

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
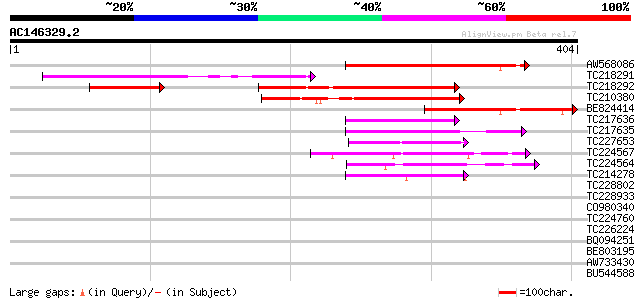
Query= AC146329.2 + phase: 0
(404 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
AW568086 202 3e-52
TC218291 similar to GB|AAM91387.1|22137084|AY133557 At3g13350/MD... 170 1e-42
TC218292 similar to GB|AAM91387.1|22137084|AY133557 At3g13350/MD... 147 1e-35
TC210380 similar to PIR|B96789|B96789 protein T23E18.4 [imported... 135 3e-32
BE824414 117 9e-27
TC217636 homologue to UP|O04235 (O04235) Transcription factor, p... 63 3e-10
TC217635 similar to UP|O04235 (O04235) Transcription factor, par... 61 8e-10
TC227653 similar to UP|O80383 (O80383) 98b, partial (78%) 51 1e-06
TC224567 similar to PIR|T51159|T51159 HMG protein [imported] - A... 47 1e-05
TC224564 similar to PIR|T51159|T51159 HMG protein [imported] - A... 45 6e-05
TC214278 similar to UP|P93704 (P93704) HMG-1, partial (98%) 43 3e-04
TC228802 similar to UP|Q6ZLD8 (Q6ZLD8) Fiber protein-like, parti... 38 0.009
TC228933 similar to UP|Q6NQ79 (Q6NQ79) At3g43240, partial (32%) 36 0.027
CO980340 36 0.027
TC224760 similar to UP|O04692 (O04692) DNA-binding protein, part... 36 0.036
TC226224 35 0.061
BQ094251 similar to GP|14550383|gb Sheath to neuron transformati... 33 0.18
BE803195 similar to GP|8920409|emb| SSRP1 protein {Zea mays}, pa... 32 0.51
AW733430 31 0.88
BU544588 29 4.4
>AW568086
Length = 435
Score = 202 bits (513), Expect = 3e-52
Identities = 104/138 (75%), Positives = 112/138 (80%), Gaps = 7/138 (5%)
Frame = +3
Query: 240 RKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKDREISRTIGELWNKLPESEKA 299
RKKSE+K+RDPAHPKPNRSGYNFFFAEQH RLK LH GKDREISR IGELWNKL ESEK
Sbjct: 3 RKKSEIKRRDPAHPKPNRSGYNFFFAEQHARLKLLHHGKDREISRMIGELWNKLKESEKT 182
Query: 300 VYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLN-------AEAD 352
VYQ+KA+KDKERY EME YREK K +VISDAVPL+QRLPEPD DML+ AE D
Sbjct: 183 VYQEKAMKDKERYRAEMEDYREKQKMGQVISDAVPLQQRLPEPDADMLDVDIKMDEAEGD 362
Query: 353 SLQTPEQSSLDGSDDYED 370
S QTPE+SS G DYED
Sbjct: 363 SPQTPEESS--GGSDYED 410
>TC218291 similar to GB|AAM91387.1|22137084|AY133557 At3g13350/MDC11_14
{Arabidopsis thaliana;} , partial (22%)
Length = 697
Score = 170 bits (430), Expect = 1e-42
Identities = 93/195 (47%), Positives = 115/195 (58%)
Frame = +3
Query: 24 YPPPMATYEEVVDNPKLFILCLEKLHTLMGTKFMIPVIGGRELDLHRLFVEVTSRGGFEK 83
YP P A Y+++V + LF L+ H ++GTK+ + +GG LDLHRLFVEVTSRGG EK
Sbjct: 201 YPAPTARYQDIVRDANLFWGTLQAFHKILGTKYKVATVGGTSLDLHRLFVEVTSRGGIEK 380
Query: 84 IIKDRKWKEVTLVFNFPSTATNASFVLRKYYTSLLYHYEQIYYFKARDWTNTTSDVLQSQ 143
+I DRKWKEV L FNF T T+AS V+RK Y S+LYHYEQ+YYF +
Sbjct: 381 VIVDRKWKEVILTFNFKDTITSASXVVRKSYLSMLYHYEQVYYF--------------GR 518
Query: 144 SSIPAPAPKMQFSHPSPQVQPAVFPMGSSSAGSQVVGVIDGKFDSGYLVTVTIGSEKLKG 203
IP P P VQ + V G ID KFD GY+VTV +GSE+LKG
