

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
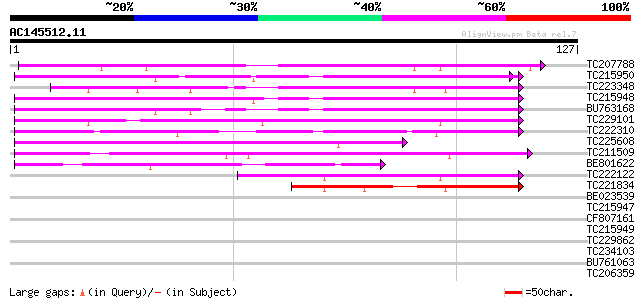
Query= AC145512.11 - phase: 0
(127 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC207788 homologue to UP|Q8W2C0 (Q8W2C0) Functional candidate re... 79 6e-16
TC215950 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candida... 70 3e-13
TC223348 UP|Q945R9 (Q945R9) NR1 (Fragment), complete 65 6e-12
TC215948 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candida... 64 2e-11
BU763168 similar to GP|18033111|gb functional candidate resistan... 63 3e-11
TC229101 disease resistance-like protein 61 2e-10
TC222310 57 2e-09
TC225608 UP|Q84ZU5 (Q84ZU5) R 8 protein, complete 49 8e-07
TC211509 47 2e-06
BE801622 similar to GP|18033111|gb functional candidate resistan... 44 3e-05
TC222122 similar to UP|Q84ZV7 (Q84ZV7) R 12 protein, partial (7%) 40 2e-04
TC221834 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candida... 40 3e-04
BE023539 38 0.001
TC215947 similar to UP|Q945R9 (Q945R9) NR1 (Fragment), partial (... 37 0.002
CF807161 35 0.009
TC215949 similar to UP|Q8W2C0 (Q8W2C0) Functional candidate resi... 32 0.078
TC229862 similar to UP|Q940L9 (Q940L9) At2g44770/F16B22.26, part... 29 0.15
TC234103 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candida... 31 0.17
BU761063 30 0.23
TC206359 similar to UP|Q9LY33 (Q9LY33) BZIP protein, partial (84%) 30 0.39
>TC207788 homologue to UP|Q8W2C0 (Q8W2C0) Functional candidate resistance
protein KR1, complete
Length = 3949
Score = 79.0 bits (193), Expect = 6e-16
Identities = 54/140 (38%), Positives = 77/140 (54%), Gaps = 22/140 (15%)
Frame = +1
Query: 3 VSFWFCNKFPSIALGVV--------SAYTWGYLEH-PARVIINDNTFFYTHGRKIDRCSR 53
+SFWF NKFP+IA+ + S+ W + + +VIIN N + C+
Sbjct: 2905 ISFWFRNKFPAIAICHIIKRVAEFSSSRGWTFRPNIRTKVIINGNANLFNSVVLGSDCTC 3084
Query: 54 PDTYHLHLFHMQVEYFNGNMDKALLENKWNHAEVDF-GFPFMF------SGIHVLKEKSN 106
LF ++ E N+D+ALLEN+WNHAEV GF F F +G+HVLK++SN
Sbjct: 3085 -------LFDLRGERVTDNLDEALLENEWNHAEVTCPGFTFTFAPTFIKTGLHVLKQESN 3243
Query: 107 MKDIRFTNP------ENDAN 120
M+DIRF++P +ND N
Sbjct: 3244 MEDIRFSDPCRKTKLDNDFN 3303
>TC215950 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidate resistance
protein KR1, partial (23%)
Length = 1891
Score = 70.1 bits (170), Expect = 3e-13
Identities = 43/116 (37%), Positives = 66/116 (56%), Gaps = 2/116 (1%)
Frame = +1
Query: 2 SVSFWFCNKFPSIALGVVSAYTWGYLEHPA--RVIINDNTFFYTHGRKIDRCSRPDTYHL 59
S+ FWF N+FP+I +V ++ Y VIIN + H R D C T
Sbjct: 1390 SIFFWFRNEFPAITFCIVKSHFEAYSSDSLVLSVIINKK-HEHKHDRFHDGCFSK-TPST 1563
