

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
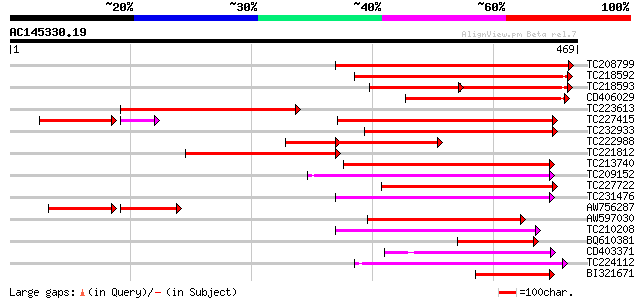
Query= AC145330.19 - phase: 0 /pseudo
(469 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC208799 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, pa... 333 7e-92
TC218592 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, pa... 295 4e-80
TC218593 158 5e-67
CD406029 217 8e-57
TC223613 weakly similar to UP|Q9FNC1 (Q9FNC1) Emb|CAB89401.1, pa... 214 9e-56
TC227415 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (89%) 211 7e-55
TC232933 similar to GB|AAN73299.1|25141209|BT002302 At3g26590/MF... 178 5e-45
TC222988 weakly similar to UP|Q9MAN5 (Q9MAN5) CDS, partial (29%) 135 1e-43
TC221812 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, pa... 167 9e-42
TC213740 weakly similar to UP|TT12_ARATH (Q9LYT3) TRANSPARENT TE... 165 5e-41
TC209152 weakly similar to PIR|B96764|B96764 protein integral me... 152 4e-37
TC227722 149 3e-36
TC231476 weakly similar to GB|AAP31960.1|30387589|BT006616 At1g1... 136 2e-32
AW756287 97 1e-31
AW597030 similar to SP|Q9LYT3|TT12 TRANSPARENT TESTA 12 protein.... 130 1e-30
TC210208 127 8e-30
BQ610381 106 3e-23
CD403371 97 1e-20
TC224112 88 9e-18
BI321671 83 2e-16
>TC208799 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, partial (46%)
Length = 924
Score = 333 bits (855), Expect = 7e-92
Identities = 163/197 (82%), Positives = 185/197 (93%)
Frame = +2
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+ +SICLNINGWEMMISLGFMAAASVRV+NELG+GSAKAAKFSI+V+VLTSLAIG LF+
Sbjct: 59 DALSICLNINGWEMMISLGFMAAASVRVANELGRGSAKAAKFSIIVSVLTSLAIGFLLFI 238
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
FFLFFRERLAYIFTSNKEVA AVG+LSPLLS+SILLNSVQPVLSGVAIGAGWQS VAYVN
Sbjct: 239 FFLFFRERLAYIFTSNKEVAFAVGDLSPLLSVSILLNSVQPVLSGVAIGAGWQSIVAYVN 418
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
+GCYY IGIPVGIVLGN++ QVKGIW+GMLFGTLIQTIVL++ITYKTNWDEQVT+A+KR
Sbjct: 419 MGCYYAIGIPVGIVLGNVLDLQVKGIWIGMLFGTLIQTIVLIVITYKTNWDEQVTIAQKR 598
Query: 450 VNRWSKVESTDQETKTK 466
++RWSKV+S D E + +
Sbjct: 599 ISRWSKVDSPDHENEVE 649
>TC218592 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, partial (37%)
Length = 715
Score = 295 bits (754), Expect = 4e-80
Identities = 150/180 (83%), Positives = 166/180 (91%)
Frame = +3
Query: 286 SLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFTSN 345
SLGFMAAASVRV+NELGKGS+KAAKFSIVVTVLTSLAIG LFLFFLF R +LAYIFTSN
