

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145767.6 - phase: 0
(533 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
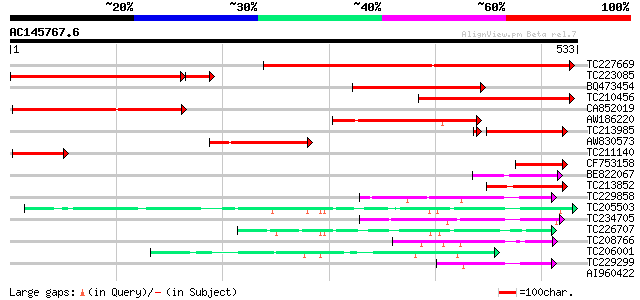
Score E
Sequences producing significant alignments: (bits) Value
TC227669 homologue to UP|Q9AVQ9 (Q9AVQ9) Phosphate transporter, ... 491 e-139
TC223085 homologue to UP|Q9AVR0 (Q9AVR0) Phosphate transporter, ... 295 4e-86
BQ473454 homologue to GP|24415831|gb phosphate transporter {Medi... 244 9e-65
TC210456 similar to UP|Q9AU01 (Q9AU01) Phosphate transporter 1, ... 201 5e-52
CA852019 homologue to GP|12697486|em phosphate transporter {Sesb... 186 2e-47
AW186220 154 9e-38
TC213985 homologue to UP|Q9AVR0 (Q9AVR0) Phosphate transporter, ... 134 2e-31
AW830573 117 2e-26
TC211140 homologue to UP|Q9AVQ9 (Q9AVQ9) Phosphate transporter, ... 102 6e-22
CF753158 85 9e-17
BE822067 64 2e-10
TC213852 62 8e-10
TC229858 weakly similar to UP|Q8VHD6 (Q8VHD6) Glucose transporte... 57 2e-08
TC205503 UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, complete 56 5e-08
TC234705 weakly similar to GB|CAD70577.1|32698459|MMU549317 solu... 55 1e-07
TC226707 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter... 49 6e-06
TC208766 similar to UP|STA_RICCO (Q10710) Sugar carrier protein ... 42 5e-04
TC206001 similar to UP|Q8VZI5 (Q8VZI5) AT5g18840/F17K4_90, parti... 42 7e-04
TC229299 similar to UP|GT10_HUMAN (O95528) Solute carrier family... 42 7e-04
AI960422 41 0.001
>TC227669 homologue to UP|Q9AVQ9 (Q9AVQ9) Phosphate transporter, partial
(55%)
Length = 1226
Score = 491 bits (1263), Expect = e-139
Identities = 243/293 (82%), Positives = 264/293 (89%)
Frame = +3
Query: 239 TARYTALVAKNAKQAAADMSKVLQVELEVEEEKVEKMTSDKRNSYGLFSKQFAARHGLAL 298
TARYTALVAKNAKQAA+DMSKVLQVE+E EE+K++ M + YGLFSK+FA RHGL L
Sbjct: 3 TARYTALVAKNAKQAASDMSKVLQVEVEAEEDKLQHMVESENQKYGLFSKEFAKRHGLHL 182
Query: 299 FGTCSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVP 358
GT TWFLLDIAFYSQNLFQKDIF+AIGWIPPA++MNAIHEVY+IARAQTLIALCSTVP
Sbjct: 183 VGTTVTWFLLDIAFYSQNLFQKDIFTAIGWIPPAQDMNAIHEVYRIARAQTLIALCSTVP 362
Query: 359 GYWFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIPYDHWSKEENRIGFVVIYSLTFFFAN 418
GYWFTVAFID +GRFAIQ+MGFFFMTVFMFALAIPY+HW K N IGFVV+YS TFFF+N
Sbjct: 363 GYWFTVAFIDIIGRFAIQLMGFFFMTVFMFALAIPYNHW-KNHNNIGFVVMYSFTFFFSN 539
Query: 419 FGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPT 478
FGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQS +P K D GYPT
Sbjct: 540 FGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSTNPNKVDHGYPT 719
Query: 479 GIGIKNSLIMLGVINFVGMLCTLLVPESKGKSLEELSGENEGEGAEATEQEGS 531
GIG+KNSLI+LGVINF GM+ TLLVPESKGKSLEELSGENE +GAEA E GS
