

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.11 + phase: 0
(327 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
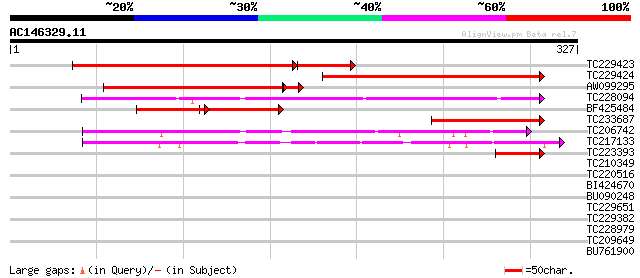
Score E
Sequences producing significant alignments: (bits) Value
TC229423 similar to UP|Q94JV5 (Q94JV5) AT4g08790/T32A17_100, par... 225 6e-73
TC229424 similar to UP|Q94JV5 (Q94JV5) AT4g08790/T32A17_100, par... 240 7e-64
AW099295 similar to GP|13926307|gb AT4g08790/T32A17_100 {Arabido... 174 5e-46
TC228094 similar to UP|Q8RUF8 (Q8RUF8) AT5g12040/F14F18_210, par... 146 1e-35
BF425484 similar to GP|13926307|gb| AT4g08790/T32A17_100 {Arabid... 79 1e-32
TC233687 similar to UP|Q94JV5 (Q94JV5) AT4g08790/T32A17_100, par... 120 6e-28
TC206742 similar to UP|Q9XGI9 (Q9XGI9) Beta-alanine synthase, co... 87 1e-17
TC217133 homologue to UP|Q42966 (Q42966) Nitrilase , partial (96%) 70 1e-12
TC223393 similar to UP|Q94JV5 (Q94JV5) AT4g08790/T32A17_100, par... 54 1e-07
TC210349 33 0.13
TC220516 homologue to UP|Q8RUF8 (Q8RUF8) AT5g12040/F14F18_210, p... 32 0.30
BI424670 homologue to SP|P51567|AFC2 Protein kinase AFC2 (EC 2.7... 31 0.67
BU090248 similar to PIR|T03739|T03 nitrilase (EC 3.5.5.1) 4B - c... 30 1.5
TC229651 28 5.6
TC229382 28 5.6
TC228979 weakly similar to UP|O23389 (O23389) Cytochrome P450 li... 27 9.6
TC209649 weakly similar to UP|Q94A64 (Q94A64) AT5g03970/F8F6_180... 27 9.6
BU761900 weakly similar to GP|15912221|gb| At1g70370/F17O7_9 {Ar... 27 9.6
>TC229423 similar to UP|Q94JV5 (Q94JV5) AT4g08790/T32A17_100, partial (48%)
Length = 566
Score = 225 bits (573), Expect(2) = 6e-73
Identities = 108/130 (83%), Positives = 121/130 (93%)
Frame = +1
Query: 37 STMATNSVRVAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSV 96
+TMA+NSVRVAAAQMTSI+DLA+N +TCSRL+KEAASAGAKLLCFPEAFS+VG KDGDSV
Sbjct: 76 ATMASNSVRVAAAQMTSISDLAANLATCSRLIKEAASAGAKLLCFPEAFSYVGTKDGDSV 255
Query: 97 SIAQPLDGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKI 156
+A+PLDGPIM YCSLARESSIWLSLGGFQEKGSDP+ L NTHV+VDDTGKI ++Y KI
Sbjct: 256 RVAEPLDGPIMSHYCSLARESSIWLSLGGFQEKGSDPQRLSNTHVIVDDTGKIISSYSKI 435
Query: 157 HLFDVDVPGG 166
HLFDVDVPGG
Sbjct: 436 HLFDVDVPGG 465
Score = 67.4 bits (163), Expect(2) = 6e-73
Identities = 31/33 (93%), Positives = 33/33 (99%)
Frame = +2
Query: 167 RVYKESNFTESGKDIVAVDSPIGRLGLSVCYDL 199
RVYKES+FTESGKDIVAVDSP+GRLGLSVCYDL
Sbjct: 467 RVYKESSFTESGKDIVAVDSPVGRLGLSVCYDL 565
>TC229424 similar to UP|Q94JV5 (Q94JV5) AT4g08790/T32A17_100, partial (44%)
