

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.15 - phase: 0
(204 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
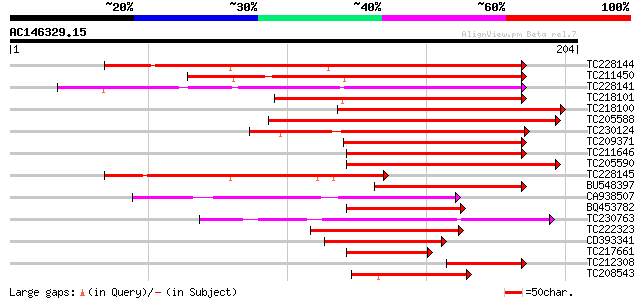
Score E
Sequences producing significant alignments: (bits) Value
TC228144 homologue to UP|Q76DY3 (Q76DY3) AG-motif binding protei... 189 1e-48
TC211450 similar to UP|Q76DY3 (Q76DY3) AG-motif binding protein-... 143 6e-35
TC228141 similar to GB|AAL77730.1|18700240|AY078029 AT3g54810/F2... 124 4e-29
TC218101 similar to GB|AAL77730.1|18700240|AY078029 AT3g54810/F2... 119 7e-28
TC218100 similar to UP|Q9AVU3 (Q9AVU3) GATA-1 zinc finger protei... 118 2e-27
TC205588 similar to UP|Q9FH57 (Q9FH57) GATA-binding transcriptio... 118 2e-27
TC230124 similar to UP|Q76DY1 (Q76DY1) AG-motif binding protein-... 117 5e-27
TC209371 similar to PIR|T52104|T52104 GATA-binding transcription... 114 4e-26
TC211646 homologue to GB|AAP37701.1|30725358|BT008342 At4g32890 ... 114 4e-26
TC205590 similar to UP|Q76DY0 (Q76DY0) AG-motif binding protein-... 107 5e-24
TC228145 similar to UP|Q76DY3 (Q76DY3) AG-motif binding protein-... 101 2e-22
BU548397 97 5e-21
CA938507 homologue to PIR|T52104|T521 GATA-binding transcription... 87 7e-18
BQ453782 80 6e-16
TC230763 67 6e-12
TC222323 similar to UP|Q9FJ10 (Q9FJ10) Gb|AAF63824.1, partial (39%) 66 1e-11
CD393341 59 2e-09
TC217661 similar to UP|Q9FJ10 (Q9FJ10) Gb|AAF63824.1, partial (47%) 54 6e-08
TC212308 similar to UP|Q76DY1 (Q76DY1) AG-motif binding protein-... 46 1e-05
TC208543 similar to GB|BAD02931.1|38603660|AB119061 GATA-type zi... 45 2e-05
>TC228144 homologue to UP|Q76DY3 (Q76DY3) AG-motif binding protein-1, partial
(27%)
Length = 1018
Score = 189 bits (479), Expect = 1e-48
Identities = 101/156 (64%), Positives = 114/156 (72%), Gaps = 4/156 (2%)
Frame = +1
Query: 35 RTRSKRPRLATFSSHHSTMQLISSTSSFVGENMQDSVISNKGAS-TEKFPDSQIAAKKQK 93
R RSKRPR ATF+ + M LIS SSFVGENMQ +VIS+K +S +E F +SQ+ K
Sbjct: 37 RARSKRPRPATFNPRPA-MNLISPASSFVGENMQPNVISSKASSDSENFAESQLVPIFLK 213
Query: 94 LSSGESKKNKKTKAPLLAAL---DHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACG 150
L+SGE KK K K PL A + NA VR+C HCE TKTPQWR GP GPKTLCNACG
Sbjct: 214 LASGEPKKKNKVKVPLPVAPADNNQNASQPVRKCMHCEITKTPQWRAGPMGPKTLCNACG 393
Query: 151 VRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMR 186
VRYKSGRL PEYRPAAS TF P +HSNSHKK+LEMR
Sbjct: 394 VRYKSGRLFPEYRPAASPTFCPSVHSNSHKKVLEMR 501
>TC211450 similar to UP|Q76DY3 (Q76DY3) AG-motif binding protein-1, partial
(23%)
Length = 825
Score = 143 bits (360), Expect = 6e-35