Sbjct: 3 SLGFMAAASVRVANELGKGSSKAAKFSIVVTVLTSLAIGFVLFLFFLFLRGKLAYIFTSN 182
Query: 346 KEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLG 405
K+VA AVG+LSPLL+ISILLNSVQPVLSGVAIGAGWQS VAYVNIGCYYIIGIPVG+VLG
Sbjct: 183 KDVADAVGDLSPLLAISILLNSVQPVLSGVAIGAGWQSIVAYVNIGCYYIIGIPVGVVLG 362
Query: 406 NIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVNRWSKVESTDQETKT 465
N+++ QVKGIW+GMLFGT IQT+VL +ITYKT+WDEQVT AR R+N+WSKVES D ET T
Sbjct: 363 NVLNLQVKGIWIGMLFGTFIQTVVLTVITYKTDWDEQVTKARNRINKWSKVES-DHETIT 539
>TC218593
Length = 991
Score = 158 bits (400), Expect(2) = 5e-67
Identities = 75/92 (81%), Positives = 84/92 (90%)
Frame = +2
Query: 374 GVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLII 433
GVA+GAGWQS VAYVNIGCYY+IGIPVGIVLGNIIH QVKGIW+GMLFGTLIQTIVL II
Sbjct: 341 GVAVGAGWQSIVAYVNIGCYYLIGIPVGIVLGNIIHLQVKGIWIGMLFGTLIQTIVLTII 520
Query: 434 TYKTNWDEQVTVARKRVNRWSKVESTDQETKT 465
TYKTNWDEQV +AR R+++WSKV+ D+ET T
Sbjct: 521 TYKTNWDEQVIIARNRISKWSKVD-LDRETVT 613
Score = 114 bits (286), Expect(2) = 5e-67
Identities = 55/78 (70%), Positives = 71/78 (90%)
Frame = +3
Query: 298 SNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFTSNKEVAAAVGELSP 357
+NELG+GS+K AKFSIVVTVLTS +IG LF+ FLF RE++AY+FTSN++VA AVG+LSP
Sbjct: 3 ANELGRGSSKDAKFSIVVTVLTSFSIGFILFVLFLFLREKVAYLFTSNEDVATAVGDLSP 182
Query: 358 LLSISILLNSVQPVLSGV 375
LL++S+LLNS+QPVLSG+
Sbjct: 183 LLAVSLLLNSIQPVLSGM 236
>CD406029
Length = 464
Score = 217 bits (553), Expect = 8e-57
Identities = 102/136 (75%), Positives = 122/136 (89%)
Frame = -2
Query: 328 FLFFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAY 387
F+ FL RE++AY+ TSN++V AVG+LSPLL++S+LLNS+QPVLSGVA+G GWQSTVAY
Sbjct: 463 FVLFLILREKVAYLLTSNEDVVTAVGDLSPLLALSLLLNSIQPVLSGVALGPGWQSTVAY 284
Query: 388 VNIGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVAR 447
VNIGCYY+IGIPVGIVLGNIIH QVKGIW+GMLFGTLIQTI+L+IITYKTNWDEQV +AR
Sbjct: 283 VNIGCYYLIGIPVGIVLGNIIHLQVKGIWIGMLFGTLIQTIILIIITYKTNWDEQVIIAR 104
Query: 448 KRVNRWSKVESTDQET 463
R+N+WSK+ D ET
Sbjct: 103 DRINKWSKM-VLDHET 59
>TC223613 weakly similar to UP|Q9FNC1 (Q9FNC1) Emb|CAB89401.1, partial (15%)
Length = 460
Score = 214 bits (544), Expect = 9e-56
Identities = 99/149 (66%), Positives = 115/149 (76%)
Frame = +3
Query: 92 VGNGDCVVNPLWTSIWCKRIWNDGSISSKIMDSFIPNCTCSSSCVCLHNPDFDTLRPR*K 151
V NGDC+VN LWTSI KRI +DGS+SSKIMDSFI C SSS V LH P+FD+LRPR*+
Sbjct: 12 VRNGDCIVNTLWTSIRRKRI*HDGSVSSKIMDSFILKCNLSSSVVHLHKPNFDSLRPR*E 191