Sbjct: 720 GIGVKNSLIVLGVINFFGMVFTLLVPESKGKSLEELSGENEDDGAEAIEMAGS 878
>TC223085 homologue to UP|Q9AVR0 (Q9AVR0) Phosphate transporter, partial
(35%)
Length = 634
Score = 295 bits (756), Expect(2) = 4e-86
Identities = 144/165 (87%), Positives = 153/165 (92%)
Frame = +3
Query: 1 MSGELGVLNALDVAKTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTR 60
M+GELGVLNALDVAKTQ YHFT IVIAGMGFFTDAYDLFCISLVTKLLGR+YYTEP +
Sbjct: 57 MAGELGVLNALDVAKTQWYHFTAIVIAGMGFFTDAYDLFCISLVTKLLGRLYYTEPGAPK 236
Query: 61 PGTLPPSAQSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGS 120
PGTLPP Q++ TGVAL GTLAGQLFFGWLGDK+GRKKVYGLTLILMVV S+ASGLSFGS
Sbjct: 237 PGTLPPRVQASATGVALCGTLAGQLFFGWLGDKMGRKKVYGLTLILMVVSSLASGLSFGS 416
Query: 121 SPKSVMATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIA 165
+ + VMATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIA
Sbjct: 417 TAEGVMATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIA 551
Score = 41.6 bits (96), Expect(2) = 4e-86
Identities = 19/27 (70%), Positives = 22/27 (81%)
Frame = +1
Query: 166 AVFAMQGFGILGGGIVALTVASIFDHK 192
AVFAMQGFGIL GG+VAL V+ +D K
Sbjct: 553 AVFAMQGFGILAGGVVALIVSYAYDQK 633
>BQ473454 homologue to GP|24415831|gb phosphate transporter {Medicago
sativa}, partial (55%)
Length = 426
Score = 244 bits (622), Expect = 9e-65
Identities = 115/125 (92%), Positives = 122/125 (97%)
Frame = +1
Query: 323 FSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGYWFTVAFIDHMGRFAIQMMGFFF 382
+SAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGYWFTVA ID+MGRFAIQ+MGFFF
Sbjct: 52 WSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGYWFTVALIDYMGRFAIQLMGFFF 231
Query: 383 MTVFMFALAIPYDHWSKEENRIGFVVIYSLTFFFANFGPNATTFVVPAEIFPARLRSTCH 442
MTVFMFALAIPY HWS+++NRIGFVV+YS TFFFANFGPNATTFVVPAEIFPARLRSTCH
Sbjct: 232 MTVFMFALAIPYHHWSEKDNRIGFVVMYSFTFFFANFGPNATTFVVPAEIFPARLRSTCH 411
Query: 443 GISAA 447
GISAA
Sbjct: 412 GISAA 426
>TC210456 similar to UP|Q9AU01 (Q9AU01) Phosphate transporter 1, partial
(29%)
Length = 617
Score = 201 bits (512), Expect = 5e-52
Identities = 96/147 (65%), Positives = 117/147 (79%)
Frame = +2
Query: 385 VFMFALAIPYDHWSKEENRIGFVVIYSLTFFFANFGPNATTFVVPAEIFPARLRSTCHGI 444
V+M ALA+PYDHW+ ++NRIGF VI ++TFFFANFGPNATTF+VPAEIFPAR RSTCHGI
Sbjct: 2 VYMCALAVPYDHWTHKDNRIGFRVIDAMTFFFANFGPNATTFIVPAEIFPARFRSTCHGI 181
Query: 445 SAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGIGIKNSLIMLGVINFVGMLCTLLVP 504
S+++GK GAIVGAFGFLY AQ+KD +K D YP GIG+KN+LI GV+N + T LVP
Sbjct: 182 SSSSGKLGAIVGAFGFLYLAQNKDKSKADARYPAGIGVKNALIA*GVVNILRFYFTFLVP 361
Query: 505 ESKGKSLEELSGENEGEGAEATEQEGS 531
E+ GKSLEE+S EN+ + E E S
Sbjct: 362 EANGKSLEEMSSENDEDVGTQEESEQS 442