Identities = 75/125 (60%), Positives = 86/125 (68%), Gaps = 3/125 (2%)
Frame = +2
Query: 65 ENMQDSVISNKGAST--EKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAALDHNALG-LV 121
EN Q + + AS+ E F +S I A KQ +SGE KK KK K + + NA +
Sbjct: 8 ENTQHNAANTSKASSDSENFAESVIKAPKQ--ASGEHKKKKKIKVTFPSGQERNAPSQAI 181
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKK 181
R+C HCE TKTPQWR GP GPKTLCNACGVRYKSGRL PEYRPAAS TF +HSNSHKK
Sbjct: 182 RKCLHCEITKTPQWRAGPMGPKTLCNACGVRYKSGRLFPEYRPAASPTFCAAMHSNSHKK 361
Query: 182 ILEMR 186
+LEMR
Sbjct: 362 VLEMR 376
>TC228141 similar to GB|AAL77730.1|18700240|AY078029 AT3g54810/F28P10_210
{Arabidopsis thaliana;} , partial (23%)
Length = 1143
Score = 124 bits (310), Expect = 4e-29
Identities = 72/176 (40%), Positives = 98/176 (54%), Gaps = 7/176 (3%)
Frame = +3
Query: 18 VLPKSNSSPTCEKTT-------VRRTRSKRPRLATFSSHHSTMQLISSTSSFVGENMQDS 70
V S+SSP+ E + +R R+KR RL+ S +S +++S + + Q
Sbjct: 441 VFESSSSSPSVENSNFELPVIPTKRPRTKRRRLSNISLLYSIPFILTSPAF---QKFQRM 611
Query: 71 VISNKGASTEKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAALDHNALGLVRQCTHCEAT 130
S T+ P ++ K +K + K + + ++ R+C HCE T
Sbjct: 612 DFSKSDIQTQ--PSGELLCKFKKKQRKKDIPLPTNKIEMKRSSSQESVA-PRKCLHCEVT 782
Query: 131 KTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMR 186
KTPQWR GP GPKTLCNACGVRY+SGRL EYRPA+S TF LHSNSHKK+LE+R
Sbjct: 783 KTPQWREGPMGPKTLCNACGVRYRSGRLFAEYRPASSPTFVASLHSNSHKKVLEIR 950
>TC218101 similar to GB|AAL77730.1|18700240|AY078029 AT3g54810/F28P10_210
{Arabidopsis thaliana;} , partial (23%)
Length = 638
Score = 119 bits (299), Expect = 7e-28
Identities = 58/96 (60%), Positives = 67/96 (69%), Gaps = 5/96 (5%)
Frame = +1
Query: 96 SGESKKNKKTKAPLLAALDHNAL-----GLVRQCTHCEATKTPQWRTGPEGPKTLCNACG 150
S + KK +K LL+ ++ G R+C HCE TKTPQWR GP GPKTLCNACG
Sbjct: 118 SNKVKKQRKKDLSLLSDVEMTRSSSPESGPPRKCMHCEVTKTPQWREGPMGPKTLCNACG 297
Query: 151 VRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMR 186
VRY+SGRL PEYRPAAS TF LHSN HKK++EMR
Sbjct: 298 VRYRSGRLFPEYRPAASPTFVASLHSNCHKKVVEMR 405
>TC218100 similar to UP|Q9AVU3 (Q9AVU3) GATA-1 zinc finger protein, partial
(27%)
Length = 1284
Score = 118 bits (296), Expect = 2e-27
Identities = 54/82 (65%), Positives = 61/82 (73%)
Frame = +2
Query: 119 GLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNS 178
G R+C HCE TKTPQWR GP GPKTLCNACGVRY+SGRL PEYRPAAS TF LHSN
Sbjct: 884 GSPRKCMHCEVTKTPQWREGPVGPKTLCNACGVRYRSGRLFPEYRPAASPTFVASLHSNC 1063
Query: 179 HKKILEMRVMRRKDNKNSGILA 200
HKK++EMR ++ +LA
Sbjct: 1064HKKVVEMRSRAIQEPVRGSMLA 1129