Query: 152 HIRSCRKHFSLVNSYYVCFYCLIHLPNIPSITKQEHHHCILGSFFHNHSCIPLLAFDNEI 211
H S + HF LVNS +C YCL LP+IPSI+KQE H+ I GSF +NHSC+PLLA N I
Sbjct: 192 HSTSGKNHFYLVNSCLICLYCLKQLPDIPSISKQECHYFIFGSFINNHSCVPLLAIHNAI 371
Query: 212 SIWDCWCNDFNYFGILDSQHWSTHICYLW 240
+WD WCNDFN FGILDS+HWST I Y+W
Sbjct: 372 QVWDSWCNDFNNFGILDSEHWSTDIYYMW 458
>TC227415 similar to UP|Q9LVD9 (Q9LVD9) Gb|AAC28507.1, partial (89%)
Length = 1748
Score = 211 bits (536), Expect = 7e-55
Identities = 101/182 (55%), Positives = 135/182 (73%)
Frame = +2
Query: 272 VSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFF 331
+SIC I+GW MIS+GF AAASVRVSNELG S K+A FS+VV + S I L
Sbjct: 1049 LSICTTISGWVFMISVGFNAAASVRVSNELGARSPKSASFSVVVVTVISFIISVIAALVV 1228
Query: 332 LFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIG 391
L R+ ++Y FT +EVAAAV +L PLL++S++LN +QPVLSGVA+G GWQ+ VAYVN+G
Sbjct: 1229 LALRDVISYAFTGGEEVAAAVSDLCPLLALSLVLNGIQPVLSGVAVGCGWQAFVAYVNVG 1408
Query: 392 CYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVN 451
CYY +GIP+G VLG + KGIW+GML GT++QTI+LL +T++T+W ++V A KR+
Sbjct: 1409 CYYGVGIPLGAVLGFYFQFGAKGIWLGMLGGTVMQTIILLWVTFRTDWTKEVEEAAKRLT 1588
Query: 452 RW 453
+W
Sbjct: 1589 KW 1594
Score = 47.0 bits (110), Expect(2) = 9e-06
Identities = 23/64 (35%), Positives = 39/64 (60%)
Frame = +1
Query: 25 LGKRVWNETKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFAN 84
+G W E KL++ +AAPA+ + + + +Q F GH+G+ ELAA +L T + FA
Sbjct: 238 VGPATWIELKLLFFLAAPAVIVYLINYLMSMSTQIFSGHLGNLELAAASLGNTGIQMFAY 417
Query: 85 GILM 88
G+++
Sbjct: 418 GLML 429
Score = 20.4 bits (41), Expect(2) = 9e-06
Identities = 10/33 (30%), Positives = 19/33 (57%)
Frame = +3
Query: 92 VGNGDCVVNPLWTSIWCKRIWNDGSISSKIMDS 124
VG G+C + + TSI ++ + S+ + + DS
Sbjct: 426 VGYGECRGDAMRTSIRSSKVRDVRSLHAAVNDS 524
>TC232933 similar to GB|AAN73299.1|25141209|BT002302 At3g26590/MFE16_11
{Arabidopsis thaliana;} , partial (30%)
Length = 545
Score = 178 bits (451), Expect = 5e-45
Identities = 81/160 (50%), Positives = 117/160 (72%)
Frame = +1
Query: 294 SVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFTSNKEVAAAVG 353
SVRVSNELG + AKFS++V ++TS IG L + + FR + F+++ EV A V
Sbjct: 55 SVRVSNELGASHPRTAKFSLLVAMITSTLIGVMLSMVLIIFRNHYPFFFSNDSEVRAMVV 234
Query: 354 ELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVK 413
EL+P+L++ I++N+VQPVLSGVA+GAGWQ+ VAYVNI CYY+ GIP+G+VLG + V
Sbjct: 235 ELTPMLALCIVINNVQPVLSGVAVGAGWQALVAYVNIACYYLFGIPLGLVLGYKLDMGVM 414
Query: 414 GIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVNRW 453
