

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145331.12 - phase: 0
(419 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
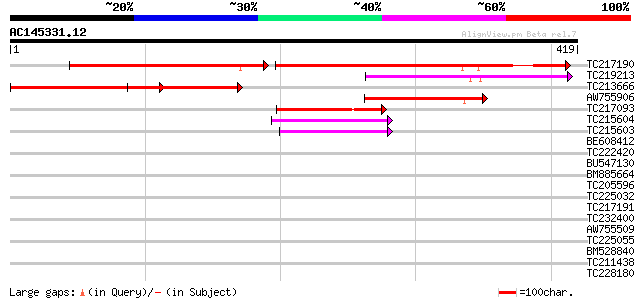
Score E
Sequences producing significant alignments: (bits) Value
TC217190 similar to UP|Q761Z6 (Q761Z6) BRI1-KD interacting prote... 169 5e-78
TC219213 114 6e-26
TC213666 114 1e-25
AW755906 85 5e-17
TC217093 53 2e-07
TC215604 similar to UP|Q84WL6 (Q84WL6) At3g23090, partial (44%) 47 1e-05
TC215603 similar to UP|Q7XYW3 (Q7XYW3) Seed specific protein Bn1... 47 2e-05
BE608412 35 0.048
TC222420 35 0.082
BU547130 34 0.14
BM885664 similar to GP|13539605|emb cyclophilin-RNA interacting ... 33 0.31
TC205596 similar to GB|AAO63377.1|28950907|BT005313 At4g34530 {A... 33 0.31
TC225032 homologue to UP|Q9LLL9 (Q9LLL9) Plasma membrane aquapor... 33 0.31
TC217191 32 0.53
TC232400 weakly similar to UP|Q8W2K5 (Q8W2K5) Phragmoplastin-int... 32 0.53
AW755509 homologue to PIR|G96681|G966 protein F1E22.8 [imported]... 32 0.70
TC225055 homologue to UP|Q9LLL9 (Q9LLL9) Plasma membrane aquapor... 31 0.91
BM528840 31 1.2
TC211438 UP|GLCX_SOYBN (P11827) Beta-conglycinin, alpha' chain p... 30 2.0
TC228180 similar to UP|Q9SR11 (Q9SR11) F7O18.11 protein, partial... 30 2.0
>TC217190 similar to UP|Q761Z6 (Q761Z6) BRI1-KD interacting protein 118
(Fragment), partial (32%)
Length = 1637
Score = 169 bits (427), Expect(2) = 5e-78
Identities = 124/233 (53%), Positives = 142/233 (60%), Gaps = 15/233 (6%)
Frame = +3
Query: 197 LRKGESSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLKSLGRA*LLKQLHYQLSIRSLLL 256
LR+ +SS+L LRKRF QR+ RRV CKQ PRR KKPRL+ L + * LKQL Q S RSLLL
Sbjct: 576 LREEKSSTLSLRKRFTQRKWRRVTCKQKPRRTKKPRLRCLEKV*DLKQLQCQASTRSLLL 755
Query: 257 LRWS*RRYQLLEQNLPSSVGRRLQQIQNLMETVAAVLDKVV*V*MRKCHRATPLQELPLP 316
L S* R QL E NLPS V RR+Q IQ+ ET+A VLDKVV*V MRKC R TPL L L
Sbjct: 756 LGRS*ERCQLPELNLPSLVARRVQ*IQSQKETLATVLDKVV*VWMRKCLRLTPLMALVLF 935
Query: 317 IKRSRYGSPFLLDSLLK---GPIQQL-HLHLRQ----------QRKILAYPKVQGKRKLR 362
I+RS S FL D LLK PIQ+L HLRQ R L YP +GKRK +
Sbjct: 936 IQRSHNESLFLHDWLLKRSVHPIQRL*EPHLRQ*MVAKPPYQV*RLKLPYPMQEGKRKFK 1115
Query: 363 -LSQPMKKTVPCQVTQMLLYLRMRFLQINQVKNFM*MGT*WLKRILSSCCHKN 414
L QP+K+T+P + QVK F+*M * LKR SS HKN
Sbjct: 1116*LQQPLKRTMP--------------Y*MRQVKFFL*MVI*SLKRNHSSTWHKN 1232