>TC205588 similar to UP|Q9FH57 (Q9FH57) GATA-binding transcription
factor-like protein (At5g66320), partial (25%)
Length = 1084
Score = 118 bits (295), Expect = 2e-27
Identities = 56/105 (53%), Positives = 70/105 (66%)
Frame = +3
Query: 94 LSSGESKKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRY 153
LS S KK K A + + + R+C+HC+ KTPQWRTGP G KTLCNACGVRY
Sbjct: 474 LSLPSSPPTKKQKKRAEAQVQPVGVQIQRRCSHCQVQKTPQWRTGPLGAKTLCNACGVRY 653
Query: 154 KSGRLCPEYRPAASSTFSPDLHSNSHKKILEMRVMRRKDNKNSGI 198
KSGRL EYRPA S TF D+HSNSH+K+LE+R + ++G+
Sbjct: 654 KSGRLFSEYRPACSPTFCSDIHSNSHRKVLEIRKRKEVAEPDTGL 788
>TC230124 similar to UP|Q76DY1 (Q76DY1) AG-motif binding protein-3, partial
(26%)
Length = 894
Score = 117 bits (292), Expect = 5e-27
Identities = 59/107 (55%), Positives = 69/107 (64%), Gaps = 6/107 (5%)
Frame = +1
Query: 87 IAAKKQKLSS------GESKKNKKTKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPE 140
+ KK+KL G+ N T + H + R+CTHC A +TPQWR GP
Sbjct: 76 VVVKKEKLGDYYYDDEGDEVNNNNTNNNNNDNVQHP---IPRRCTHCLAQRTPQWRAGPL 246
Query: 141 GPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMRV 187
GPKTLCNACGVRYKSGRL PEYRPA S TF LHSNSHKK++EMR+
Sbjct: 247 GPKTLCNACGVRYKSGRLLPEYRPAKSPTFVSYLHSNSHKKVMEMRM 387
>TC209371 similar to PIR|T52104|T52104 GATA-binding transcription factor
homolog 2 [imported] - Arabidopsis thaliana {Arabidopsis
thaliana;} , partial (40%)
Length = 857
Score = 114 bits (284), Expect = 4e-26
Identities = 49/66 (74%), Positives = 56/66 (84%)
Frame = +2
Query: 121 VRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHK 180
VR+C+HC KTPQWRTGP GPKTLCNACGVR+KSGRL PEYRPAAS TF HSNSH+
Sbjct: 404 VRRCSHCATDKTPQWRTGPLGPKTLCNACGVRFKSGRLVPEYRPAASPTFVMTQHSNSHR 583
Query: 181 KILEMR 186
K++E+R
Sbjct: 584 KVMELR 601
>TC211646 homologue to GB|AAP37701.1|30725358|BT008342 At4g32890 {Arabidopsis
thaliana;} , partial (26%)
Length = 778
Score = 114 bits (284), Expect = 4e-26
Identities = 50/65 (76%), Positives = 54/65 (82%)
Frame = +2
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKK 181
R+C HC KTPQWRTGP GPKTLCNACGVRYKSGRL PEYRPAAS TF HSNSH+K
Sbjct: 251 RKCLHCATDKTPQWRTGPMGPKTLCNACGVRYKSGRLVPEYRPAASPTFVLTKHSNSHRK 430
Query: 182 ILEMR 186
+LE+R
Sbjct: 431 VLELR 445
>TC205590 similar to UP|Q76DY0 (Q76DY0) AG-motif binding protein-4, partial
(25%)
Length = 1249
Score = 107 bits (266), Expect = 5e-24
Identities = 47/77 (61%), Positives = 57/77 (73%)
Frame = +3
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKK 181
R+C+HC KTPQWRTGP G KT NACGVRYKSGRL EYRPA S TF D+HSNSH+K
