

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145329.13 - phase: 2 /pseudo
(528 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
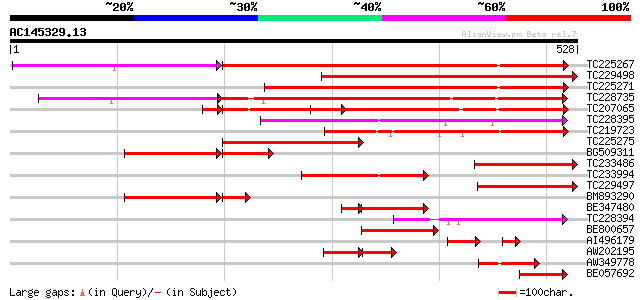
Score E
Sequences producing significant alignments: (bits) Value
TC225267 similar to UP|Q7XJH8 (Q7XJH8) Auxin-induced beta-glucos... 429 e-142
TC229498 similar to UP|Q94KD8 (Q94KD8) At1g02640/T14P4_11, parti... 408 e-114
TC225271 similar to UP|Q6RXY3 (Q6RXY3) Beta xylosidase (Fragment... 374 e-104
TC228735 similar to UP|Q76MS5 (Q76MS5) LEXYL1 protein, partial (... 313 e-103
TC207065 beta-glucosidase 243 1e-64
TC228395 weakly similar to UP|Q8VZG5 (Q8VZG5) At1g78060/F28K19_3... 216 2e-56
TC219723 weakly similar to UP|Q9LXA8 (Q9LXA8) Beta-xylosidase-li... 189 3e-48
TC225275 similar to GB|AAM00218.1|19879972|AF362990 beta-D-xylos... 187 1e-47
BG509311 similar to GP|19879972|gb beta-D-xylosidase {Prunus per... 110 1e-42
TC233486 similar to UP|Q94KD8 (Q94KD8) At1g02640/T14P4_11, parti... 160 1e-39
TC233994 similar to UP|Q8W011 (Q8W011) Beta-D-xylosidase, partia... 157 8e-39
TC229497 similar to UP|Q94KD8 (Q94KD8) At1g02640/T14P4_11, parti... 155 4e-38
BM893290 similar to GP|19879972|gb beta-D-xylosidase {Prunus per... 93 4e-27
BE347480 similar to GP|19879972|gb| beta-D-xylosidase {Prunus pe... 114 1e-25
TC228394 similar to UP|Q8VZG5 (Q8VZG5) At1g78060/F28K19_32, part... 86 3e-17
BE800657 weakly similar to GP|4100433|gb|A beta-glucosidase {Gly... 82 6e-16
AI496179 weakly similar to GP|14194121|gb| At1g02640/T14P4_11 {A... 59 2e-12
AW202195 similar to GP|19879972|gb| beta-D-xylosidase {Prunus pe... 67 2e-11
AW349778 65 6e-11
BE057692 similar to GP|10178060|dbj beta-xylosidase {Arabidopsis... 54 1e-07
>TC225267 similar to UP|Q7XJH8 (Q7XJH8) Auxin-induced beta-glucosidase, partial
(69%)
Length = 2006
Score = 429 bits (1104), Expect(2) = e-142
Identities = 208/324 (64%), Positives = 252/324 (77%), Gaps = 2/324 (0%)
Frame = +3
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTS 258
PLQGI RY KT HQ GC VACR ++ FG A AR ADA +LV+GLDQ++EAET DR
Sbjct: 756 PLQGIARYVKTAHQVGCKGVACRGNELFGAAETIARQADAIVLVMGLDQTVEAETRDRVG 935
Query: 259 LLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAI 318
LLLPG QQ+LV++VA A+KGP IL++MSGGPVDI+FAKNDPK++ ILW GYPGQAGG AI
Sbjct: 936 LLLPGLQQELVTRVARAAKGPVILLIMSGGPVDISFAKNDPKISAILWVGYPGQAGGTAI 1115