Score = 140 bits (354), Expect(2) = 5e-78
Identities = 82/164 (50%), Positives = 106/164 (64%), Gaps = 17/164 (10%)
Frame = +2
Query: 45 ETFVPDGNSENINQLESTATGNSAMKEI-EGSNDNVDGSNLTVSKEKEVKIKVSTEQSRA 103
ET +GN EN Q +STAT S+ +EI EGSNDN+ +N+T+SKE+E +I TEQ +
Sbjct: 2 ETAATNGNLENFIQYDSTATDYSSKEEIKEGSNDNIYMNNVTISKEEEAEIIDRTEQLKV 181
Query: 104 QKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSND 163
KGP KNKNAK S G +AS VK +K GKD++ + +VSNGT A DS PRQPIKNRS +D
Sbjct: 182 GKGPAKNKNAKPPSPRGSHASSVKKNKDGKDEEVASSVSNGTFASDSHPRQPIKNRSLSD 361
Query: 164 RQSQLS----------------KKKPKSLKKGPLDKVQGEGESS 191
+Q++LS K +P+ KK P D +QGE ESS
Sbjct: 362 KQARLSKHPGKSNAAHSEESMEKTRPQLSKKDPHDNLQGEAESS 493
>TC219213
Length = 883
Score = 114 bits (286), Expect = 6e-26
Identities = 88/171 (51%), Positives = 102/171 (59%), Gaps = 18/171 (10%)
Frame = +3
Query: 264 YQLLEQNLPSSVGRRLQQIQNLMETVAAVLDKVV*V*MRKCHRATPLQE-LPLPIKRSRY 322
YQL EQN PSSV RR+QQIQ+ MET AAV D +V*V MRKC R T L++ L L IKRS
Sbjct: 3 YQLQEQNHPSSVVRRVQQIQSQMETTAAVHD*LV*VWMRKCQRVTSLRDLLLLSIKRSHN 182
Query: 323 GSPFLLDSLLKGPIQQL----HLHLRQQ-----------RKILAYPKVQGKRKLRLSQPM 367
FLL LKG + Q+ LH R + I YP +Q +RK R Q M
Sbjct: 183 EDLFLLGWPLKGTVYQIQGLHQLHPRLSKMKSPPCLVLLKNIPTYPMLQERRKPRRLQQM 362
Query: 368 KKTVPCQVTQMLLYLRM-RFLQINQVKN-FM*MGT*WLKRILSSCCHKNRS 416
K+ VPC V Q++L M FLQI+QVK M*M T* LKRILSS KN S
Sbjct: 363 KRKVPCPVKQVMLCC*MWFFLQISQVKKCLM*MVT*QLKRILSSPWQKNLS 515
>TC213666
Length = 548
Score = 114 bits (284), Expect = 1e-25
Identities = 63/86 (73%), Positives = 71/86 (82%), Gaps = 1/86 (1%)
Frame = +1
Query: 88 KEKEVKIKVSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSA 147
++KEVKI T+QSRA KG VKNKNAK SSSGV+ASLV SKIGKDK+AS +VSNGTSA
Sbjct: 286 RKKEVKISDQTKQSRAPKGLVKNKNAKAPSSSGVHASLVNKSKIGKDKEASSSVSNGTSA 465
Query: 148 LDSRPRQPIK-NRSSNDRQSQLSKKK 172
LDSRPRQ K +RS NDRQ+QLSK K
Sbjct: 466 LDSRPRQSTKSSRSFNDRQTQLSKPK 543
Score = 111 bits (278), Expect = 5e-25
Identities = 57/115 (49%), Positives = 78/115 (67%), Gaps = 1/115 (0%)
Frame = +3
Query: 1 MDPVNSLPADGLDDVHQNGVHDEPSNSGED-AVSNDLDPHVTVNTETFVPDGNSENINQL 59
MDP+N LPADG++ VHQNGVHDEPSNSG+D VS DLDP VT T +GN +N +Q
Sbjct: 21 MDPINLLPADGVEVVHQNGVHDEPSNSGDDDGVSYDLDPSVTETAATVALNGNFDNFHQS 200
Query: 60 ESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVKNKNAK 114
+S A+ NS + EI+ SNDN+DG+N+T+ KE+ S + ++ KG + + K