>CA852019 homologue to GP|12697486|em phosphate transporter {Sesbania
rostrata}, partial (21%)
Length = 793
Score = 186 bits (473), Expect = 2e-47
Identities = 102/164 (62%), Positives = 114/164 (69%)
Frame = +2
Query: 3 GELGVLNALDVAKTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTRPG 62
G+LGVLNALDVAKTQ YHFT IVIAGMGFFTDAYDLFCISLVTKLLGR+YYT+ +PG
Sbjct: 305 GQLGVLNALDVAKTQWYHFTAIVIAGMGFFTDAYDLFCISLVTKLLGRLYYTDITNPKPG 484
Query: 63 TLPPSAQSAVTGVALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSP 122
LPP+ Q+AVTGVAL GTLAGQLFFGWLGDKLGRK ++ L SV G + P
Sbjct: 485 VLPPNVQAAVTGVALCGTLAGQLFFGWLGDKLGRKGLW-LNSYAYGGGSVP*GSLLETPP 661
Query: 123 KSVMATLCFFRFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAA 166
L F LGFG+ G YP E A KKT+G FI +
Sbjct: 662 GCHGHPLVSSGFGLGFGLAGTYPFLVQSWLEIATKKTQGHFIGS 793
>AW186220
Length = 425
Score = 154 bits (389), Expect = 9e-38
Identities = 73/143 (51%), Positives = 101/143 (70%), Gaps = 3/143 (2%)
Frame = +1
Query: 304 TWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGYWFT 363
TWFL+DI FYSQ LFQ +I+ ++ ++++ E + A Q +IA+CST+PGY+F+
Sbjct: 1 TWFLVDIVFYSQVLFQSEIYKR--YLDKKEDVDVYQETFHAAWIQAVIAVCSTIPGYFFS 174
Query: 364 VAFIDHMGRFAIQMMGFFFMTVFMFALAIPY-DHWSKEENRIG--FVVIYSLTFFFANFG 420
+ FID GR MMGFFFM + F++ IPY +W+ +E+ F+V+Y L FFFANFG
Sbjct: 175 MYFIDKWGRVKNPMMGFFFMALAFFSIGIPYYSYWTTKEHHKNKVFMVLYGLAFFFANFG 354
Query: 421 PNATTFVVPAEIFPARLRSTCHG 443
PN TTF+VPAE+FPAR RS+CHG
Sbjct: 355 PNTTTFIVPAELFPARFRSSCHG 423
>TC213985 homologue to UP|Q9AVR0 (Q9AVR0) Phosphate transporter, partial
(15%)
Length = 583
Score = 134 bits (337), Expect(2) = 2e-31
Identities = 67/76 (88%), Positives = 70/76 (91%)
Frame = +2
Query: 449 GKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGIGIKNSLIMLGVINFVGMLCTLLVPESKG 508
GKAGAIVGAFGFLYAAQSKDP+KTD GYPTGIGIKNSLIMLGVINF+GML TLLVPE+KG
Sbjct: 71 GKAGAIVGAFGFLYAAQSKDPSKTDAGYPTGIGIKNSLIMLGVINFIGMLFTLLVPEAKG 250
Query: 509 KSLEELSGENEGEGAE 524
KSLEELSGEN AE
Sbjct: 251 KSLEELSGENNENDAE 298
Score = 20.4 bits (41), Expect(2) = 2e-31
Identities = 7/7 (100%), Positives = 7/7 (100%)
Frame = +3
Query: 437 LRSTCHG 443
LRSTCHG
Sbjct: 33 LRSTCHG 53
>AW830573
Length = 304
Score = 117 bits (292), Expect = 2e-26
Identities = 58/96 (60%), Positives = 73/96 (75%)
Frame = +2
Query: 189 FDHKYKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTALVAK 248
+DHK+K+P++E++P ASL P F Y WR++ GA PAALTYYWRM MPET RYT LVAK
Sbjct: 20 YDHKFKLPSYEQHPYASL-DPAFVYDWRILTKAGA*PAALTYYWRMNMPETPRYTLLVAK 196
Query: 249 NAKQAAADMSKVLQVELEVEEEKVEKMTSDKRNSYG 284
AKQAAADMS VL +ELE E+E V+ T ++ +YG
Sbjct: 197 TAKQAAADMSGVLHLELEAEDENVKISTENESFTYG 304
>TC211140 homologue to UP|Q9AVQ9 (Q9AVQ9) Phosphate transporter, partial