Query: 60 HLFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMFSGIHVLKEKSNMKDIRFTNP 115
+F +Q++ N+D+ + +++WNHAE+ + GIHVLKE+S+M+DIRFT+P
Sbjct: 1564 SIFRLQMK---DNLDEEISKSEWNHAEIVCNLSWDECGIHVLKEQSSMEDIRFTDP 1722
Score = 62.0 bits (149), Expect = 7e-11
Identities = 40/113 (35%), Positives = 63/113 (55%), Gaps = 1/113 (0%)
Frame = +1
Query: 2 SVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSR-PDTYHLH 60
S+ FWF NKFP+I + + + + + L V IND + C P T
Sbjct: 85 SIVFWFRNKFPAIIVCIDTEFCFDELA--VNVFINDEDESNHVSLDVTECCMGPSTA--- 249
Query: 61 LFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMFSGIHVLKEKSNMKDIRFT 113
+F +++E + +D+ LL+N+WN AE+ + + IHVLKE+S+M+DIRFT
Sbjct: 250 VFRLKMEDY---LDEELLKNEWNRAEIGCEYLWDEYAIHVLKEQSSMEDIRFT 399
>TC223348 UP|Q945R9 (Q945R9) NR1 (Fragment), complete
Length = 688
Score = 65.5 bits (158), Expect = 6e-12
Identities = 50/123 (40%), Positives = 68/123 (54%), Gaps = 17/123 (13%)
Frame = +3
Query: 10 KFPSIAL--------GVVSAYTWGYL-EHPARVIINDNT-FFYTHGRKIDRCSRPDTYHL 59
KFP+IA+ S+ W Y + +VIIN N FY+ D CS
Sbjct: 3 KFPAIAICHSIKRVPEFSSSRGWTYRPDIRTKVIINGNANLFYSMDLGSD-CSV------ 161
Query: 60 HLFHMQVEYFNGNMDKALLENKWNHAEVDF-GFPFMFS------GIHVLKEKSNMKDIRF 112
LF + + N+D+ALLEN+WNHAEV GF F F+ G+HVLK++SNM+DIRF
Sbjct: 162 -LFDPRDDKVTDNLDEALLENEWNHAEVTCPGFTFTFAPTFIKTGLHVLKQESNMEDIRF 338
Query: 113 TNP 115
++P
Sbjct: 339 SDP 347
>TC215948 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidate
resistance protein KR1, partial (19%)
Length = 1464
Score = 63.5 bits (153), Expect = 2e-11
Identities = 41/116 (35%), Positives = 65/116 (55%), Gaps = 2/116 (1%)
Frame = +2
Query: 2 SVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSR--PDTYHL 59
S+ FWF NKFP I + +V++ Y + +I + + H R S P T
Sbjct: 836 SIFFWFPNKFPVITVCIVTSGPKKYSNYLVLNVIINKKHKHRHQRFYSNGSNAIPSTT-- 1009
Query: 60 HLFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMFSGIHVLKEKSNMKDIRFTNP 115
+F +Q++ N+D+ L +++WN AE+ + GIHVLKEKS+M+DIRF++P
Sbjct: 1010-VFRLQMK---DNLDEELSKSEWNLAEIVCEDSWAAYGIHVLKEKSSMEDIRFSDP 1165
>BU763168 similar to GP|18033111|gb functional candidate resistance protein
KR1 {Glycine max}, partial (5%)
Length = 421
Score = 63.2 bits (152), Expect = 3e-11
Identities = 45/124 (36%), Positives = 66/124 (52%), Gaps = 10/124 (8%)
Frame = +3
Query: 2 SVSFWFCNKFPSIALGVVSAYTWGYLEHPA-RVIINDNT---------FFYTHGRKIDRC 51
S+ FWF NKFP+I + V + VIIN + FF+ C
Sbjct: 57 SIFFWFRNKFPAIVVCFVKEDFLNFTSDLVLSVIINGHEHQHKPLFGGFFFE-----SPC 221
Query: 52 SRPDTYHLHLFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMFSGIHVLKEKSNMKDIR 111
+ LFH+Q++ N+D+ALLEN+WN AE+ +G +GIHVLK++S++ DIR
Sbjct: 222 TA-------LFHLQMK---DNLDEALLENEWNLAEIVYGDLCDENGIHVLKKQSSVDDIR 371
Query: 112 FTNP 115
F +P
Sbjct: 372 FIDP 383
>TC229101 disease resistance-like protein
Length = 2755
Score = 60.8 bits (146), Expect = 2e-10
Identities = 43/127 (33%), Positives = 64/127 (49%), Gaps = 13/127 (10%)
Frame = +1
Query: 2 SVSFWFCNKFPSIAL---GVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSRPD-TY 57
S+SFWF NKFP I+L G++ + +G V IN N R+ P T
Sbjct: 2236 SISFWFRNKFPVISLCLAGLMHKHPFGL---KPIVSINGNKMKTEFQRRWFYFEFPVLTD 2406
Query: 58 HLHLFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMF---------SGIHVLKEKSNMK 108
H+ +F + F N+D+ + EN WNH V F + +G+HV+K KS+++
Sbjct: 2407 HILIFGERQIKFEDNVDEVVSENDWNHVVVSVDVDFKWNPTEPLVVRTGLHVIKPKSSVE 2586
Query: 109 DIRFTNP 115
DIRF +P
Sbjct: 2587 DIRFIDP 2607
>TC222310
Length = 811
Score = 57.0 bits (136), Expect = 2e-09
Identities = 44/119 (36%), Positives = 62/119 (51%), Gaps = 5/119 (4%)
Frame = +1
Query: 2 SVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIIN-DNTFFYTHGRKIDRCSRPDTYHLH 60
S+SFWF N FPSI+LGVV A YL +R IN +N + G + + +
Sbjct: 169 SISFWFRNNFPSISLGVV-AGPKSYLNVHSRFRININNNITFNLGIR--------NHLVQ 321
Query: 61 LFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFM----FSGIHVLKEKSNMKDIRFTNP 115
+ H +VE ++ NKWN E +P GIHVL++ +NM+DI+FTNP
Sbjct: 322 VRHHRVEL--ASIPIGFWGNKWNRVECTV-YPLQEFIKQIGIHVLEQGNNMEDIQFTNP 489
>TC225608 UP|Q84ZU5 (Q84ZU5) R 8 protein, complete
Length = 3084
Score = 48.5 bits (114), Expect = 8e-07
Identities = 30/90 (33%), Positives = 40/90 (44%), Gaps = 2/90 (2%)
Frame = +1
Query: 2 SVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSRPDTYHLHL 61
S+SFWF NKFP+ L + A + G+ V IN + D S H H+
Sbjct: 2746 SISFWFRNKFPAKLLCLHIAPSTGFFIRYPEVFINGKFQEFESHETDDTESMLGLDHTHI 2925
Query: 62 FHMQVEYFNGN--MDKALLENKWNHAEVDF 89
F +Q F N K E +WNH EV +
Sbjct: 2926 FDLQAYAFKNNNLFKKVAWEKEWNHVEVTY 3015
>TC211509
Length = 880
Score = 47.4 bits (111), Expect = 2e-06
Identities = 36/135 (26%), Positives = 60/135 (43%), Gaps = 19/135 (14%)
Frame = +2
Query: 2 SVSFWFCNKFPSIALGVVSAYTWGYLEHPARVIINDNTFFYTHGRK---IDRCS--RPDT 56
S++FW +FP I + V ++G LE+ +G K +RC T
Sbjct: 41 SITFWGRERFPRICVCV----SFGMLENSLHHHFQVTFCIVINGHKRILSNRCYDWSVQT 208
Query: 57 YHLHLFHMQVEYFNGNMDKALLENKWNHAEVDFGFPFMFSG--------------IHVLK 102
H+ LF + ++ L+++ WNH E++ + G IHV +
Sbjct: 209 DHVWLFDLTALVSYEDLRGTLVKSDWNHVEIEMEWNCCIQGDHGPTRMAIVKWYGIHVYR 388
Query: 103 EKSNMKDIRFTNPEN 117
++S M+DI FTNP+N
Sbjct: 389 QESKMEDISFTNPKN 433