Query: 319 ADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPS-KIGYPGRTYRFYKGPVVYPFGH 377
AD++FGT +PGG+LP+TWYPQ YL + MTNM MRP+ GYPGRTYRFYKGPVV+PFGH
Sbjct: 1116 ADVIFGTTNPGGRLPMTWYPQGYLAKVPMTNMDMRPNPTTGYPGRTYRFYKGPVVFPFGH 1295
Query: 378 GLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARC-GKLSIALHVDVKNV 436
GL+Y+ F H L+ AP VSVP+ + N+ +S+KA++V+HA C L + HVDVKN
Sbjct: 1296 GLSYSRFSHSLALAPKQVSVPIMSLQALTNSTLSSKAVKVSHANCDDSLEMEFHVDVKNE 1475
Query: 437 GSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKS 496
GS DGTHTLL+FS PP+G W K LV F K HV A +KQRV+V +HVCK LSVVD+
Sbjct: 1476 GSMDGTHTLLIFSQPPHG--KWSQIKQLVGFHKTHVLAGSKQRVKVGVHVCKHLSVVDQF 1649
Query: 497 GIRRIPMGEHSLHIGDVKHSVSLQ 520
G+RRIP GEH LHIGDVKHS+S+Q
Sbjct: 1650 GVRRIPTGEHELHIGDVKHSISVQ 1721
Score = 94.4 bits (233), Expect(2) = e-142
Identities = 70/206 (33%), Positives = 92/206 (43%), Gaps = 11/206 (5%)
Frame = +2
Query: 3 GE*ARHRRHI*CTIQDVCEGRKSS*CHVFLQSSKWGPYMC*S*PLKENSSWSMGS*WVHC 62
GE* ++*CTI V GR S C V LQS +W ++C* ++ SW M WV+C
Sbjct: 113 GE*TGFGGYL*CTI*SVRVGRASGECDVLLQSGEWQAHVC*PRSPSQHYSWPMAPQWVYC 292
Query: 63 I*L*LCGSALQ*PTLYINAGRSCCRCHQSSAHAG-----------CGEKRFVN*SRCE*C 111
+ L* S L PTL+ N R C H S + G C +R C
Sbjct: 293 LRL*FSWSFL*QPTLHKNT*RGSC*GH*SGSRLGLWPIFGYPH*FCH*ERAYFRK*S*PC 472
Query: 112 FGEYIEGANEIRDV*WRAISSSLWKIRP*RCV*TSSSRASP*SC*TRNCASQKHWSYFAS 171
G+ G NEI V R ++ ++ K RP RCV S P*S * + K
Sbjct: 473 LGQPYFGPNEIGYVRRRTVNPTIRKPRPKRCVHLGPSTIGP*SS*REYSLTTKQRKLPTI 652
Query: 172 LPTTPPNCGCHWAQFRCHSYHDWKLC 197
+P C W Q C+ Y+DW+LC
Sbjct: 653 IPFKTSYYRCRWTQC*CYRYNDWELC 730
>TC229498 similar to UP|Q94KD8 (Q94KD8) At1g02640/T14P4_11, partial (24%)
Length = 792
Score = 408 bits (1049), Expect = e-114
Identities = 198/239 (82%), Positives = 214/239 (88%), Gaps = 1/239 (0%)
Frame = +1
Query: 291 DITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNM 350
DITFAKN+P++ GILWAGYPGQAGGAAIADILFGTA+PGGKLPVTWYP+EYL L MTNM
Sbjct: 1 DITFAKNNPRIVGILWAGYPGQAGGAAIADILFGTANPGGKLPVTWYPEEYLTKLPMTNM 180
Query: 351 AMRPSK-IGYPGRTYRFYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTN 409
AMR +K GYPGRTYRFY GPVVYPFGHGLTYTHFVH L+SAPTVVSVP++GHR N TN
Sbjct: 181 AMRATKSAGYPGRTYRFYNGPVVYPFGHGLTYTHFVHTLASAPTVVSVPLNGHRRANVTN 360
Query: 410 ISNKAIRVTHARCGKLSIALHVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEK 469