(10%)
Length = 466
Score = 102 bits (253), Expect = 6e-22
Identities = 48/53 (90%), Positives = 51/53 (95%)
Frame = +1
Query: 3 GELGVLNALDVAKTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTE 55
G+LGVLNALDVAKTQ YHFT IVIAGMGFFTDAYDLFCISLVTKLLGR+YYT+
Sbjct: 301 GQLGVLNALDVAKTQWYHFTAIVIAGMGFFTDAYDLFCISLVTKLLGRLYYTD 459
>CF753158
Length = 473
Score = 84.7 bits (208), Expect = 9e-17
Identities = 42/49 (85%), Positives = 44/49 (89%)
Frame = -3
Query: 476 YPTGIGIKNSLIMLGVINFVGMLCTLLVPESKGKSLEELSGENEGEGAE 524
YPTGIGIKNSLIMLGVINF+GML TLLVPE+KGKSLEELSGEN AE
Sbjct: 387 YPTGIGIKNSLIMLGVINFIGMLFTLLVPEAKGKSLEELSGENNENDAE 241
>BE822067
Length = 610
Score = 63.5 bits (153), Expect = 2e-10
Identities = 37/85 (43%), Positives = 51/85 (59%), Gaps = 1/85 (1%)
Frame = -1
Query: 436 RLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGIGIKNSLIMLGVINFV 495
R RS+CHGIS GAI+G+ GFL+A+ K + GY IG+K SLI+LG + +
Sbjct: 610 RFRSSCHGISGXVXXVGAIIGSVGFLWASH----RKKEDGYXKXIGMKVSLIILGGVCLL 443
Query: 496 GMLCT-LLVPESKGKSLEELSGENE 519
GM+ T E+ G+SL+E E E
Sbjct: 442 GMVITYFFTRETMGRSLQENEVEIE 368
>TC213852
Length = 411
Score = 61.6 bits (148), Expect = 8e-10
Identities = 35/77 (45%), Positives = 49/77 (63%), Gaps = 1/77 (1%)
Frame = +3
Query: 449 GKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGIGIKNSLIMLGVINFVGMLCT-LLVPESK 507
GK GAI+G+ GFL+A+ K + GYP GIG++ +LI+LGV+ +GML T L E+
Sbjct: 36 GKVGAIIGSVGFLWASHKKK----ENGYPKGIGMEVTLIILGVVCLLGMLVTYLFTRETM 203
Query: 508 GKSLEELSGENEGEGAE 524
G+SLEE E+ E
Sbjct: 204 GRSLEENEVESVNHEVE 254
>TC229858 weakly similar to UP|Q8VHD6 (Q8VHD6) Glucose transporter 10 (Mus
musculus 13 days embryo heart cDNA, RIKEN full-length
enriched library, clone:D330001H08 product:solute
carrier family 2 (facilitated glucose transporter),
member 10, full insert sequence), partial (4%)
Length = 715
Score = 57.0 bits (136), Expect = 2e-08
Identities = 48/190 (25%), Positives = 83/190 (43%), Gaps = 5/190 (2%)
Frame = +3
Query: 330 PPAKEMNAIHEVYKIARAQTLIALCSTVPGYWFTVAFIDHMGR--FAIQMMGFFFMTVFM 387
P +M H ++A +LI G + IDH GR A+ +G +++ +
Sbjct: 87 PTIVQMAGFH-ANELALLLSLIVAGMNAAGTILGIYLIDHAGRKKLALSSLGGVIVSLVI 263
Query: 388 FALAIPYDHWSKEENRIGFVVIYSLTFFFANFGPNA--TTFVVPAEIFPARLRSTCHGIS 445
A A Y S G++ + L + F P + + +EI+P R C G+S
Sbjct: 264 LAFAF-YKQSSTSNELYGWLAVVGLALYIGFFSPGMGPVPWTLSSEIYPEEYRGICGGMS 440
Query: 446 AAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGIGIKNSLIMLGVINFVGMLCTLL-VP 504
A ++ + FL A+ GIGI ++ +++GVI V + L+ VP
Sbjct: 441 ATVCWVSNLIVSETFLSIAE-------------GIGIGSTFLIIGVIAVVAFVFVLVYVP 581
Query: 505 ESKGKSLEEL 514
E+KG + +E+
Sbjct: 582 ETKGLTFDEV 611
>TC205503 UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, complete