Sbjct: 201 DSAASDNSFVAEIKESNDNIDGTNMTIPKEEGS*NIRSNKAIKSTKGSCQEQECK 365
>AW755906
Length = 440
Score = 85.1 bits (209), Expect = 5e-17
Identities = 57/96 (59%), Positives = 66/96 (68%), Gaps = 5/96 (5%)
Frame = +1
Query: 263 RYQLLEQNLPSSVGRRLQQIQNLMETVAAVLDKVV*V*MRKCHRATPLQE-LPLPIKRSR 321
RYQL EQN PS V RR+QQIQ+ M T+AAV D +V*V MRKC R PL++ L L IKRSR
Sbjct: 106 RYQLQEQNHPSLVVRRVQQIQSQMATIAAVHD*LV*VWMRKCQRVIPLRDLLLLSIKRSR 285
Query: 322 YGSPFLLDSLLKG---PIQQLH-LHLRQQRKILAYP 353
FLLD LLKG IQ+LH LH RQ K+ + P
Sbjct: 286 NEDLFLLDWLLKGTVYQIQELH*LHPRQLSKMKSLP 393
>TC217093
Length = 1829
Score = 53.1 bits (126), Expect = 2e-07
Identities = 32/81 (39%), Positives = 52/81 (63%)
Frame = +3
Query: 198 RKGESSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLKSLGRA*LLKQLHYQLSIRSLLLL 257
++G SSS +++FR+R++R++ K+N R+ ++ R + GR *L K L Q+S R+ LL
Sbjct: 1011 KRGRSSSQS*KRKFRKRKQRKLTSKRNQRKTRRLRSSN*GRP*LSKPLQCQVSTRN-LLQ 1187
Query: 258 RWS*RRYQLLEQNLPSSVGRR 278
+ +*RRY QNL S G +
Sbjct: 1188 KLN*RRYL*RAQNLQSLEGTK 1250
>TC215604 similar to UP|Q84WL6 (Q84WL6) At3g23090, partial (44%)
Length = 1145
Score = 47.4 bits (111), Expect = 1e-05
Identities = 36/91 (39%), Positives = 47/91 (51%), Gaps = 1/91 (1%)
Frame = +3
Query: 194 PFSLRK-GESSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLKSLGRA*LLKQLHYQLSIR 252
P +++K G S RK +Q + RR+ K PR+ + S GR *LLKQ Q
Sbjct: 855 PLNVQKNGRSFIQN*RKSSKQWKLRRIKVKLGPRKRWRKLSSS*GRV*LLKQALCQAFTM 1034
Query: 253 SLLLLRWS*RRYQLLEQNLPSSVGRRLQQIQ 283
LLR +*R +Q EQNL S VG R +Q
Sbjct: 1035KGPLLRLN*RSFQPREQNLQSWVGERATMVQ 1127
>TC215603 similar to UP|Q7XYW3 (Q7XYW3) Seed specific protein Bn15D14A
(Fragment), partial (29%)
Length = 663
Score = 47.0 bits (110), Expect = 2e-05
Identities = 33/84 (39%), Positives = 40/84 (47%)
Frame = +1
Query: 200 GESSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLKSLGRA*LLKQLHYQLSIRSLLLLRW 259
G RK +Q + RR+ K PR+ + S GR *LLKQ Q LLR
Sbjct: 1 GRXXXXN*RKSSKQWKLRRIKVKLGPRKRWRKLSSS*GRV*LLKQALCQAFTMKGPLLRL 180
Query: 260 S*RRYQLLEQNLPSSVGRRLQQIQ 283
+*R YQ EQNL S G R +Q
Sbjct: 181 N*RSYQQREQNLQSWAGERATMVQ 252
>BE608412
Length = 441
Score = 35.4 bits (80), Expect = 0.048
Identities = 21/53 (39%), Positives = 32/53 (59%), Gaps = 3/53 (5%)
Frame = +2
Query: 198 RKGESSSLRLRKRFRQRRRRRVACKQNPRRVKKPRLKSLGRA*LLKQ---LHY 247