Sbjct: 519 QGIPPPT-------PGTPVQ---------AHDDMVSGTIDAKFDGGYVVTVILGSEQLKG 650
Query: 204 VLYQAPQNPVLPASH 218
VL+ P N V +SH
Sbjct: 651 VLFHVPDN-VSQSSH 692
>TC218292 similar to GB|AAM91387.1|22137084|AY133557 At3g13350/MDC11_14
{Arabidopsis thaliana;} , partial (37%)
Length = 972
Score = 147 bits (370), Expect = 1e-35
Identities = 74/143 (51%), Positives = 99/143 (68%)
Frame = +2
Query: 178 VVGVIDGKFDSGYLVTVTIGSEKLKGVLYQAPQNPVLPASHHSVPANNNNVTASVGVHRR 237
V G ID KFD GY+VTVT+GSE+LKGVL+ P N V +SH +++ N+ G
Sbjct: 320 VSGTIDAKFDVGYVVTVTLGSEQLKGVLFHVPDN-VSQSSHAEGTSSSQNL--GDGTSNS 490
Query: 238 RRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKDREISRTIGELWNKLPESE 297
+ RK+++ RDP PK NRSGYNFFFAE + RLKP + G++R IS+ IG LWN L E+E
Sbjct: 491 QSRKRAKYAPRDPFRPKSNRSGYNFFFAENYARLKPSYHGQERAISKRIGFLWNNLSEAE 670
Query: 298 KAVYQDKAVKDKERYITEMEYYR 320
+ VYQ+K ++DKERY TE+ Y+
Sbjct: 671 RQVYQEKGMRDKERYRTELMEYK 739
Score = 81.6 bits (200), Expect = 6e-16
Identities = 38/53 (71%), Positives = 42/53 (78%)
Frame = +3
Query: 58 IPVIGGRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTATNASFVL 110
I +GG LDLHRLFVEVTSRGG EK+I DRKWKEV L FNF T TNASF++
Sbjct: 48 ISTVGGTPLDLHRLFVEVTSRGGIEKVIVDRKWKEVILTFNFKDTITNASFMI 206
>TC210380 similar to PIR|B96789|B96789 protein T23E18.4 [imported] -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(32%)
Length = 807
Score = 135 bits (341), Expect = 3e-32
Identities = 75/150 (50%), Positives = 99/150 (66%), Gaps = 5/150 (3%)
Frame = +2
Query: 180 GVIDGKFDSGYLVTVTIGSEKLKGVLYQAPQNPVLPAS---HHS--VPANNNNVTASVGV 234
G I+GKFD GYLV+V +GSE L+GVLY P+ V P S H S VP N
Sbjct: 104 GTIEGKFDCGYLVSVKLGSEVLRGVLYH-PEQLVPPPSIPKHESAIVPINRKP------- 259
Query: 235 HRRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKDREISRTIGELWNKLP 294
HR RRKK++ ++ DP +PKPNRSGYNFFFAE+H LK L+ ++RE ++ IG+ WN L
Sbjct: 260 HRSGRRKKNQ-RRWDPNYPKPNRSGYNFFFAEKHYTLKTLYPNREREFTKMIGQSWNSLS 436
Query: 295 ESEKAVYQDKAVKDKERYITEMEYYREKLK 324
E+ VYQ+ ++DKERY E+ Y+EK+K
Sbjct: 437 PEERMVYQNIGLRDKERYKRELTEYKEKMK 526
>BE824414
Length = 627
Score = 117 bits (293), Expect = 9e-27
Identities = 70/119 (58%), Positives = 83/119 (68%), Gaps = 10/119 (8%)
Frame = -2
Query: 296 SEKAVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLN------- 348
SEK VYQ+KA+ KERY EM REK K +VISDAVPL QRLPEPD DML+
Sbjct: 626 SEKTVYQEKAMXXKERYRXEMXXXREKQKMGQVISDAVPLXQRLPEPDADMLDVDIKMDE 447
Query: 349 AEADSLQTPEQSSLDGSDDYED-DKAKEKDFSVDSLPVIGLGAES--MDSVERSSKLGL 404
AE DS QTPE+SS G DYED +K E+ F +D+LPVIG+GAE+ + S E+SSK GL
Sbjct: 446 AEGDSPQTPEESS--GXSDYEDYNKTAERGFDMDALPVIGMGAETSYLGSEEKSSKGGL 276
>TC217636 homologue to UP|O04235 (O04235) Transcription factor, partial (45%)
Length = 1101
Score = 62.8 bits (151), Expect = 3e-10
Identities = 32/82 (39%), Positives = 47/82 (57%), Gaps = 1/82 (1%)