Sbjct: 762 RRCSHCHVQKTPQWRTGPLGAKTXXNACGVRYKSGRLFSEYRPACSPTFCSDIHSNSHRK 941
Query: 182 ILEMRVMRRKDNKNSGI 198
+LE+R + ++G+
Sbjct: 942 VLEIRKRKEVAQPDTGL 992
>TC228145 similar to UP|Q76DY3 (Q76DY3) AG-motif binding protein-1, partial
(17%)
Length = 446
Score = 101 bits (252), Expect = 2e-22
Identities = 61/106 (57%), Positives = 71/106 (66%), Gaps = 4/106 (3%)
Frame = +2
Query: 35 RTRSKRPRLATFSSHHSTMQLISSTSSFVGENMQDSVISNKGAS-TEKFPDSQIAAKKQK 93
RTRSKRPR ATF+ M LIS SSFVGENMQ +VIS+K +S +E F +SQ+ K K
Sbjct: 131 RTRSKRPRPATFNP-RPAMNLISPASSFVGENMQPNVISSKSSSDSENFAESQLVPKMPK 307
Query: 94 LSSGESKKNKKTKAPL-LAALDH--NALGLVRQCTHCEATKTPQWR 136
+S E KK KK K PL L D+ NA VR+C HCE TKTPQWR
Sbjct: 308 QASEEPKKKKKVKLPLPLVPADNNQNASQPVRKCMHCEITKTPQWR 445
>BU548397
Length = 605
Score = 97.1 bits (240), Expect = 5e-21
Identities = 43/55 (78%), Positives = 46/55 (83%)
Frame = -2
Query: 132 TPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMR 186
TPQWRTGP GPKT CNACGVRYKSGRL PEYRPAAS TF HSNSH+K+ E+R
Sbjct: 604 TPQWRTGPMGPKTXCNACGVRYKSGRLVPEYRPAASPTFVLTKHSNSHRKVXELR 440
>CA938507 homologue to PIR|T52104|T521 GATA-binding transcription factor
homolog 2 [imported] - Arabidopsis thaliana, partial
(23%)
Length = 449
Score = 86.7 bits (213), Expect = 7e-18
Identities = 46/118 (38%), Positives = 65/118 (54%)
Frame = +1
Query: 45 TFSSHHSTMQLISSTSSFVGENMQDSVISNKGASTEKFPDSQIAAKKQKLSSGESKKNKK 104
+F S +T + S ++SF+ N N ++P S +++ S+ K +
Sbjct: 133 SFHSFPATASIGSHSTSFLSNN-------NNRNDNNEYPKSSLSSNIPCSSAVAGKSRAR 291
Query: 105 TKAPLLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEY 162
+ + G VR+C+HC + KTPQWR GP GPKTLCNACGVR+KSGRL PEY
Sbjct: 292 REGSVTGD------GGVRRCSHCASEKTPQWRAGPLGPKTLCNACGVRFKSGRLVPEY 447
>BQ453782
Length = 424
Score = 80.1 bits (196), Expect = 6e-16
Identities = 31/43 (72%), Positives = 38/43 (88%)
Frame = -1
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRP 164
R+C+HC+A KTPQWRTGP G TLCNACG+R+K+G+L PEYRP
Sbjct: 406 RRCSHCDAIKTPQWRTGPFGRNTLCNACGIRFKAGKLYPEYRP 278
>TC230763
Length = 1058
Score = 67.0 bits (162), Expect = 6e-12
Identities = 40/128 (31%), Positives = 65/128 (50%)
Frame = +1
Query: 69 DSVISNKGASTEKFPDSQIAAKKQKLSSGESKKNKKTKAPLLAALDHNALGLVRQCTHCE 128
D+ ++ +T KF D +KQ+LSS N + +++ VR C+ C
Sbjct: 118 DTYTNSDNNTTHKFDD-----QKQQLSSPLGTDNSSSNN-----YSNHSNNTVRVCSDCH 267
Query: 129 ATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAASSTFSPDLHSNSHKKILEMRVM 188
TKTP WR+GP GPK+LCNACG+R + R AAS++ + + + K + +
Sbjct: 268 TTKTPLWRSGPRGPKSLCNACGIRQRKARRAMA-AAAASASGNGTVIVEAKKSVKGRNKL 444
Query: 189 RRKDNKNS 196