Length = 454
Score = 240 bits (612), Expect = 7e-64
Identities = 118/128 (92%), Positives = 122/128 (95%)
Frame = +3
Query: 181 IVAVDSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARA 240
IVAVDSPIGRLGLSVCYDLRFPE+YQLLRFQH AQ+LLVPAAFT VTGEAHWEILLRAR
Sbjct: 3 IVAVDSPIGRLGLSVCYDLRFPEMYQLLRFQHEAQVLLVPAAFTTVTGEAHWEILLRARV 182
Query: 241 IENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVGRLPDRLSTGIVVADIDLSLVDSV 300
IE QCYVIAAAQAG HNDKRESYG+TLIIDPWGT+VGRLPDRLSTGIVVADIDLS VDSV
Sbjct: 183 IETQCYVIAAAQAGKHNDKRESYGNTLIIDPWGTIVGRLPDRLSTGIVVADIDLSFVDSV 362
Query: 301 REKMPIAK 308
REKMPIAK
Sbjct: 363 REKMPIAK 386
>AW099295 similar to GP|13926307|gb AT4g08790/T32A17_100 {Arabidopsis
thaliana}, partial (32%)
Length = 410
Score = 174 bits (441), Expect(2) = 5e-46
Identities = 82/106 (77%), Positives = 95/106 (89%)
Frame = +3
Query: 55 TDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSIAQPLDGPIMDQYCSLA 114
++LA+N +TCS L+KEAASAGAKLLCFPEAFS+VG DGDSV +A+PLDGPIM+ YCSLA
Sbjct: 45 SNLAANMATCSHLLKEAASAGAKLLCFPEAFSYVGTMDGDSVLVAEPLDGPIMNHYCSLA 224
Query: 115 RESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHLFD 160
RES IWLSLGGFQE GSDP+HL NTHV+VDDTGKI ++YRKIHL D
Sbjct: 225 RESDIWLSLGGFQEIGSDPQHLCNTHVIVDDTGKIISSYRKIHLQD 362
Score = 28.1 bits (61), Expect(2) = 5e-46
Identities = 11/11 (100%), Positives = 11/11 (100%)
Frame = +2
Query: 159 FDVDVPGGRVY 169
FDVDVPGGRVY
Sbjct: 377 FDVDVPGGRVY 409
>TC228094 similar to UP|Q8RUF8 (Q8RUF8) AT5g12040/F14F18_210, partial (82%)
Length = 1538
Score = 146 bits (369), Expect = 1e-35
Identities = 86/270 (31%), Positives = 141/270 (51%), Gaps = 3/270 (1%)
Frame = +1
Query: 42 NSVRVAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFSFVGAKDGDSVSIAQP 101
++ ++ Q++ D SN + +++AAS GA+L+ PE ++ + D V A+
Sbjct: 262 SNFKIGLCQLSVSPDKDSNIAHARTAIQDAASKGAQLVLLPEIWNSPYSNDSFPV-YAED 438
Query: 102 LDG---PIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHL 158
+D P L+R I + G E+ L+NT V G + +RKIHL
Sbjct: 439 IDAGASPSTAMLSELSRLLKITIVGGSIPERSGGL--LYNTCCVFGTDGNLLAKHRKIHL 612
Query: 159 FDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILL 218
FD+D+PG + ES +G+ VD+ +GR+G+ +CYD+RFPEL ++ GA +L
Sbjct: 613 FDIDIPGKITFIESKTLTAGETPTIVDTEVGRIGIGICYDIRFPEL-AMIYAARGAHLLC 789
Query: 219 VPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVGR 278
P AF TG HWE+L RARA +NQ YV + A ++G + ++ P+G V+
Sbjct: 790 YPGAFNMTTGPLHWELLQRARATDNQLYVATCSPARDTGSGYVAWGHSTLVGPFGEVLAT 969