ISN+AIRVTHARC KLSI+L VD+KNVGSRDGTHTLLVFSAPP G HW +K LVAFEK
Sbjct: 361 ISNRAIRVTHARCDKLSISLEVDIKNVGSRDGTHTLLVFSAPPAGFGHWALEKQLVAFEK 540
Query: 470 VHVPAKTKQRVRVNIHVCKLLSVVDKSGIRRIPMGEHSLHIGDVKHSVSLQAEALGIIK 528
+HVPAK QRV VNIHVCKLLSVVDKSGIRRIP+GEHS +IGDVKHSVSLQA ALGIIK
Sbjct: 541 IHVPAKGLQRVGVNIHVCKLLSVVDKSGIRRIPLGEHSFNIGDVKHSVSLQAAALGIIK 717
>TC225271 similar to UP|Q6RXY3 (Q6RXY3) Beta xylosidase (Fragment), partial
(55%)
Length = 952
Score = 374 bits (959), Expect = e-104
Identities = 180/285 (63%), Positives = 223/285 (78%), Gaps = 2/285 (0%)
Frame = +3
Query: 238 ATILVIGLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKN 297
AT+LV+GLDQ+IEAET DR LLLPG QQ+LV++VA A+KGP ILV+MSGGPVD++FAKN
Sbjct: 3 ATVLVMGLDQTIEAETRDRVGLLLPGLQQELVTRVARAAKGPVILVIMSGGPVDVSFAKN 182
Query: 298 DPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPS-K 356
+PK++ ILW GYPGQAGG AIAD++FG +PGG+LP+TWYPQ YL + MTNM MRP+
Sbjct: 183 NPKISAILWVGYPGQAGGTAIADVIFGATNPGGRLPMTWYPQGYLAKVPMTNMDMRPNPA 362
Query: 357 IGYPGRTYRFYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIR 416
GYPGRTYRFYKGPVV+PFGHGL+Y+ F L+ AP VSV + + N+ +S+KA++
Sbjct: 363 TGYPGRTYRFYKGPVVFPFGHGLSYSRFSQSLALAPKQVSVQILSLQALTNSTLSSKAVK 542
Query: 417 VTHARC-GKLSIALHVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAK 475
V+HA C L HVDVKN GS DGTHTLL+FS PP G W K LV F K HVPA
Sbjct: 543 VSHANCDDSLETEFHVDVKNEGSMDGTHTLLIFSKPPPG--KWSQIKQLVTFHKTHVPAG 716
Query: 476 TKQRVRVNIHVCKLLSVVDKSGIRRIPMGEHSLHIGDVKHSVSLQ 520
+KQR++VN+H CK LSVVD+ G+RRIP GEH LHIGD+KHS+++Q
Sbjct: 717 SKQRLKVNVHSCKHLSVVDQFGVRRIPTGEHELHIGDLKHSINVQ 851
>TC228735 similar to UP|Q76MS5 (Q76MS5) LEXYL1 protein, partial (65%)
Length = 1599
Score = 313 bits (802), Expect(2) = e-103
Identities = 161/330 (48%), Positives = 219/330 (65%), Gaps = 5/330 (1%)
Frame = +1
Query: 195 KLCRPLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARH----ADATILVIGLDQSIE 250
K PLQG+ +A T + GC +V C + P LD A+ ADAT++V+G +IE
Sbjct: 565 KYISPLQGLTAFAPTSYAAGCLDVRCPN-----PVLDDAKKIAASADATVMVVGASLAIE 729
Query: 251 AETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYP 310
AE++DR ++LLPG QQ LVS+VA ASKGP ILV+MSGG +D++FAKN+ K+ ILW GYP
Sbjct: 730 AESLDRVNILLPGQQQLLVSEVANASKGPVILVIMSGGGMDVSFAKNNNKITSILWVGYP 909
Query: 311 GQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMRPS-KIGYPGRTYRFYKG 369