>BE801622 similar to GP|18033111|gb functional candidate resistance protein
KR1 {Glycine max}, partial (6%)
Length = 391
Score = 43.5 bits (101), Expect = 3e-05
Identities = 31/88 (35%), Positives = 43/88 (48%), Gaps = 5/88 (5%)
Frame = +2
Query: 2 SVSFWFCNKFPSIALGVVSAYTWGYLEHP-----ARVIINDNTFFYTHGRKIDRCSRPDT 56
S+SFWF NKFP G V G ++ ++VIIN N +F G +
Sbjct: 158 SISFWFRNKFP----GKVLCLVIGPMDDDSGMLISKVIINGNKYFRGSGYFMMGMD---- 313
Query: 57 YHLHLFHMQVEYFNGNMDKALLENKWNH 84
H +LF +Q+ F N+ LEN+WNH
Sbjct: 314 -HTYLFDLQIMEFEDNL-YVPLENEWNH 391
>TC222122 similar to UP|Q84ZV7 (Q84ZV7) R 12 protein, partial (7%)
Length = 473
Score = 40.4 bits (93), Expect = 2e-04
Identities = 25/85 (29%), Positives = 37/85 (43%), Gaps = 21/85 (24%)
Frame = +1
Query: 52 SRPDTYHLHLFHMQVEYF--NGNMDKALLENKWNHAEVDFGFPFMF-------------- 95
SR + H ++F +Q F N ++ E +WNH EV + +
Sbjct: 22 SRLNLDHTYIFDLQASAFINNNRFEEMAREKEWNHVEVRYQSVLAYEKEKREEGVLDLES 201
Query: 96 -----SGIHVLKEKSNMKDIRFTNP 115
SGIH+ KE S +DIRF +P
Sbjct: 202 SIIKASGIHIFKESSMEEDIRFDDP 276
>TC221834 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidate resistance
protein KR1, partial (45%)
Length = 3472
Score = 40.0 bits (92), Expect = 3e-04
Identities = 27/58 (46%), Positives = 36/58 (61%), Gaps = 6/58 (10%)
Frame = +3
Query: 64 MQVEYF---NGNMDKALL--ENKWNHAEVDFGFPFMFS-GIHVLKEKSNMKDIRFTNP 115
M + YF NG+M +LL + KWN E PF+ G+HV+K KS+M DIRFT+P
Sbjct: 2973 MWMRYFRKMNGSMWWSLLIFDFKWNPIE-----PFVVQIGLHVIKPKSSMDDIRFTDP 3131
Score = 25.8 bits (55), Expect = 5.6
Identities = 10/32 (31%), Positives = 16/32 (49%)
Frame = +2
Query: 62 FHMQVEYFNGNMDKALLENKWNHAEVDFGFPF 93
F + + + N+D+ L EN+W H V F
Sbjct: 2942 FWWKTDRISNNVDEVLSENEWKHVVVSIDI*F 3037
>BE023539
Length = 324
Score = 38.1 bits (87), Expect = 0.001
Identities = 26/93 (27%), Positives = 45/93 (47%), Gaps = 5/93 (5%)
Frame = +2
Query: 2 SVSFWFCNKFPSIALGVVSA---YTWGYLEHPARVIINDNTFFYTHGRK--IDRCSRPDT 56
S+SFWF N+FP L ++ A YT+ + V IN G + + R
Sbjct: 8 SISFWFRNEFPDNVLCLLLARVEYTYKCISK-LTVFINGKRHKIASGWEDWMTTEVRKAK 184
Query: 57 YHLHLFHMQVEYFNGNMDKALLENKWNHAEVDF 89
+ +LF ++ + G++ + LE +WNH E+ +
Sbjct: 185LNTYLFDLKSSFRLGDLSEVGLEKEWNHVEISY 283
>TC215947 similar to UP|Q945R9 (Q945R9) NR1 (Fragment), partial (11%)
Length = 536
Score = 37.0 bits (84), Expect = 0.002
Identities = 15/22 (68%), Positives = 21/22 (95%)
Frame = +2
Query: 97 GIHVLKEKSNMKDIRFTNPEND 118
GIHVLKE+S+M+DIRF++P+ D
Sbjct: 8 GIHVLKEQSSMEDIRFSDPDPD 73
>CF807161
Length = 420
Score = 35.0 bits (79), Expect = 0.009
Identities = 15/19 (78%), Positives = 18/19 (93%)