Length = 1928
Score = 55.8 bits (133), Expect = 5e-08
Identities = 121/558 (21%), Positives = 210/558 (36%), Gaps = 39/558 (6%)
Frame = +1
Query: 15 KTQLYHFTTIVIAGMGFFTDAYDLFCISLVTKLLGRIYYTEPNPTRPGTLPPSAQSAVTG 74
K Y F ++A M YD+ +S G Y + R + + G
Sbjct: 241 KRNKYAFACAMLASMTSILLGYDIGVMS------GAAIYIK----RDLKVSDEQIEILLG 390
Query: 75 VALVGTLAGQLFFGWLGDKLGRKKVYGLTLILMVVCSVASGLSFGSSPKSVMATLCFFRF 134
+ + +L G G D +GR+ GL + +V S G S L RF
Sbjct: 391 IINLYSLIGSCLAGRTSDWIGRRYTIGLGGAIFLVGSTLMGFYPHYS------FLMCGRF 552
Query: 135 WLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVASIFDHKYK 194
G GIG ++ +E + +RG + GIL G I
Sbjct: 553 VAGIGIGYALMIAPVYTAEVSPASSRGFLTSFPEVFINGGILLGYI-------------- 690
Query: 195 VPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTAL-----VAKN 249
N S L + WR++L GA+P+ + + MPE+ R+ + A+
Sbjct: 691 -----SNYGFSKLTLKVG--WRMMLGVGAIPSVVLTEGVLAMPESPRWLVMRGRLGEARK 849
Query: 250 AKQAAADMSKVLQVEL-EVEEEK-VEKMTSD-------KRNSYGLFSKQF-----AARH- 294
+D + Q+ L E+++ + + +D + N G++ + F A RH
Sbjct: 850 VLNKTSDSKEEAQLRLAEIKQAAGIPESCNDDVVQVNKQSNGEGVWKELFLYPTPAIRHI 1029
Query: 295 -----GLALFGTCSTWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQT 349
G+ F S + + YS +F+K + + + + + +T
Sbjct: 1030VIAALGIHFFQQASG--VDAVVLYSPRIFEKAGIT--------NDTHKLLATVAVGFVKT 1179
Query: 350 LIALCSTVPGYWFTVAFIDHMGRFAIQMMGFFFMTVFMFALAIP---YDHWSKE-----E 401
+ L +T FT +D +GR + + M + + LAI DH ++
Sbjct: 1180VFILAAT-----FT---LDRVGRRPLLLSSVGGMVLSLLTLAISLTVIDHSERKLMWAVG 1335
Query: 402 NRIGFVVIYSLTFFFANFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFL 461
+ I V+ Y TF + G T+V +EIFP RLR+ A + + V + FL
Sbjct: 1336SSIAMVLAYVATF---SIGAGPITWVYSSEIFPLRLRAQGAAAGVAVNRTTSAVVSMTFL 1506
Query: 462 YAAQSKDPTKTDKGYPTGIGIKNSLIMLGVINFVGMLCTLLV-PESKGKSLEELSG---- 516
++ I I + + I VG + V PE++GK+LE++ G
Sbjct: 1507SLTRA-------------ITIGGAFFLYCGIATVGWIFFYTVLPETRGKTLEDMEGSFGT 1647
Query: 517 -ENEGEGAEATEQEGSRV 533
++ ++A E E +V
Sbjct: 1648FRSKSNASKAVENENGQV 1701
>TC234705 weakly similar to GB|CAD70577.1|32698459|MMU549317 solute carrier
family 2 (facilitated glucose transporter), member 12
{Mus musculus;} , partial (5%)
Length = 992
Score = 54.7 bits (130), Expect = 1e-07
Identities = 48/198 (24%), Positives = 87/198 (43%), Gaps = 6/198 (3%)
Frame = +1
Query: 330 PPAKEMNAIHEVYKIARAQTLIALCSTVPGYWFTVAFIDHMGRFAIQMMGFFFMTVFMFA 389
P + I + K+ A + + T+ + ID +GR + M+ MTV +F
Sbjct: 1 PEIFQAAGIEDNSKLLAATVAVGVAKTI-FILVAIILIDKLGRKPLLMISTIGMTVCLFC 177
Query: 390 LAIPYDHWSKEENRIGFVVIY---SLTFFFANFGPNATTFVVPAEIFPARLRSTCHGISA 446
+ K I +++ ++ FF GP +V+ +EIFP R+R+ + A
Sbjct: 178 MGATLALLGKGSFAIALAILFVCGNVAFFSVGLGP--VCWVLTSEIFPLRVRAQASALGA 351