G+AGGAAIAD++FG +P G+LP+TWYPQ Y+ + MTNM MRP GYPGRTYRFYKG
Sbjct: 910 GEAGGAAIADVIFGFHNPSGRLPMTWYPQSYVDKVPMTNMNMRPDPATGYPGRTYRFYKG 1089
Query: 370 PVVYPFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNNTNISNKAIRVTHARCGKLSIAL 429
V+ FG GL+Y+ VH+L AP +VSV + ++ K+I V C L +
Sbjct: 1090ETVFAFGDGLSYSSIVHKLVKAPQLVSVQLAEDHVCRSSEC--KSIDVVGEHCQNLVFDI 1263
Query: 430 HVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKL 489
H+ +KN G HT+ +FS PP H PQK L+ FEKVH+ K++ V + VCK
Sbjct: 1264HLRIKNKGKMSSAHTVFLFSTPP--AVHNAPQKHLLGFEKVHLIGKSEALVSFKVDVCKD 1437
Query: 490 LSVVDKSGIRRIPMGEHSLHIGDVKHSVSL 519
LS+VD+ G R++ +G+H LH+GD+KH +S+
Sbjct: 1438LSIVDELGNRKVALGQHLLHVGDLKHPLSV 1527
Score = 82.4 bits (202), Expect(2) = e-103
Identities = 60/182 (32%), Positives = 83/182 (44%), Gaps = 11/182 (6%)
Frame = +3
Query: 28 CHVFLQSSKWGPYMC*S*PLKENSSWSMGS*WVHCI*L*LCGSALQ*PTLYINAGRSCCR 87
CHVFLQ W +C *P K W M + W++ *L*L SALQ L+ ++ RSC
Sbjct: 9 CHVFLQQG*WKANLCRP*PPKGCCPW*METKWIYSF*L*LGRSALQRSALHQDS*RSCSH 188
Query: 88 CHQSSA-----------HAGCGEKRFVN*SRCE*CFGEYIEGANEIRDV*WRAISSSLWK 136
H H C + R *S C + + + R +*W + ++LWK
Sbjct: 189 IHSGRVGFELWKISWPIHRRCCQARAYR*SIH*QCCHQQLRHVDASRIL*W*SKKATLWK 368
Query: 137 IRP*RCV*TSSSRASP*SC*TRNCASQKHWSYFASLPTTPPNCGCHWAQFRCHSYHDWKL 196
RC+ + P SC R+C ++K S AS + G +W Q C+ HDWKL
Sbjct: 369 PWSKRCLHSRKPGTCPRSCKARDCVAEK*SSITASQCQSH*IVGSYWTQC*CY*GHDWKL 548
Query: 197 CR 198
R
Sbjct: 549 *R 554
>TC207065 beta-glucosidase
Length = 1328
Score = 243 bits (621), Expect = 1e-64
Identities = 116/241 (48%), Positives = 166/241 (68%), Gaps = 1/241 (0%)
Frame = +2
Query: 281 ILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQE 340
ILV+MSGG +D++FAK++ K+ ILW GYPG+AGGAAIAD++FG +P G+LP+TWYPQ
Sbjct: 491 ILVIMSGGGMDVSFAKSNDKITSILWVGYPGEAGGAAIADVIFGFYNPSGRLPMTWYPQA 670
Query: 341 YLKNLAMTNMAMRPS-KIGYPGRTYRFYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPV 399
Y+ + MTNM MR GYPGRTYRFYKG V+ FG G++++ H++ AP +VSVP+
Sbjct: 671 YVNKVPMTNMNMRADPATGYPGRTYRFYKGETVFSFGDGISFSSIEHKIVKAPQLVSVPL 850
Query: 400 HGHRHGNNTNISNKAIRVTHARCGKLSIALHVDVKNVGSRDGTHTLLVFSAPPNGGNHWV 459
++ + I H C L+ +H+ VKN G +H +L+F PP+ H
Sbjct: 851 AEDHECRSSECMSLDIADEH--CQNLAFDIHLGVKNTGKMSTSHVVLLFFTPPD--VHNA 1018
Query: 460 PQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKSGIRRIPMGEHSLHIGDVKHSVSL 519
PQK L+ FEKVH+P K++ +VR + VCK LSVVD+ G R++P+G+H LH+G++KH +SL