Query: 279 LPDRLSTGIVVADIDLSLVDSVREKMPIAK 308
I++A+ID S+++ R +P+ K
Sbjct: 970 TEH--EEAIIIAEIDYSILEQRRTNLPVTK 1053
>BF425484 similar to GP|13926307|gb| AT4g08790/T32A17_100 {Arabidopsis
thaliana}, partial (17%)
Length = 415
Score = 79.3 bits (194), Expect(2) = 1e-32
Identities = 38/49 (77%), Positives = 42/49 (85%)
Frame = +3
Query: 110 YCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHL 158
YC RESSIWLSLGGFQEKGSDP+ L NTHV+VDDTGKI ++Y KIHL
Sbjct: 216 YC---RESSIWLSLGGFQEKGSDPQRLSNTHVIVDDTGKIISSYSKIHL 353
Score = 78.6 bits (192), Expect(2) = 1e-32
Identities = 35/42 (83%), Positives = 38/42 (90%)
Frame = +1
Query: 74 AGAKLLCFPEAFSFVGAKDGDSVSIAQPLDGPIMDQYCSLAR 115
AGAKLLCFPEAFS+VG KDGDSV +A+PLDGPIM YCSLAR
Sbjct: 1 AGAKLLCFPEAFSYVGTKDGDSVRVAEPLDGPIMSHYCSLAR 126
>TC233687 similar to UP|Q94JV5 (Q94JV5) AT4g08790/T32A17_100, partial (22%)
Length = 507
Score = 120 bits (302), Expect = 6e-28
Identities = 59/65 (90%), Positives = 60/65 (91%)
Frame = +2
Query: 244 QCYVIAAAQAGTHNDKRESYGDTLIIDPWGTVVGRLPDRLSTGIVVADIDLSLVDSVREK 303
QCYVIAAAQAG HNDKRES G LIIDPWGT+VGRLPDRLSTGIVVADIDLS VDSVREK
Sbjct: 5 QCYVIAAAQAGKHNDKRESXGXXLIIDPWGTIVGRLPDRLSTGIVVADIDLSFVDSVREK 184
Query: 304 MPIAK 308
MPIAK
Sbjct: 185 MPIAK 199
>TC206742 similar to UP|Q9XGI9 (Q9XGI9) Beta-alanine synthase, complete
Length = 1245
Score = 87.0 bits (214), Expect = 1e-17
Identities = 73/277 (26%), Positives = 130/277 (46%), Gaps = 18/277 (6%)
Frame = +2
Query: 43 SVRVAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPEAFS---FVGAKDGDSVSIA 99
+V V+A Q D+++N +T RLV+ A GA ++ E F F A+ + + A
Sbjct: 95 TVVVSALQFACTDDVSTNVATAERLVRAAHKQGANIILIQELFEGYYFCQAQREEFIQRA 274
Query: 100 QP-LDGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDTGKIQTTYRKIHL 158
+P D P + + LA+E + + + F+E + +N+ ++D G YRK H
Sbjct: 275 KPHKDHPTILRMQKLAKELGVVIPVSFFEEANNAH---YNSTAIIDADGTDLGIYRKSH- 442
Query: 159 FDVDVPGGRVYKESNFTESGKDIVAV-DSPIGRLGLSVCYDLRFPELYQLLRFQHGAQIL 217
+P G Y+E + G V + ++G+++C+D FPE + + Q GA+IL
Sbjct: 443 ----IPDGPGYEEKFYFNPGDTGFKVFQTKFAKIGVAICWDQWFPEAARAMVLQ-GAEIL 607
Query: 218 LVPAAF------TKVTGEAHWEILLRARAIENQCYVIAAAQAG-----THNDKRE--SYG 264
P A + HW+ +++ A N ++A+ + G T + K E YG
Sbjct: 608 FYPTAIGSEPHDGSIDSRDHWKRVMQGHAGANLVPLVASNRIGKEIIETEHGKSEITFYG 787
Query: 265 DTLIIDPWGTVVGRLPDRLSTGIVVADIDLSLVDSVR 301
++ I P G +V + D +++A DL + S+R
Sbjct: 788 NSFIAGPTGEIVS-IADDKEEAVLIAQFDLDNIKSMR 895