Query: 447 AAGKAGAIVGAFGFLYAAQSKDPTKTDKGYPTGIGIKNSLIMLGVINFVGMLCTL-LVPE 505
A + + + A FL +++ I + + + I+ + + + LVPE
Sbjct: 352 VANRVCSGLVAMSFLSVSEA-------------ISVAGTFFVFAAISALAIAFVVTLVPE 492
Query: 506 SKGKSLE--ELSGENEGE 521
+KGKSLE E+ +NE E
Sbjct: 493 TKGKSLEQIEMMFQNEYE 546
>TC226707 similar to UP|Q7XA50 (Q7XA50) Sorbitol-like transporter, partial
(62%)
Length = 2338
Score = 48.9 bits (115), Expect = 6e-06
Identities = 77/337 (22%), Positives = 131/337 (38%), Gaps = 37/337 (10%)
Frame = +3
Query: 215 WRLILMFGALPAALTYYWRMKMPETARYTALVAKN------------------AKQAAAD 256
WRL+L GA+P+ L + MPE+ R+ LVAK A+ AD
Sbjct: 60 WRLMLGVGAIPSILIGVAVLAMPESPRW--LVAKGRLGEAKRVLYKISESEEEARLRLAD 233
Query: 257 MSKVLQVELEVEEEKVEKMTSDKRNSYGLFSKQF-----AARH------GLALFGTCSTW 305
+ + + +++ V + S + + +G++ + F A RH G+ F +
Sbjct: 234 IKDTAGIPQDCDDDVV--LVSKQTHGHGVWRELFLHPTPAVRHIFIASLGIHFFAQATG- 404
Query: 306 FLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGYWFTVA 365
+ + YS +F+K + + + + +T+ L +T
Sbjct: 405 -IDAVVLYSPRIFEKAGIKSDNY--------RLLATVAVGFVKTVSILVATF-------- 533
Query: 366 FIDHMGRFAIQMMGFFFMTVFMFALAIPY---DHWSKEEN-----RIGFVVIYSLTFFFA 417
F+D GR + + + + + L + DH N I V+ Y TF
Sbjct: 534 FLDRAGRRVLLLCSVSGLILSLLTLGLSLTVVDHSQTTLNWAVGLSIAAVLSYVATF--- 704
Query: 418 NFGPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDKGYP 477
+ G T+V +EIFP RLR+ I AA + + V A FL K
Sbjct: 705 SIGSGPITWVYSSEIFPLRLRAQGVAIGAAVNRVTSGVIAMTFL---------SLQKAIT 857
Query: 478 TGIGIKNSLIMLGVINFVGMLCTLLVPESKGKSLEEL 514
G + GV + L+PE++GK+LEE+
Sbjct: 858 IGGAF---FLFAGVAAVAWIFHYTLLPETRGKTLEEI 959
>TC208766 similar to UP|STA_RICCO (Q10710) Sugar carrier protein A, partial
(55%)
Length = 1202
Score = 42.4 bits (98), Expect = 5e-04
Identities = 42/161 (26%), Positives = 70/161 (43%), Gaps = 6/161 (3%)
Frame = +1
Query: 361 WFTVAFIDHMGRFAIQMMGFFFMTV--FMFALAIPYDHWSKEENRIGF--VVIYSLTFFF 416
+ ++A +D +GR + + G M + A+ + + +E GF +V+ + F
Sbjct: 382 FISIATVDRLGRRVLLVSGGLQMITCQIIVAIILGVKFGADQELSKGFSILVVVVICLFV 561
Query: 417 ANFGPN--ATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGFLYAAQSKDPTKTDK 474
FG + + VP+EIFP +RS GI+ A + A FL S
Sbjct: 562 VAFGWSWGPLGWTVPSEIFPLEIRSAGQGITVAVNLLFTFIIAQAFLALLCS-------- 717
Query: 475 GYPTGIGIKNSLIMLGVINFVGMLCTLLVPESKGKSLEELS 515
+ GI L G I + + L +PE+KG +EE+S
Sbjct: 718 -FKFGI----FLFFAGWITIMTIFVYLFLPETKGIPIEEMS 825
>TC206001 similar to UP|Q8VZI5 (Q8VZI5) AT5g18840/F17K4_90, partial (66%)
Length = 951
Score = 42.0 bits (97), Expect = 7e-04
Identities = 68/341 (19%), Positives = 123/341 (35%), Gaps = 13/341 (3%)
Frame = +3
Query: 133 RFWLGFGIGGDYPLSATIMSEYANKKTRGAFIAAVFAMQGFGILGGGIVALTVASIFDHK 192
RF+ G+GIG + ++E A K RG + I+ GG V+ + S+ +