Sbjct: 1019PQKHLLGFEKVHLPGKSEAQVRFKVDVCKDLSVVDELGNRKVPLGQHLLHVGNLKHPLSL 1198
Query: 520 Q 520
+
Sbjct: 1199R 1201
Score = 115 bits (289), Expect(2) = 5e-27
Identities = 57/115 (49%), Positives = 79/115 (68%)
Frame = +1
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTS 258
PLQ + T + GC NV C + + A A ADAT++++G +IEAE++DR +
Sbjct: 154 PLQTLTALVPTSYAAGCPNVQCAN-AELDDATQIAASADATVIIVGASLAIEAESLDRIN 330
Query: 259 LLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQA 313
+LLPG QQ LVS+VA ASKGP ILV+MSGG +D++FAK++ K+ ILW GYPG +
Sbjct: 331 ILLPGQQQLLVSEVANASKGPVILVIMSGGGMDVSFAKSNDKITSILWIGYPGDS 495
Score = 23.9 bits (50), Expect(2) = 5e-27
Identities = 10/19 (52%), Positives = 12/19 (62%)
Frame = +3
Query: 180 GCHWAQFRCHSYHDWKLCR 198
G + AQ C+ HDWKL R
Sbjct: 75 GSYRAQC*CYQGHDWKL*R 131
>TC228395 weakly similar to UP|Q8VZG5 (Q8VZG5) At1g78060/F28K19_32, partial
(22%)
Length = 1011
Score = 216 bits (551), Expect = 2e-56
Identities = 123/293 (41%), Positives = 174/293 (58%), Gaps = 7/293 (2%)
Frame = +3
Query: 234 RHADATILVIGLDQSIEAETVDRTSLLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDIT 293
R + +LV+GLDQS E E+ DR L LPG Q++L+ VA ASK P +LVL+ GGPVDIT
Sbjct: 3 RSSRPVVLVMGLDQSQERESHDREYLGLPGKQEELIKSVARASKRPVVLVLLCGGPVDIT 182
Query: 294 FAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMR 353
AK D KV GILWAGYPG+ GG A+A ++FG +PGGKLP+TWYP++++K + MT+M MR
Sbjct: 183 SAKFDDKVGGILWAGYPGELGGVALAQVVFGDHNPGGKLPITWYPKDFIK-VPMTDMRMR 359
Query: 354 PSKI-GYPGRTYRFYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPVHGHRH---GNNTN 409
GYPGRTYRFY GP VY FG+GL+YT + ++L S H N+
Sbjct: 360 ADPASGYPGRTYRFYTGPKVYEFGYGLSYTKYSYKLLSLSHSTLHINQSSTHLMTQNSET 539
Query: 410 ISNKAI-RVTHARCGKLSIALHVDVKNVGSRDGTHTLLVF--SAPPNGGNHWVPQKSLVA 466
I K + + C + +++ + V N G+ G H +L+F N+ P K LV
Sbjct: 540 IRYKLVSELAEETCQTMLLSIALGVTNRGNLAGKHPVLLFVRQGKVRNINNGNPVKQLVG 719
Query: 467 FEKVHVPAKTKQRVRVNIHVCKLLSVVDKSGIRRIPMGEHSLHIGDVKHSVSL 519
F+ V V A +V + C+ LSV +++G I G + +GD ++ + +
Sbjct: 720 FQSVKVNAGETVQVGFELSPCEHLSVANEAGSMVIEEGSYLFIVGDQEYPIEV 878
>TC219723 weakly similar to UP|Q9LXA8 (Q9LXA8) Beta-xylosidase-like protein
(AT5g10560/F12B17_90), partial (19%)
Length = 1013
Score = 189 bits (479), Expect = 3e-48
Identities = 96/238 (40%), Positives = 147/238 (61%), Gaps = 11/238 (4%)
Frame = +1