>TC217133 homologue to UP|Q42966 (Q42966) Nitrilase , partial (96%)
Length = 1337
Score = 70.5 bits (171), Expect = 1e-12
Identities = 79/322 (24%), Positives = 134/322 (41%), Gaps = 44/322 (13%)
Frame = +2
Query: 43 SVRVAAAQMTSIT-DLASNFSTCSRLVKEAASAGAKLLCFPEAF-------SFVGAKDGD 94
+VR Q ++I D + RL+ EA S G++L+ FPEAF S G G+
Sbjct: 128 TVRATVVQASTIFYDTPATLDKAERLLAEATSYGSQLVVFPEAFVGGYPRGSAFGLSIGN 307
Query: 95 SV-----------SIAQPLDGPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVV 143
S A + GP +D+ ++A + + L +G + G L+ T +
Sbjct: 308 RTVKGREEFRKYHSAAIDVPGPEVDRLAAMAGKYKVHLVMGVIERDGYT---LYCTVLFF 478
Query: 144 DDTGKIQTTYRKIHLFDVDVPGGRVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRFPE 203
D G +RKI +P F + G I ++P+G++G ++C++ R P
Sbjct: 479 DSQGHYLGKHRKI------MPTALERVIWGFGD-GSTIPVFETPVGKIGAAICWENRMP- 634
Query: 204 LYQLLRFQHGAQILLVPAAFTKVTGEAHWEILLRARAIENQCYVIAAAQ----------- 252
L + + G +I P A ++ W+ + A+E C+V++A Q
Sbjct: 635 LLRTAMYAKGVEIYCAPTADSRDV----WQASMTHIALEGGCFVLSANQFCRRKDYPPPP 802
Query: 253 ----AGTHNDKRES----YGDTLIIDPWGTVVGRLPDRLSTGIVVADIDLSLVDSVREKM 304
AGT D G ++II P G V+ P+ ++ AD+DL + +
Sbjct: 803 EYVFAGTEEDLTPDSVVCAGGSVIISPSGAVLAG-PNYDGEALISADLDLGEIARAKFDF 979
Query: 305 PIA------KVIVCNVYIHPTN 320
+ +V+ V HPTN
Sbjct: 980 DVVGHYSRPEVLSLIVKDHPTN 1045
>TC223393 similar to UP|Q94JV5 (Q94JV5) AT4g08790/T32A17_100, partial (10%)
Length = 711
Score = 53.9 bits (128), Expect = 1e-07
Identities = 27/28 (96%), Positives = 27/28 (96%)
Frame = +2
Query: 281 DRLSTGIVVADIDLSLVDSVREKMPIAK 308
DRLSTGIVVADIDLS VDSVREKMPIAK
Sbjct: 500 DRLSTGIVVADIDLSFVDSVREKMPIAK 583
>TC210349
Length = 605
Score = 33.5 bits (75), Expect = 0.13
Identities = 27/87 (31%), Positives = 43/87 (49%), Gaps = 2/87 (2%)
Frame = -2
Query: 42 NSVRVAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLC--FPEAFSFVGAKDGDSVSIA 99
+S+ + + M S +D S ST SR ++S + + FP++F F A D + +
Sbjct: 424 SSISSSVS*MDSKSDSKSRSSTSSRGRMRSSSMSSSSVSS*FPDSFRFRIASMIDGSTFS 245
Query: 100 QPLDGPIMDQYCSLARESSIWLSLGGF 126
PL + + SL SSIW + GGF
Sbjct: 244 SPLK*VSICAFSSLLFTSSIWGTGGGF 164
>TC220516 homologue to UP|Q8RUF8 (Q8RUF8) AT5g12040/F14F18_210, partial (7%)
Length = 585
Score = 32.3 bits (72), Expect = 0.30
Identities = 14/27 (51%), Positives = 16/27 (58%)
Frame = +2
Query: 213 GAQILLVPAAFTKVTGEAHWEILLRAR 239
GA +L P AF TG WE+L RAR
Sbjct: 83 GAHLLCYPGAFNMTTGSLPWELLQRAR 163
>BI424670 homologue to SP|P51567|AFC2 Protein kinase AFC2 (EC 2.7.1.-).