+ G SS+ +K+ QR++R + CKQ R+ +K LK+ GR LKQ LH+
Sbjct: 8 KDGNSST*SWKKKCMQRKQRLIKCKQCHRKRQKLILKNSGRVSTLKQHQCLHF 166
>TC222420
Length = 582
Score = 34.7 bits (78), Expect = 0.082
Identities = 30/126 (23%), Positives = 53/126 (41%), Gaps = 1/126 (0%)
Frame = +2
Query: 51 GNSENI-NQLESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSRAQKGPVK 109
GN+E N L T G + ++++G++D D S + + E +IK + K K
Sbjct: 23 GNTETEENPLNQTEGGKNQQEKVQGTSDKEDDSGVDNADSLE-QIKTKSNAEHMDKRQRK 199
Query: 110 NKNAKVGSSSGVNASLVKNSKIGKDKQASPAVSNGTSALDSRPRQPIKNRSSNDRQSQLS 169
N+K S+S +S++ ++ LDS+ P ++ + S
Sbjct: 200 KSNSKQTSTSKSISSMLTKDQV----------------LDSKKPSPSSGSGTHGKPSVAV 331
Query: 170 KKKPKS 175
KK KS
Sbjct: 332 TKKSKS 349
>BU547130
Length = 697
Score = 33.9 bits (76), Expect = 0.14
Identities = 26/53 (49%), Positives = 30/53 (56%), Gaps = 1/53 (1%)
Frame = -2
Query: 365 QPMKKTVPCQVTQMLLYLRMRFLQINQVK-NFM*MGT*WLKRILSSCCHKNRS 416
Q MK+ C V Q+LL F QI VK + M*M T* K ILSS +KN S
Sbjct: 669 QQMKRKXXCPVKQVLLCR*XXFXQIXXVKKSLM*MVT*QSKSILSSPWNKNLS 511
>BM885664 similar to GP|13539605|emb cyclophilin-RNA interacting protein
{Paramecium tetraurelia}, partial (5%)
Length = 420
Score = 32.7 bits (73), Expect = 0.31
Identities = 24/70 (34%), Positives = 35/70 (49%), Gaps = 4/70 (5%)
Frame = -2
Query: 165 QSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGESSSLRLRKRFRQRRRRRVACKQN 224
+S +SKKKP+S + P +G S KG+ S+L KR R+R R R ++
Sbjct: 419 KSAISKKKPRSSQSMPSSSSRG----SCKEREKEKGKESNLEREKREREREREREREREG 252
Query: 225 PR----RVKK 230
R R+KK
Sbjct: 251 GREQR*RLKK 222
>TC205596 similar to GB|AAO63377.1|28950907|BT005313 At4g34530 {Arabidopsis
thaliana;} , partial (28%)
Length = 1029
Score = 32.7 bits (73), Expect = 0.31
Identities = 21/85 (24%), Positives = 44/85 (51%), Gaps = 5/85 (5%)
Frame = +3
Query: 56 INQLESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQ--SRAQKGPVKNKNA 113
++ E+ A+G K+ + N V + + +K+K+ ++KV+ E+ S+ + +NKNA
Sbjct: 411 VSPKENMASGKENAKKRKPQNSKVVVAEIDNNKDKDKRVKVTGEEGESKVTEHHTRNKNA 590
Query: 114 KVGSSSG---VNASLVKNSKIGKDK 135
K ++ +A K S++ K
Sbjct: 591 KSNANKNNRETSADTSKGSEVQNQK 665
>TC225032 homologue to UP|Q9LLL9 (Q9LLL9) Plasma membrane aquaporin, partial
(60%)
Length = 729
Score = 32.7 bits (73), Expect = 0.31
Identities = 34/115 (29%), Positives = 53/115 (45%), Gaps = 2/115 (1%)