GIW GML GT++QT VL + Y+T+W+++ ++A R+ +W
Sbjct: 415 GIWSGMLSGTILQTCVLFFLVYRTDWNKEASLAEDRIKQW 534
>TC222988 weakly similar to UP|Q9MAN5 (Q9MAN5) CDS, partial (29%)
Length = 474
Score = 135 bits (339), Expect(2) = 1e-43
Identities = 66/89 (74%), Positives = 79/89 (88%)
Frame = +2
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+ +SIC+NINGWEMMI+ GFMAAASVRV+NELG+GS+KAAKFSIVVTVLTS IG LFL
Sbjct: 206 DALSICININGWEMMIAFGFMAAASVRVANELGRGSSKAAKFSIVVTVLTSFVIGFILFL 385
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPL 358
FLF RE++AY+ TSN++VA AVG+LSPL
Sbjct: 386 LFLFVREKVAYLSTSNEDVATAVGDLSPL 472
Score = 60.1 bits (144), Expect(2) = 1e-43
Identities = 28/45 (62%), Positives = 35/45 (77%)
Frame = +1
Query: 229 SQHWSTHICYLWLVS*NLEWFLIFSIQRPLACCQTFPFSRRNVVS 273
S++WST+I Y+WLV *N+E FL FSIQR LACCQ F F +V+S
Sbjct: 4 SEYWSTNIYYMWLVP*NMERFLCFSIQRSLACCQAFHFIWCHVMS 138
>TC221812 weakly similar to UP|Q9LP32 (Q9LP32) T9L6.1 protein, partial (37%)
Length = 470
Score = 167 bits (423), Expect = 9e-42
Identities = 79/128 (61%), Positives = 97/128 (75%)
Frame = +2
Query: 146 LRPR*KHIRSCRKHFSLVNSYYVCFYCLIHLPNIPSITKQEHHHCILGSFFHNHSCIPLL 205
LRPR*++ RS RK+FSLVNSY +CFYCL HLP P+I+KQ++HH LGS +++S +PLL
Sbjct: 5 LRPR*EYSRSGRKYFSLVNSYDICFYCLFHLPEFPAISKQKYHHFFLGSILNSYSFVPLL 184
Query: 206 AFDNEISIWDCWCNDFNYFGILDSQHWSTHICYLWLVS*NLEWFLIFSIQRPLACCQTFP 265
A DN I D WCNDFN FGILDS +WST+I +WLV +E FLIF IQR LACCQ FP
Sbjct: 185 AIDNSIQARDSWCNDFNKFGILDS*YWSTNIYNMWLVLRYMERFLIFGIQRSLACCQAFP 364
Query: 266 FSRRNVVS 273
F +V+S
Sbjct: 365 FIWYHVMS 388
>TC213740 weakly similar to UP|TT12_ARATH (Q9LYT3) TRANSPARENT TESTA 12
protein, partial (20%)
Length = 733
Score = 165 bits (417), Expect = 5e-41
Identities = 77/174 (44%), Positives = 113/174 (64%)
Frame = +3
Query: 277 NINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFLFFRE 336
N+ GW+ M+ LG A SVRVSN LG +AA +S VT+ SL +G F ++
Sbjct: 3 NVQGWDDMLRLGINTAISVRVSNTLGMSHPRAAIYSFCVTMFQSLLLGILFMTVIFFSKD 182
Query: 337 RLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYII 396
A IFT ++++ A +L+ LL ++I+LNS V+SGVAIG+GWQ V Y+N+ CYYI+
Sbjct: 183 EFAKIFTDSEDMILAAADLAYLLGVTIVLNSASQVMSGVAIGSGWQVMVGYINLACYYIV 362
Query: 397 GIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRV 450
G+P+GI LG +H VKG+W G + G+++QT+VL I +KTNW ++V R+
Sbjct: 363 GLPIGIFLGFKLHLGVKGLWGGTMCGSILQTLVLFTIIWKTNWSKEVEQTAHRM 524
>TC209152 weakly similar to PIR|B96764|B96764 protein integral membrane