[Mouse-ear cress] {Arabidopsis thaliana}, partial (33%)
Length = 462
Score = 31.2 bits (69), Expect = 0.67
Identities = 16/50 (32%), Positives = 25/50 (50%), Gaps = 1/50 (2%)
Frame = -3
Query: 172 SNFTESGKDIVAVDSPIGRLGLSVCYDLRFPELYQLLRFQHGA-QILLVP 220
S F + D++ + P+ L ++ + FP+L Q L F HG I L P
Sbjct: 160 SKFLKHNTDMITIVKPVPYLHTTIATFIVFPKLLQYLNFYHGCLSIFLYP 11
>BU090248 similar to PIR|T03739|T03 nitrilase (EC 3.5.5.1) 4B - common
tobacco, partial (29%)
Length = 415
Score = 30.0 bits (66), Expect = 1.5
Identities = 11/42 (26%), Positives = 26/42 (61%)
Frame = +3
Query: 178 GKDIVAVDSPIGRLGLSVCYDLRFPELYQLLRFQHGAQILLV 219
G I ++P+G++G ++C++ R P L + + G +I+++
Sbjct: 282 GSTIPVFETPVGKIGAAICWENRMP-LLRTAMYAXGCEIIVL 404
>TC229651
Length = 1257
Score = 28.1 bits (61), Expect = 5.6
Identities = 12/16 (75%), Positives = 13/16 (81%)
Frame = +3
Query: 108 DQYCSLARESSIWLSL 123
+QYC L R SSIWLSL
Sbjct: 1041 NQYCCLIRWSSIWLSL 1088
>TC229382
Length = 1290
Score = 28.1 bits (61), Expect = 5.6
Identities = 20/53 (37%), Positives = 26/53 (48%), Gaps = 6/53 (11%)
Frame = -3
Query: 156 IHLFD-----VDVPGG-RVYKESNFTESGKDIVAVDSPIGRLGLSVCYDLRFP 202
+HLF +D GG R+ + S D ++ DS LGLS C D RFP
Sbjct: 931 LHLFKAFSLIIDKGGGVRLMPVPPWNFSSSDKISPDSLTVTLGLSECIDPRFP 773
>TC228979 weakly similar to UP|O23389 (O23389) Cytochrome P450 like
protein, partial (26%)
Length = 1207
Score = 27.3 bits (59), Expect = 9.6
Identities = 17/46 (36%), Positives = 22/46 (46%)
Frame = +2
Query: 36 ESTMATNSVRVAAAQMTSITDLASNFSTCSRLVKEAASAGAKLLCF 81
E T TN+V A TS + + +LVKE+ AK LCF
Sbjct: 35 EFTKFTNNVTCRTAMSTSCAEKCEDAERIRKLVKESFELAAK-LCF 169
>TC209649 weakly similar to UP|Q94A64 (Q94A64) AT5g03970/F8F6_180, partial
(8%)
Length = 941
Score = 27.3 bits (59), Expect = 9.6
Identities = 13/43 (30%), Positives = 22/43 (50%)
Frame = +1
Query: 104 GPIMDQYCSLARESSIWLSLGGFQEKGSDPRHLFNTHVVVDDT 146
G + +CS+AR + IW G+ + +P HLF ++ T
Sbjct: 25 GSHLGGFCSMAR-AIIWAWRYGYLRRNQEPTHLFTATALI*ST 150
>BU761900 weakly similar to GP|15912221|gb| At1g70370/F17O7_9 {Arabidopsis
thaliana}, partial (14%)
Length = 445
Score = 27.3 bits (59), Expect = 9.6
Identities = 16/36 (44%), Positives = 22/36 (60%)
Frame = +3
Query: 48 AAQMTSITDLASNFSTCSRLVKEAASAGAKLLCFPE 83
AA+ + LAS + +RL + A+A KLLCFPE
Sbjct: 339 AAEAAAFVKLASGNALSTRLPEFCAAA--KLLCFPE 440
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.320 0.135 0.401
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,225,116
Number of Sequences: 63676
Number of extensions: 175410
Number of successful extensions: 715
Number of sequences better than 10.0: 36
Number of HSP's better than 10.0 without gapping: 713
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 713
length of query: 327
length of database: 12,639,632
effective HSP length: 98
effective length of query: 229
effective length of database: 6,399,384
effective search space: 1465458936
effective search space used: 1465458936
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 59 (27.3 bits)
Medicago: description of AC146329.11