Query: 294 FAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGKLPVTWYPQEYLKNLAMTNMAMR 353
FA+ +P++A I+W GYPG+AGG A+A+I+FG +P G+LP+TWYP+ + N+ M M+MR
Sbjct: 1 FAEKNPQIASIIWLGYPGEAGGKALAEIIFGEFNPAGRLPMTWYPEAF-TNVPMNEMSMR 177
Query: 354 --PSKIGYPGRTYRFYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVP---VHGHRHGNNT 408
PS+ GYPGRTYRFY G VY FGHGL+++ F + SAP+ +S+ G R
Sbjct: 178 ADPSR-GYPGRTYRFYTGGRVYGFGHGLSFSDFSYNFLSAPSKISLSRTIKDGSRKRLLY 354
Query: 409 NISNKAIRVTHA------RCGKLSIALHVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQK 462
+ N+ V + C KLS ++H+ V N+G DG+H +++FS P + P+
Sbjct: 355 QVENEVYGVDYVPVNQLQNCNKLSFSVHISVMNLGGLDGSHVVMLFSKGPKVVD-GSPET 531
Query: 463 SLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDKSGIRRIPMGEHSLHIGDVKHSVSLQ 520
LV F ++H + + +H C+ LS DK G R +P+G H+L +GD++H VS++
Sbjct: 532 QLVGFSRLHTISSKPTETSILVHPCEHLSFADKQGKRILPLGPHTLSVGDLEHVVSIE 705
>TC225275 similar to GB|AAM00218.1|19879972|AF362990 beta-D-xylosidase
{Prunus persica;} , partial (32%)
Length = 933
Score = 187 bits (474), Expect = 1e-47
Identities = 88/131 (67%), Positives = 106/131 (80%)
Frame = +3
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGLDQSIEAETVDRTS 258
PLQGI RY KT HQ GC VACR ++ FG A AR DAT+LV+GLDQ+IEAET DR
Sbjct: 54 PLQGIARYVKTAHQVGCRGVACRGNELFGAAEIIARQVDATVLVMGLDQTIEAETRDRVG 233
Query: 259 LLLPGHQQDLVSKVAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAI 318
LLLPG QQ+LV++VA A+KGP ILV+MSGGPVD++FAKN+PK++ ILW GYPGQAGG AI
Sbjct: 234 LLLPGLQQELVTRVARAAKGPVILVIMSGGPVDVSFAKNNPKISAILWVGYPGQAGGTAI 413
Query: 319 ADILFGTASPG 329
AD++FG +PG
Sbjct: 414 ADVIFGATNPG 446
>BG509311 similar to GP|19879972|gb beta-D-xylosidase {Prunus persica},
partial (31%)
Length = 443
Score = 110 bits (276), Expect(2) = 1e-42
Identities = 51/90 (56%), Positives = 61/90 (67%)
Frame = +3
Query: 108 CE*CFGEYIEGANEIRDV*WRAISSSLWKIRP*RCV*TSSSRASP*SC*TRNCASQKHWS 167
C+ CFGEY +G NE+RDV*WRA S+W + P RCV T R *SC TRNC S +HWS
Sbjct: 6 CQWCFGEYTDGPNEVRDV*WRAHGPSIWSLGPKRCVQTGPPRTGS*SCQTRNCTS*EHWS 185
Query: 168 YFASLPTTPPNCGCHWAQFRCHSYHDWKLC 197
FA+L T + GCH AQF + Y+DWKLC
Sbjct: 186 CFATLLTASSHRGCHRAQF*SYYYNDWKLC 275
Score = 81.3 bits (199), Expect(2) = 1e-42
Identities = 35/47 (74%), Positives = 43/47 (91%)
Frame = +1
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDKQFGPALDAARHADATILVIGL 245