protein F25P22.12 [imported] - Arabidopsis thaliana
{Arabidopsis thaliana;} , partial (41%)
Length = 1166
Score = 152 bits (383), Expect = 4e-37
Identities = 83/204 (40%), Positives = 123/204 (59%)
Frame = +1
Query: 247 EWFLIFSIQRPLACCQTFPFSRRNVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSA 306
EW+ F + LA P V+SICLN I A+AS RVSNELG G+
Sbjct: 217 EWWS-FEVLTLLAGILPNPQLETAVLSICLNTTTLHYFIPYAVGASASTRVSNELGAGNP 393
Query: 307 KAAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLN 366
K AK ++ V V+ +A + + F+ R L Y ++++KEV V E++PLL +S+ +
Sbjct: 394 KTAKGAVRVVVILGVAEAAIVSTVFISCRHVLGYAYSNDKEVIDYVAEMAPLLCVSVTAD 573
Query: 367 SVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQ 426
S+ LSG+A G G+Q AYVN+G YY++GIP+G++LG + + KG+WMG L G+L Q
Sbjct: 574 SLIGALSGIARGGGFQEIGAYVNLGAYYLVGIPMGLLLGFHLQLRAKGLWMGTLSGSLTQ 753
Query: 427 TIVLLIITYKTNWDEQVTVARKRV 450
I+L I+T +W ++ T AR+RV
Sbjct: 754 VIILAIVTALIDWQKEATKARERV 825
>TC227722
Length = 676
Score = 149 bits (375), Expect = 3e-36
Identities = 71/146 (48%), Positives = 103/146 (69%)
Frame = +1
Query: 308 AAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNS 367
AAK+S V V SL +G F L R+ A IFT+++ + AV +L LL+++++LNS
Sbjct: 4 AAKYSFYVIVFQSLFLGIFFMAIILATRDYYAIIFTNSEVLHKAVAKLGYLLAVTMVLNS 183
Query: 368 VQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQT 427
VQPV+SGVAIG GWQ+ VAY+NIGCYY+ G+P+G +LG + V+G+W GM+ G +IQT
Sbjct: 184 VQPVVSGVAIGGGWQALVAYINIGCYYLFGLPLGFLLGYEANLGVEGLWGGMICGIVIQT 363
Query: 428 IVLLIITYKTNWDEQVTVARKRVNRW 453
++LL+I YKTNW ++V +R+ W
Sbjct: 364 LLLLLILYKTNWKKEVEQTTERMRIW 441
>TC231476 weakly similar to GB|AAP31960.1|30387589|BT006616 At1g15170
{Arabidopsis thaliana;} , partial (32%)
Length = 734
Score = 136 bits (342), Expect = 2e-32
Identities = 71/181 (39%), Positives = 106/181 (58%)
Frame = +2
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
+V+SICLN I G AAAS R+SNELG G+ AA +++ + ++ + +
Sbjct: 2 SVLSICLNTISTLFSIPFGIAAAASTRISNELGAGNPHAAHVAVLAAMSFAIMETAIVSG 181
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
R YIF++ KEV V ++PL+ IS++L+S+Q VL+GVA G GWQ YVN
Sbjct: 182 TLFVCRHDFGYIFSNEKEVVDYVTVMAPLICISVILDSIQGVLAGVARGCGWQHIGVYVN 361
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKR 449
+G +Y+ GIPV L + + KG+W+G+ G +Q I+ IT NW++Q ARKR
Sbjct: 362 LGAFYLCGIPVAATLAFLAKMRGKGLWIGVQVGAFVQCILFSTITSCINWEQQAIKARKR 541
Query: 450 V 450
+
Sbjct: 542 L 544
>AW756287
Length = 319