Frame = +2
Query: 240 RKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEK 298
+KK + KK+DP PK SG+ FF + LK + G ++ R +GE W KL EK
Sbjct: 590 KKKKQKKKKDPNAPKRAMSGFMFFSKLERENLKKTNPGISFTDVGRVLGEKWKKLSAEEK 769
Query: 299 AVYQDKAVKDKERYITEMEYYR 320
Y+ KA +DK+RY+ E+ Y+
Sbjct: 770 EPYEAKAREDKKRYMDEISGYK 835
>TC217635 similar to UP|O04235 (O04235) Transcription factor, partial (22%)
Length = 759
Score = 61.2 bits (147), Expect = 8e-10
Identities = 41/130 (31%), Positives = 59/130 (44%), Gaps = 1/130 (0%)
Frame = +2
Query: 240 RKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKD-REISRTIGELWNKLPESEK 298
+K+ + K++DP PK SG+ FF + LK + G ++SR +GE W KL EK
Sbjct: 140 KKRKQKKRKDPNAPKRAMSGFMFFSKLERENLKKTNPGISFTDVSRVLGEKWKKLSVEEK 319
Query: 299 AVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADSLQTPE 358
Y+ KA +DK+RY E+ Y+ P+P E+DS P
Sbjct: 320 EPYEAKAREDKKRYKDEISGYKN------------------PQPMNIDSGNESDSA*APY 445
Query: 359 QSSLDGSDDY 368
S G D Y
Sbjct: 446 ISYWSGDDIY 475
>TC227653 similar to UP|O80383 (O80383) 98b, partial (78%)
Length = 1654
Score = 50.8 bits (120), Expect = 1e-06
Identities = 28/86 (32%), Positives = 46/86 (52%)
Frame = +1
Query: 242 KSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKDREISRTIGELWNKLPESEKAVY 301
K K++DP PK S Y F ++ L ++ E+ + E W + E +K Y
Sbjct: 814 KKNKKEKDPLKPKHPMSAYFLFTNDRRAALAAENKNF-LEVPKITSEEWKNMTEEQKRPY 990
Query: 302 QDKAVKDKERYITEMEYYREKLKNDE 327
++ A K+KE+Y EME Y++K K++E
Sbjct: 991 EEMAKKNKEQYALEMEAYKQK-KDEE 1065
>TC224567 similar to PIR|T51159|T51159 HMG protein [imported] - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial (71%)
Length = 876
Score = 47.4 bits (111), Expect = 1e-05
Identities = 45/165 (27%), Positives = 70/165 (42%), Gaps = 8/165 (4%)
Frame = +2
Query: 215 PASHHSVPANNNNV---TASVGVHRRRRRKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRL 271
P+ P ++ V AS + R KK + K+DP PK S + F E
Sbjct: 161 PSKESLKPVDDRKVGKRKASGKPEKSRAPKKEKKAKKDPNKPKRPPSAFFVFLEEFRKTF 340
Query: 272 K---PLHRGKDREISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYYREKLKN--D 326
K PL + + + GE W L +EKA Y+ KA K K Y ++ Y +K + D
Sbjct: 341 KAENPLVKAVS-VVGKAGGEKWKSLSSAEKAPYEAKAAKRKAEYEKLIKAYEKKQASSAD 517
Query: 327 EVISDAVPLRQRLPEPDTDMLNAEADSLQTPEQSSLDGSDDYEDD 371
+ SD + + D D + E D + ++ + DD +DD
Sbjct: 518 DDESD----KSKSEVNDEDDASGEEDHQE--DEDDEEEEDDEDDD 634
>TC224564 similar to PIR|T51159|T51159 HMG protein [imported] - Arabidopsis
thaliana {Arabidopsis thaliana;} , partial (82%)
Length = 833
Score = 45.1 bits (105), Expect = 6e-05
Identities = 35/144 (24%), Positives = 61/144 (42%), Gaps = 7/144 (4%)
Frame = +1
Query: 241 KKSEMKKRDPAHPKPNRSGYNFFFAE-------QHPRLKPLHRGKDREISRTIGELWNKL 293
KK + K+DP PK S + F E ++P +K + + + GE W L
Sbjct: 205 KKEKKAKKDPNKPKRPPSAFFVFLEEFRKTFKAENPNVKAVS-----VVGKAGGEKWKSL 369
