAIM 121
CL S S + QV+ + +I+
Sbjct: 1597 SITMCLCKNNKLHSTSPMLQVWLLNFKSIL 1686
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.132 0.396
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 18,657,744
Number of Sequences: 63676
Number of extensions: 267011
Number of successful extensions: 1935
Number of sequences better than 10.0: 46
Number of HSP's better than 10.0 without gapping: 1766
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1865
length of query: 402
length of database: 12,639,632
effective HSP length: 99
effective length of query: 303
effective length of database: 6,335,708
effective search space: 1919719524
effective search space used: 1919719524
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.9 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146329.8


