

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC146329.10 - phase: 0
(68 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
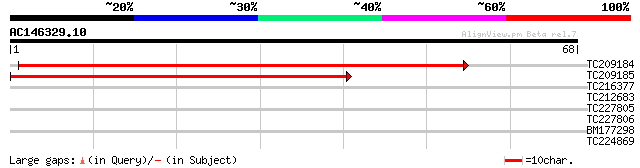
Score E
Sequences producing significant alignments: (bits) Value
TC209184 69 4e-13
TC209185 weakly similar to GB|AAL15380.1|16323334|AY059155 AT3g1... 62 4e-11
TC216377 28 0.91
TC212683 similar to UP|Q6R2J8 (Q6R2J8) Strubbelig receptor famil... 27 1.5
TC227805 26 3.5
TC227806 26 3.5
BM177298 25 5.9
TC224869 weakly similar to UP|ST14_SOLTU (Q41495) STS14 protein ... 25 5.9
>TC209184
Length = 466
Score = 69.3 bits (168), Expect = 4e-13
Identities = 31/54 (57%), Positives = 43/54 (79%)
Frame = +2
Query: 2 VAGNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLARHINVAAPNFTGIGS 55
VA AY+EGAQVGCLVKGGVQD+D+GD+S S+QF +SL+ ++N+ F +G+
Sbjct: 2 VACKAYMEGAQVGCLVKGGVQDVDEGDRSCSQQFKDSLSGYMNMLVKEFAKVGA 163
>TC209185 weakly similar to GB|AAL15380.1|16323334|AY059155 AT3g15355/MJK13_1
{Arabidopsis thaliana;} , partial (21%)
Length = 812
Score = 62.4 bits (150), Expect = 4e-11
Identities = 28/41 (68%), Positives = 36/41 (87%)
Frame = +3
Query: 1 MVAGNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLAR 41
+VA AY+EGAQVGCLVKGGVQD+D+GD+S S++F +SL R
Sbjct: 336 LVACKAYMEGAQVGCLVKGGVQDVDEGDRSCSQRFKDSLCR 458
>TC216377
Length = 2020
Score = 28.1 bits (61), Expect = 0.91
Identities = 15/41 (36%), Positives = 23/41 (55%)
Frame = +2
Query: 1 MVAGNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLAR 41
++A AY+EGA +GC GG + + K +S F LA+
Sbjct: 1529 LLACKAYLEGAPIGCGFGGGKAE-HENQKGTSTGFKIMLAK 1648
>TC212683 similar to UP|Q6R2J8 (Q6R2J8) Strubbelig receptor family 8, partial
(20%)
Length = 456
Score = 27.3 bits (59), Expect = 1.5
Identities = 18/57 (31%), Positives = 27/57 (46%)
Frame = +3
Query: 4 GNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLARHINVAAPNFTGIGSMLVQA 60
GN Y E Q+ ++ GG RQF+ S ++ +AAP IG +L +A
Sbjct: 258 GNGY*EDRQLSSVITGG------------RQFS*SCF*YVTLAAPKHCDIGWILCRA 392
>TC227805
Length = 1849
Score = 26.2 bits (56), Expect = 3.5
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = +2
Query: 1 MVAGNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLAR 41
++A AY+EGA +GC G + + K +S F LA+
Sbjct: 1400 LLACKAYLEGASIGCAFGSG-KTGHENQKGTSTGFKIMLAK 1519
>TC227806
Length = 683
Score = 26.2 bits (56), Expect = 3.5
Identities = 14/41 (34%), Positives = 22/41 (53%)
Frame = +2
Query: 1 MVAGNAYIEGAQVGCLVKGGVQDLDDGDKSSSRQFNNSLAR 41
++A AY+EGA +GC G + + K +S F LA+
Sbjct: 239 LLACKAYLEGASIGCAFGSG-KTGHENQKGTSTGFKIMLAK 358
>BM177298
Length = 437
Score = 25.4 bits (54), Expect = 5.9
Identities = 9/19 (47%), Positives = 14/19 (73%)
Frame = -2
Query: 15 CLVKGGVQDLDDGDKSSSR 33
C +K G+QDL+ G+K+ R
Sbjct: 166 CFLKYGIQDLNRGEKAKER 110
>TC224869 weakly similar to UP|ST14_SOLTU (Q41495) STS14 protein precursor,
partial (65%)
Length = 1276
Score = 25.4 bits (54), Expect = 5.9
Identities = 14/31 (45%), Positives = 19/31 (61%), Gaps = 1/31 (3%)
Frame = +2
Query: 4 GNAYIEG-AQVGCLVKGGVQDLDDGDKSSSR 33
G ++ G A+V L GGV++LD G SSR
Sbjct: 920 GEQHVRGEARVRGLYAGGVEELDGGGVRSSR 1012
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.135 0.382
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 2,202,265
Number of Sequences: 63676
Number of extensions: 19126
Number of successful extensions: 82
Number of sequences better than 10.0: 16
Number of HSP's better than 10.0 without gapping: 82
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 82
length of query: 68
length of database: 12,639,632
effective HSP length: 44
effective length of query: 24
effective length of database: 9,837,888
effective search space: 236109312
effective search space used: 236109312
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC146329.10