++K K +
Sbjct: 445 QKKKEKKT 468
>TC222323 similar to UP|Q9FJ10 (Q9FJ10) Gb|AAF63824.1, partial (39%)
Length = 368
Score = 66.2 bits (160), Expect = 1e-11
Identities = 28/55 (50%), Positives = 34/55 (60%)
Frame = +2
Query: 109 LLAALDHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGRLCPEYR 163
+L +D N + C C+ TKTP WR GP GPKTLCNACG+RY+ R C R
Sbjct: 143 ILEMMDLNVNEKKKCCADCKTTKTPLWRGGPAGPKTLCNACGIRYRKRRACSRKR 307
>CD393341
Length = 612
Score = 58.5 bits (140), Expect = 2e-09
Identities = 22/44 (50%), Positives = 30/44 (68%)
Frame = -3
Query: 114 DHNALGLVRQCTHCEATKTPQWRTGPEGPKTLCNACGVRYKSGR 157
++N R C+ C + TP WRTGP+GPK+LCNACG+R + R
Sbjct: 574 NNNGSSTPRVCSDCNTSTTPLWRTGPKGPKSLCNACGIRQRKAR 443
>TC217661 similar to UP|Q9FJ10 (Q9FJ10) Gb|AAF63824.1, partial (47%)
Length = 765
Score = 53.5 bits (127), Expect = 6e-08
Identities = 20/31 (64%), Positives = 23/31 (73%)
Frame = +2
Query: 122 RQCTHCEATKTPQWRTGPEGPKTLCNACGVR 152
+ C C TKTP WR GP GPK+LCNACG+R
Sbjct: 284 KTCADCGTTKTPLWRGGPAGPKSLCNACGIR 376
>TC212308 similar to UP|Q76DY1 (Q76DY1) AG-motif binding protein-3, partial
(15%)
Length = 542
Score = 45.8 bits (107), Expect = 1e-05
Identities = 20/29 (68%), Positives = 23/29 (78%)
Frame = +3
Query: 158 LCPEYRPAASSTFSPDLHSNSHKKILEMR 186
L PEYRPAAS TF HSNSH+K+LE+R
Sbjct: 3 LVPEYRPAASPTFMSTKHSNSHRKVLELR 89
>TC208543 similar to GB|BAD02931.1|38603660|AB119061 GATA-type zinc finger
protein {Arabidopsis thaliana;} , partial (50%)
Length = 929
Score = 45.1 bits (105), Expect = 2e-05
Identities = 20/45 (44%), Positives = 28/45 (61%), Gaps = 2/45 (4%)
Frame = +2
Query: 124 CTHCEATK--TPQWRTGPEGPKTLCNACGVRYKSGRLCPEYRPAA 166
C HC ++ TP R GPEGP+TLCNACG+ + + + + AA
Sbjct: 440 CRHCGISEKSTPMMRRGPEGPRTLCNACGLMWANKGILRDLSRAA 574
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.311 0.124 0.353
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 9,100,426
Number of Sequences: 63676
Number of extensions: 123621
Number of successful extensions: 702
Number of sequences better than 10.0: 67
Number of HSP's better than 10.0 without gapping: 695
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 697
length of query: 204
length of database: 12,639,632
effective HSP length: 93
effective length of query: 111
effective length of database: 6,717,764
effective search space: 745671804
effective search space used: 745671804
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.2 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.9 bits)
S2: 56 (26.2 bits)
Medicago: description of AC146329.15