Query: 294 PESEKAVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDMLNAEADS 353
+EKA Y+ KA K K Y ++ Y +K + + ++D +E +
Sbjct: 370 SSAEKAPYEAKAAKRKAEYEKLIKAYDKKQASS------------ADDEESDKSKSEVN- 510
Query: 354 LQTPEQSSLDGSDDYEDDKAKEKD 377
++ G ++ EDD+ +E D
Sbjct: 511 ----DEDDASGEEEEEDDEEEEDD 570
>TC214278 similar to UP|P93704 (P93704) HMG-1, partial (98%)
Length = 932
Score = 42.7 bits (99), Expect = 3e-04
Identities = 28/92 (30%), Positives = 42/92 (45%), Gaps = 4/92 (4%)
Frame = +1
Query: 240 RKKSEMKKRDPAHPKPNRSGYNFFFAEQHPRLKPLHRGKDRE--ISRTIGELWNKLPESE 297
RK+S +DP PK S + F +E + K H + + GE W L ++E
Sbjct: 358 RKQSRKAAKDPNKPKRPPSAFFVFMSEFREQFKKEHPNNKSVAVVGKAGGEKWKSLSDAE 537
Query: 298 KAVYQDKAVKDKERYITEMEYYREKL--KNDE 327
KA + A K K+ Y + Y ++L KN E
Sbjct: 538 KAPFVATAEKKKQEYEKTISAYNKQLEGKNSE 633
>TC228802 similar to UP|Q6ZLD8 (Q6ZLD8) Fiber protein-like, partial (21%)
Length = 873
Score = 37.7 bits (86), Expect = 0.009
Identities = 23/73 (31%), Positives = 39/73 (52%), Gaps = 4/73 (5%)
Frame = +1
Query: 54 TKFMIPVIGGRELDLHRLFVEVTSRGGFEKIIKDRKWKE--VTLVFNFPST--ATNASFV 109
T+F ++ G+ LDL+ L+ EV +RGGF + WK + + N+ +T T
Sbjct: 16 TEFPDAILNGKRLDLYNLYKEVVTRGGFH-VGNGINWKGQIFSKMRNYTTTNRMTGVGNT 192
Query: 110 LRKYYTSLLYHYE 122
L+++Y + L YE
Sbjct: 193 LKRHYETYLLEYE 231
>TC228933 similar to UP|Q6NQ79 (Q6NQ79) At3g43240, partial (32%)
Length = 1202
Score = 36.2 bits (82), Expect = 0.027
Identities = 37/124 (29%), Positives = 53/124 (41%), Gaps = 9/124 (7%)
Frame = +1
Query: 8 SKSPLPMKETALSHG-EYPPPMATYEEVVDNPKLFILCLEKLHTLMG----TKFMIPVIG 62
S +PLPMK+ HG + P A EE + L L + L+ +F ++
Sbjct: 298 SLNPLPMKK----HGCDRAPIRACSEEEFLRDVMQFLILRGHNRLIPPGGLAEFPDAILN 465
Query: 63 GRELDLHRLFVEVTSRGGFEKIIKDRKWKEVTLVFNFPSTATN----ASFVLRKYYTSLL 118
+ LDL L+ EV SRGGF + WK T TN L+++Y + L
Sbjct: 466 AKRLDLFNLYREVVSRGGFH-VGNGINWKGQVFSKMRNHTMTNRMTGVGNTLKRHYETYL 642
Query: 119 YHYE 122
YE
Sbjct: 643 LEYE 654
>CO980340
Length = 792
Score = 36.2 bits (82), Expect = 0.027
Identities = 28/108 (25%), Positives = 44/108 (39%), Gaps = 14/108 (12%)
Frame = -2
Query: 175 GSQVVGVIDGKFDSGYLVTVTIGSEKLKGVLYQAP-----QNPVLPASHHSVPANNNNVT 229
G++V G +DG FDSG L+T + +GVL+ AP V P S +
Sbjct: 599 GAEVRGKVDGAFDSGLLMTAAVNGRVFRGVLF-APGAGVFSTSVEPKCSVSCSFPSTQPF 423
Query: 230 ASVGVHRRRRRKKSEMKKRDPAH---------PKPNRSGYNFFFAEQH 268
+ H R ++ + + +P H P P G A++H
Sbjct: 422 MNSNHHVRDSQQVANCSRAEPCHGISQPVLAQPTPIMRGTTVSLAKEH 279
>TC224760 similar to UP|O04692 (O04692) DNA-binding protein, partial (73%)
Length = 654
Score = 35.8 bits (81), Expect = 0.036
Identities = 26/92 (28%), Positives = 43/92 (46%), Gaps = 2/92 (2%)
Frame = +3
Query: 282 ISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYYREKLKN--DEVISDAVPLRQRL 339
+ + GE W L +EKA Y+ KA K K Y ++ Y +K + D+ SD + +