Frame = -1
Query: 97 GIHVLKEKSNMKDIRFTNP 115
GIHVLKE S+M+DIRFT+P
Sbjct: 393 GIHVLKELSSMEDIRFTDP 337
>TC215949 similar to UP|Q8W2C0 (Q8W2C0) Functional candidate resistance protein
KR1, partial (30%)
Length = 1805
Score = 32.0 bits (71), Expect = 0.078
Identities = 11/23 (47%), Positives = 20/23 (86%)
Frame = +2
Query: 2 SVSFWFCNKFPSIALGVVSAYTW 24
S+SFWF N+FP+I + +V+++T+
Sbjct: 1619 SISFWFRNEFPAIIVCIVTSFTF 1687
>TC229862 similar to UP|Q940L9 (Q940L9) At2g44770/F16B22.26, partial (89%)
Length = 1295
Score = 29.3 bits (64), Expect(2) = 0.15
Identities = 16/46 (34%), Positives = 21/46 (44%)
Frame = +1
Query: 16 LGVVSAYTWGYLEHPARVIINDNTFFYTHGRKIDRCSRPDTYHLHL 61
LGVV Y W + N + FFY ++ID+C HL L
Sbjct: 361 LGVVVLYHWRIFSSLLGISRNPSRFFYASRKEIDQCGNT---HLQL 489
Score = 20.4 bits (41), Expect(2) = 0.15
Identities = 7/27 (25%), Positives = 13/27 (47%)
Frame = +3
Query: 91 FPFMFSGIHVLKEKSNMKDIRFTNPEN 117
+PF +G+++ M D+ P N
Sbjct: 474 YPFAVAGVNITYMLIQMLDLEAVKPRN 554
>TC234103 weakly similar to UP|Q8W2C0 (Q8W2C0) Functional candidate
resistance protein KR1, partial (3%)
Length = 444
Score = 30.8 bits (68), Expect = 0.17
Identities = 12/24 (50%), Positives = 18/24 (75%)
Frame = +1
Query: 99 HVLKEKSNMKDIRFTNPENDANIV 122
H+LK++S+M+D RFTNP +V
Sbjct: 1 HLLKQESSMEDFRFTNPFRKRKLV 72
>BU761063
Length = 443
Score = 30.4 bits (67), Expect = 0.23
Identities = 13/15 (86%), Positives = 15/15 (99%)
Frame = +2
Query: 97 GIHVLKEKSNMKDIR 111
GIHVLKEKS+M+DIR
Sbjct: 44 GIHVLKEKSSMEDIR 88
>TC206359 similar to UP|Q9LY33 (Q9LY33) BZIP protein, partial (84%)
Length = 1541
Score = 29.6 bits (65), Expect = 0.39
Identities = 19/50 (38%), Positives = 24/50 (48%), Gaps = 2/50 (4%)
Frame = -2
Query: 47 KIDRCSRPDTYHLHLFHMQVEYFNGNMDKALLENKWNHAE--VDFGFPFM 94
K RCS + L LFH+ Y N+DKAL + N V FPF+
Sbjct: 946 KFLRCSLGSFFDLELFHLLGRYHVDNIDKALDGFRCNRVRDTVKCEFPFI 797
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.326 0.140 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 7,424,212
Number of Sequences: 63676
Number of extensions: 111145
Number of successful extensions: 642
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 622
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 625
length of query: 127
length of database: 12,639,632
effective HSP length: 86
effective length of query: 41
effective length of database: 7,163,496
effective search space: 293703336
effective search space used: 293703336
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.7 bits)
S2: 53 (25.0 bits)
Medicago: description of AC145512.11