Score = 97.4 bits (241), Expect(2) = 1e-31
Identities = 43/56 (76%), Positives = 55/56 (97%)
Frame = +2
Query: 33 TKLMWVVAAPAIFTRFSTFGIQIISQAFVGHIGSRELAAFALVFTVLIRFANGILM 88
+K+MW+VAAPAIFT+F+TFG+ +ISQAF+GHIGS+ELAA+ALVFTV+IRFANGIL+
Sbjct: 2 SKVMWIVAAPAIFTKFTTFGLSVISQAFIGHIGSKELAAYALVFTVIIRFANGILL 169
Score = 57.4 bits (137), Expect(2) = 1e-31
Identities = 31/51 (60%), Positives = 36/51 (69%)
Frame = +1
Query: 92 VGNGDCVVNPLWTSIWCKRIWNDGSISSKIMDSFIPNCTCSSSCVCLHNPD 142
V N CVVN L TSI KRI +DGS+SSKI+DS+I NCT SS LH P+
Sbjct: 166 VRNVKCVVNTLRTSIRSKRI*HDGSVSSKIIDSYILNCTLPSSVDHLHKPN 318
>AW597030 similar to SP|Q9LYT3|TT12 TRANSPARENT TESTA 12 protein. [Mouse-ear
cress] {Arabidopsis thaliana}, partial (25%)
Length = 401
Score = 130 bits (328), Expect = 1e-30
Identities = 66/130 (50%), Positives = 85/130 (64%)
Frame = +1
Query: 297 VSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFTSNKEVAAAVGELS 356
VSNELG + AKFS+ V TS+ I L FR L+ +FTS+ +V AV L+
Sbjct: 10 VSNELGASHPRVAKFSVFVVNGTSILISVVFCTIILIFRVSLSKLFTSDSDVIDAVSNLT 189
Query: 357 PLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKGIW 416
PLL+IS+ N +QP+LSGVAIG+GWQ+ VAYVN+ YY++G+ VG VLG V GIW
Sbjct: 190 PLLAISVFFNGIQPILSGVAIGSGWQALVAYVNLASYYVVGLTVGCVLGFKTSLGVAGIW 369
Query: 417 MGMLFGTLIQ 426
GM+ G LIQ
Sbjct: 370 WGMILGVLIQ 399
>TC210208
Length = 743
Score = 127 bits (320), Expect = 8e-30
Identities = 65/170 (38%), Positives = 101/170 (59%)
Frame = +2
Query: 270 NVVSICLNINGWEMMISLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFL 329
++++IC+N MI+ G AAAS RVSNELG G+ + AK ++ VT+ SL +G L
Sbjct: 185 SLIAICINTEFIAYMITYGLSAAASTRVSNELGAGNPERAKHAMSVTLKLSLLLGLCFVL 364
Query: 330 FFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVN 389
F F+ + + ++PLL+ISILL+++Q VLSGV+ G GWQ AY+N
Sbjct: 365 ALGFGHNIWIQFFSDSSTIKKEFASVTPLLAISILLDAIQGVLSGVSRGCGWQHLAAYIN 544
Query: 390 IGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNW 439
+ +Y+IG+P+ LG + Q KG+W+G++ G L Q+ L + + W
Sbjct: 545 LATFYLIGLPISCFLGFKTNLQYKGLWIGLICGLLCQSGTLFLFIRRKXW 694
>BQ610381
Length = 411
Score = 106 bits (264), Expect = 3e-23
Identities = 47/67 (70%), Positives = 57/67 (84%)
Frame = +1
Query: 371 VLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVL 430
VLSGVAIGAGWQ+ VAYVNI CYY+ GIPVG+VLG + W VKGIW+GM+ GT++QT VL
Sbjct: 211 VLSGVAIGAGWQALVAYVNIACYYLFGIPVGLVLGYKLDWGVKGIWLGMISGTILQTCVL 390
Query: 431 LIITYKT 437
L++ YKT
Sbjct: 391 LVLIYKT 411
>CD403371
Length = 592
Score = 97.4 bits (241), Expect = 1e-20