Frame = -3
Query: 20 VHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLE-STATGNSAMKEIEGSNDN 78
+H +S E SN +P V + V DG+SE + + S+ GN + + E S
Sbjct: 490 IHSSKGSSNER--SNPENPLVIPCMVSVVDDGSSETPSWVNASSCDGNGSQVDQEHSK-- 323
Query: 79 VDGSNLTVSKEKEVKIK-VSTEQSRAQKGPVKNKNAKVGSSSGVNASLVKNSKIG 132
SN S+ + V++ VS S + G KNK+A SS + + + KIG
Sbjct: 322 ---SNGKWSQNRNVRVSSVSLGISGRENGVDKNKSANNLSSKSITFGVSRIHKIG 167
>TC217191
Length = 710
Score = 32.0 bits (71), Expect = 0.53
Identities = 27/70 (38%), Positives = 40/70 (56%), Gaps = 1/70 (1%)
Frame = +1
Query: 346 QRKILAYPKVQGKRKLRLSQPMKKTVPCQVTQMLLYLRMRFLQINQVK-NFM*MGT*WLK 404
Q K+ YP + KRKL+ QP+K+T+ + Q+ L+L + +Q+K N +*M * K
Sbjct: 100 QLKLPPYPIQERKRKLK*LQPLKRTMSY*MRQVKLFL*I----*SQMKQNLL*MVI*SFK 267
Query: 405 RILSSCCHKN 414
R SS KN
Sbjct: 268 RNHSSTWRKN 297
>TC232400 weakly similar to UP|Q8W2K5 (Q8W2K5) Phragmoplastin-interacting
protein PHIP1, partial (8%)
Length = 621
Score = 32.0 bits (71), Expect = 0.53
Identities = 18/54 (33%), Positives = 31/54 (57%), Gaps = 2/54 (3%)
Frame = +3
Query: 206 RLRKRFRQRRRRRVACKQNPRRVKKPRLKSLGRA*LLKQ--LHYQLSIRSLLLL 257
R R+R + RRR R ++NP+R KK L G+ L+ QL ++++++L
Sbjct: 309 RTRRRSKNRRRTRTRTRRNPQRRKKTNLMVEGKCQRLQS*PKPIQLPVKTMVML 470
>AW755509 homologue to PIR|G96681|G966 protein F1E22.8 [imported] -
Arabidopsis thaliana, partial (4%)
Length = 431
Score = 31.6 bits (70), Expect = 0.70
Identities = 27/107 (25%), Positives = 47/107 (43%), Gaps = 6/107 (5%)
Frame = +2
Query: 110 NKNAKVGSSSGVNASLVK-NSKIGKDKQASPAVSNGTSA-----LDSRPRQPIKNRSSND 163
N + + G+++G+ VK N + SPA +NG A +QP +RS++
Sbjct: 74 NGSGESGATTGIKRIAVKRNVGAASPRSQSPARANGNGANGNKAFSENQQQPSLSRSNSR 253
Query: 164 RQSQLSKKKPKSLKKGPLDKVQGEGESSLYPFSLRKGESSSLRLRKR 210
+ Q K+ PL ++ E S +P S SS ++ R +
Sbjct: 254 KAEQ------SPYKRNPLSEI--EPNSLAFPHSTTNNSSSRVQNRPK 370
>TC225055 homologue to UP|Q9LLL9 (Q9LLL9) Plasma membrane aquaporin, complete
Length = 1148
Score = 31.2 bits (69), Expect = 0.91
Identities = 39/128 (30%), Positives = 57/128 (44%), Gaps = 2/128 (1%)
Frame = -1
Query: 7 LPADGLDDVHQNGVHDEPSNSGEDAVSNDLDPHVTVNTETFVPDGNSENINQLE-STATG 65
L DG DD +H +S E SN +P V + V DG+SE + + S+ G
Sbjct: 896 LERDGSDDYLV--IHSSKGSSNER--SNPENPLVIPCMVSVVDDGSSETPSWVNASSCDG 729
Query: 66 NSAMKEIEGSNDNVDGSNLTVSKEKEVKIK-VSTEQSRAQKGPVKNKNAKVGSSSGVNAS 124