Sbjct: 51 RFFTGYGIGVISYVVPVYIAEIAPKNLRGGLATTNQLL----IVTGGSVSFLLGSVIN-- 212
Query: 193 YKVPTFEENPAASLLVPQFDYVWRLILMFGALPAALTYYWRMKMPETARYTALVAKNAKQ 252
WR + + G +P +PE+ R+ A V + K+
Sbjct: 213 ----------------------WRELALAGLVPCICLLVGLCFIPESPRWLAKVGRE-KE 323
Query: 253 AAADMSKVLQVELEVEEEKVEKM-------TSDKRNSYGLFSKQF--AARHGLALFGTCS 303
+S++ ++ +E E + + K LF ++ + G+ L
Sbjct: 324 FQLALSRLRGKHADISDEAAEILDYIETLESLPKTKLLDLFQSKYVHSVVIGVGLMACQQ 503
Query: 304 TWFLLDIAFYSQNLFQKDIFSAIGWIPPAKEMNAIHEVYKIARAQTLIALCSTVPGYWFT 363
+ + I FY+ +IF A G +A T+ C +P
Sbjct: 504 SVGINGIGFYT-----AEIFVAAG--------------LSSGKAGTIAYACIQIPFTLLG 626
Query: 364 VAFIDHMGRFAIQMMGF--FFMTVFMFALAIPYDHWSKEENRIGFVVIYSLTFFFANF-- 419
+D GR + M+ F+ F+ A A S + + + + A F
Sbjct: 627 AILMDKSGRRPLVMVSAAGTFLGCFVAAFAFFLKDQSLLPEWVPILAFAGVLIYIAAFSI 806
Query: 420 GPNATTFVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFGF 460
G + +V+ +EIFP L+ T + GA V ++ F
Sbjct: 807 GLGSVPWVIMSEIFPIHLKGTAGSLVVLVAWLGAWVVSYTF 929
>TC229299 similar to UP|GT10_HUMAN (O95528) Solute carrier family 2,
facilitated glucose transporter, member 10 (Glucose
transporter type 10), partial (6%)
Length = 1060
Score = 42.0 bits (97), Expect = 7e-04
Identities = 28/116 (24%), Positives = 54/116 (46%), Gaps = 3/116 (2%)
Frame = +2
Query: 402 NRIGFVVIYSLTFFFANFGPNATT--FVVPAEIFPARLRSTCHGISAAAGKAGAIVGAFG 459
++ G+ + L + F P T +VV +EI+P R R C GI++ ++ +
Sbjct: 512 SKYGWAALIGLALYIIFFSPGMGTVPWVVNSEIYPLRYRGVCGGIASTTVWISNLIVSES 691
Query: 460 FLYAAQSKDPTKTDKGYPTGIGIKNSLIMLGVINFVGMLCTLL-VPESKGKSLEEL 514
FL ++ +G + +M G++ V + ++ VPE+KG +EE+
Sbjct: 692 FLSLTKA-------------LGTAWTFMMFGIVAIVAIFFVIIFVPETKGVPMEEV 820
>AI960422
Length = 130
Score = 41.2 bits (95), Expect = 0.001
Identities = 18/43 (41%), Positives = 27/43 (61%)
Frame = +2
Query: 281 NSYGLFSKQFAARHGLALFGTCSTWFLLDIAFYSQNLFQKDIF 323
++Y L S ++ +HG LF TWFL+DIA YSQ L +++
Sbjct: 2 HAYPLLSWEYLRKHGPDLFACSFTWFLVDIAVYSQVLVHSELY 130
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.139 0.420
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 22,278,226
Number of Sequences: 63676
Number of extensions: 298977
Number of successful extensions: 2205
Number of sequences better than 10.0: 72
Number of HSP's better than 10.0 without gapping: 2170
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2191
length of query: 533
length of database: 12,639,632
effective HSP length: 102
effective length of query: 431
effective length of database: 6,144,680
effective search space: 2648357080
effective search space used: 2648357080
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 61 (28.1 bits)
Medicago: description of AC145767.6