Identities = 48/142 (33%), Positives = 86/142 (59%), Gaps = 1/142 (0%)
Frame = -1
Query: 311 FSIVVTVLTSLAI-GSFLFLFFLFFRERLAYIFTSNKEVAAAVGELSPLLSISILLNSVQ 369
F+++V T + S LF F R L + F++ EV V ++ P+L +S +++
Sbjct: 589 FAVIVLAFTDAVVFSSVLFCF----RHVLGFAFSNEMEVVHYVAKIVPVLCLSFMVDGFL 422
Query: 370 PVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIV 429
VL G+ G+GWQ A N+ YY +GIPV ++ +++ KG+W+G+L G+ +QTI+
Sbjct: 421 GVLCGIVRGSGWQKIGAITNLVAYYAVGIPVSLLFXFGLNFNGKGLWIGILTGSTLQTII 242
Query: 430 LLIITYKTNWDEQVTVARKRVN 451
L ++T TNW++Q ++A +R++
Sbjct: 241 LALLTAFTNWEKQASLAIERLS 176
>TC224112
Length = 708
Score = 87.8 bits (216), Expect = 9e-18
Identities = 49/176 (27%), Positives = 83/176 (46%)
Frame = +1
Query: 286 SLGFMAAASVRVSNELGKGSAKAAKFSIVVTVLTSLAIGSFLFLFFLFFRERLAYIFTSN 345
SL F + S RV N+LG A+ S +V + S G +F L R A +FT +
Sbjct: 13 SLSF--SVSTRVGNKLGAQKPSKARLSAIVGLSCSFMSGVLALVFALMVRNTWASMFTKD 186
Query: 346 KEVAAAVGELSPLLSISILLNSVQPVLSGVAIGAGWQSTVAYVNIGCYYIIGIPVGIVLG 405
K++ + P++ + L N Q GV G A +N+GC+Y++G+PV I L
Sbjct: 187 KDIITLTSMVLPIIGLCELGNCPQTTGCGVLRGTARPKVGANINLGCFYLVGMPVSIWLA 366
Query: 406 NIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTVARKRVNRWSKVESTDQ 461
+ +G+W+G+L + +L++ +T+W+ + A+K DQ
Sbjct: 367 FFTGYDFQGLWLGLLAAQGSCAVTMLVVLCRTDWEFEAQRAKKLTGMGGAASGVDQ 534
>BI321671
Length = 419
Score = 83.2 bits (204), Expect = 2e-16
Identities = 33/65 (50%), Positives = 54/65 (82%)
Frame = +2
Query: 386 AYVNIGCYYIIGIPVGIVLGNIIHWQVKGIWMGMLFGTLIQTIVLLIITYKTNWDEQVTV 445
AYVN+G +Y++GIPVGI+LG + H++ KG+W+G++ G+++Q+I+L +IT TNW +Q +
Sbjct: 14 AYVNLGAFYLVGIPVGILLGFVAHFRAKGLWIGIVTGSIVQSILLSLITALTNWKKQAMM 193
Query: 446 ARKRV 450
AR+RV
Sbjct: 194 ARERV 208
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.329 0.141 0.463
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 28,187,752
Number of Sequences: 63676
Number of extensions: 572815
Number of successful extensions: 5033
Number of sequences better than 10.0: 80
Number of HSP's better than 10.0 without gapping: 4955
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 5023
length of query: 469
length of database: 12,639,632
effective HSP length: 101
effective length of query: 368
effective length of database: 6,208,356
effective search space: 2284675008
effective search space used: 2284675008
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.8 bits)
S2: 61 (28.1 bits)
Medicago: description of AC145330.19