N + + E S DG S+ + V + VS S + G KNK+A SS +
Sbjct: 728 NGSQVDQEHSKS--DGK---WSQNRNV*VSSVSLGISGRENGVDKNKSANNLSSKSITLG 564
Query: 125 LVKNSKIG 132
+ + KIG
Sbjct: 563 VSRIHKIG 540
>BM528840
Length = 406
Score = 30.8 bits (68), Expect = 1.2
Identities = 25/85 (29%), Positives = 41/85 (47%), Gaps = 1/85 (1%)
Frame = -1
Query: 302 RKCHRATPLQELPLPIKRSRYGSPFLLDSLLKGPIQQLHLHLRQQRKILAYPKVQGKRKL 361
R+C +T LP +G + LL+ +++LHL+ QQR+ L + QG R +
Sbjct: 352 RRCRSSTGTPALP-------FGLGA*IFFLLRTSLRKLHLNQIQQRRALPEAQTQGARLI 194
Query: 362 RLSQPMKKTVPCQVTQM-LLYLRMR 385
Q ++T + + LL LR R
Sbjct: 193 EYVQMGRRTDRTIIRSIPLLRLRQR 119
>TC211438 UP|GLCX_SOYBN (P11827) Beta-conglycinin, alpha' chain precursor,
complete
Length = 1959
Score = 30.0 bits (66), Expect = 2.0
Identities = 28/108 (25%), Positives = 44/108 (39%), Gaps = 4/108 (3%)
Frame = +2
Query: 43 NTETFVPDGNSENINQLESTATGNSAMKEIEGSNDNVDGSNLTVSKEKEVKIKVSTEQSR 102
NTE N N+N T K G+ V K+K+ K ++ ++ R
Sbjct: 428 NTEEREVKRNKMNVNTHAHTNLIKRKRKSTNGNTSRKSTKERKVKKKKKTKTRMKSKTKR 607
Query: 103 AQKGPV---KNKNAKVGSSSGVNASLVKNSKI-GKDKQASPAVSNGTS 146
A+K V K + K + S +K SK+ K A+ A S G++
Sbjct: 608 AKKVKVLSLKEEPTKT*E*GTLFTSTLKGSKLSSKTNMATFASSRGST 751
>TC228180 similar to UP|Q9SR11 (Q9SR11) F7O18.11 protein, partial (40%)
Length = 1690
Score = 30.0 bits (66), Expect = 2.0
Identities = 22/61 (36%), Positives = 32/61 (52%)
Frame = +3
Query: 216 RRRVACKQNPRRVKKPRLKSLGRA*LLKQLHYQLSIRSLLLLRWS*RRYQLLEQNLPSSV 275
+R+ + ++ R+ K+ S R LKQ+ YQ+S L LR +*R L QN S V
Sbjct: 960 KRKTSMRRGLRKNKRQPSSS*ERTW*LKQIQYQVSTMKRLPLRLN*RSCH*LGQNHQS*V 1139
Query: 276 G 276
G
Sbjct: 1140 G 1142
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.324 0.137 0.398
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 15,939,852
Number of Sequences: 63676
Number of extensions: 208200
Number of successful extensions: 2215
Number of sequences better than 10.0: 55
Number of HSP's better than 10.0 without gapping: 2122
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2180
length of query: 419
length of database: 12,639,632
effective HSP length: 100
effective length of query: 319
effective length of database: 6,272,032
effective search space: 2000778208
effective search space used: 2000778208
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 60 (27.7 bits)
Medicago: description of AC145331.12