Sbjct: 195 VGKAGGEKWKSLSSAEKAPYEAKAAKRKAEYEKLIKAYEKKQASSADDDESD----KSKS 362
Query: 340 PEPDTDMLNAEADSLQTPEQSSLDGSDDYEDD 371
D D + E D + ++ + DD +DD
Sbjct: 363 EVNDEDDASGEEDHQE--DEDDEEEEDDEDDD 452
>TC226224
Length = 1804
Score = 35.0 bits (79), Expect = 0.061
Identities = 22/66 (33%), Positives = 35/66 (52%), Gaps = 9/66 (13%)
Frame = +3
Query: 171 SSSAGSQVVGVIDGKFDSGYLVTVTIGSEK--LKGVLYQ-------APQNPVLPASHHSV 221
++ G V GVI+ FD+GYL++V +G L+G++++ +P N V P SV
Sbjct: 393 NAMVGQLVSGVIEAAFDAGYLLSVRVGDSNTTLRGLIFKPGRYVPISPDNDVAP----SV 560
Query: 222 PANNNN 227
P N
Sbjct: 561 PMIQRN 578
>BQ094251 similar to GP|14550383|gb Sheath to neuron transformation protein 1
isoform b {Caenorhabditis elegans}, partial (47%)
Length = 341
Score = 33.5 bits (75), Expect = 0.18
Identities = 16/52 (30%), Positives = 25/52 (47%)
Frame = +1
Query: 268 HPRLKPLHRGKDREISRTIGELWNKLPESEKAVYQDKAVKDKERYITEMEYY 319
H + P + EI++ E W + + EK + + A KD ERY E+ Y
Sbjct: 1 HKKKYPTENVQVTEITKKCSEKWKTMSDDEKRRFYELAQKDAERYQAEVAAY 156
>BE803195 similar to GP|8920409|emb| SSRP1 protein {Zea mays}, partial (2%)
Length = 253
Score = 32.0 bits (71), Expect = 0.51
Identities = 13/25 (52%), Positives = 18/25 (72%)
Frame = +2
Query: 240 RKKSEMKKRDPAHPKPNRSGYNFFF 264
+KK++ KK+DP PK SG+ FFF
Sbjct: 98 KKKNQKKKKDPIAPKGAMSGFLFFF 172
>AW733430
Length = 184
Score = 31.2 bits (69), Expect = 0.88
Identities = 15/38 (39%), Positives = 24/38 (62%), Gaps = 2/38 (5%)
Frame = +2
Query: 175 GSQVVGVIDGKFDSGYLVTVTIGSEK--LKGVLYQAPQ 210
G V GVIDG F++GYL++V + L+G+++ Q
Sbjct: 14 GKVVTGVIDGTFNAGYLLSVKVADTDAFLRGLVFVEDQ 127
>BU544588
Length = 414
Score = 28.9 bits (63), Expect = 4.4
Identities = 23/89 (25%), Positives = 44/89 (48%), Gaps = 1/89 (1%)
Frame = -2
Query: 288 ELWNKLPESEKAVYQDKAVKDKERYITEMEYYREKLKNDEVISDAVPLRQRLPEPDTDML 347
E+ + E+E+ +DK VK++ +E Y +KN I D L +L + + +
Sbjct: 404 EIERMVREAEEFAEEDKKVKERXXARNSLETYVYNMKNQ--IGDKDKLXXKLXXDEKEKI 231
Query: 348 -NAEADSLQTPEQSSLDGSDDYEDDKAKE 375
NA ++L+ + + ++YE +K KE
Sbjct: 230 ENAVKEALEWLDDNQSVEKEEYE-EKLKE 147
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.314 0.132 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 17,947,614
Number of Sequences: 63676
Number of extensions: 267043
Number of successful extensions: 1622
Number of sequences better than 10.0: 65
Number of HSP's better than 10.0 without gapping: 1558
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1610
length of query: 404
length of database: 12,639,632
effective HSP length: 99
effective length of query: 305
effective length of database: 6,335,708
effective search space: 1932390940
effective search space used: 1932390940
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (22.0 bits)
S2: 60 (27.7 bits)
Medicago: description of AC146329.2