PLQGIGRYA+T+HQ GC NVAC++DK FGPA++AAR ADAT+LV+GL
Sbjct: 301 PLQGIGRYARTVHQLGCQNVACKNDKLFGPAINAARQADATVLVMGL 441
>TC233486 similar to UP|Q94KD8 (Q94KD8) At1g02640/T14P4_11, partial (7%)
Length = 677
Score = 160 bits (406), Expect = 1e-39
Identities = 75/95 (78%), Positives = 83/95 (86%)
Frame = +2
Query: 434 KNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVV 493
KNVGS+DGTHTLLVFSAPP G HW P K LVAF+K+H+P+K +QRV VNIHVCKLLSVV
Sbjct: 2 KNVGSKDGTHTLLVFSAPPAGNGHWAPHKQLVAFQKLHIPSKAQQRVNVNIHVCKLLSVV 181
Query: 494 DKSGIRRIPMGEHSLHIGDVKHSVSLQAEALGIIK 528
D+SG RR+PMG HSLHIGDVKH VSLQAE LGIIK
Sbjct: 182 DRSGTRRVPMGLHSLHIGDVKHYVSLQAETLGIIK 286
>TC233994 similar to UP|Q8W011 (Q8W011) Beta-D-xylosidase, partial (15%)
Length = 462
Score = 157 bits (398), Expect = 8e-39
Identities = 75/120 (62%), Positives = 94/120 (77%), Gaps = 1/120 (0%)
Frame = +2
Query: 272 VAAASKGPTILVLMSGGPVDITFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPGGK 331
VA ASK P ILVL+SGGP+DIT AK + K+ GILWAGYPG+ GG A+A I+FG +PGG+
Sbjct: 2 VAEASKKPVILVLLSGGPLDITSAKYNHKIGGILWAGYPGELGGIALAQIIFGDHNPGGR 181
Query: 332 LPVTWYPQEYLKNLAMTNMAMRPS-KIGYPGRTYRFYKGPVVYPFGHGLTYTHFVHELSS 390
LP TWYP++Y+K + MT+M MR GYPGRTYRFYKGP VY FG+GL+Y+ + +EL S
Sbjct: 182 LPTTWYPKDYIK-VPMTDMRMRADPSTGYPGRTYRFYKGPKVYEFGYGLSYSKYSYELVS 358
>TC229497 similar to UP|Q94KD8 (Q94KD8) At1g02640/T14P4_11, partial (12%)
Length = 651
Score = 155 bits (392), Expect = 4e-38
Identities = 77/93 (82%), Positives = 82/93 (87%)
Frame = +3
Query: 436 VGSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVVDK 495
VGSRDGTHTLLVFSAPP G HW +K LVAFEKVHVPAK + RV VNIHVCKLLSVVD+
Sbjct: 3 VGSRDGTHTLLVFSAPPAGFGHWALEKQLVAFEKVHVPAKGQHRVGVNIHVCKLLSVVDR 182
Query: 496 SGIRRIPMGEHSLHIGDVKHSVSLQAEALGIIK 528
SGIRRIP+GEHS +IGDVKHSVSLQA ALGIIK
Sbjct: 183 SGIRRIPLGEHSFNIGDVKHSVSLQAAALGIIK 281
>BM893290 similar to GP|19879972|gb beta-D-xylosidase {Prunus persica},
partial (27%)
Length = 403
Score = 92.8 bits (229), Expect(2) = 4e-27
Identities = 49/90 (54%), Positives = 60/90 (66%)
Frame = +3
Query: 108 CE*CFGEYIEGANEIRDV*WRAISSSLWKIRP*RCV*TSSSRASP*SC*TRNCASQKHWS 167
CE F E+I NE+RDV*W AI SS+ + RCV S RA P*S TR+CAS K S
Sbjct: 30 CEWGFIEHINSPNEVRDV*W*AIKSSI*QPWSKRCVHPKSPRACP*SRKTRDCAS*KQRS 209
Query: 168 YFASLPTTPPNCGCHWAQFRCHSYHDWKLC 197
+FASL P+C C+WAQF CH ++DW+LC
Sbjct: 210 FFASLH*AWPHCSCYWAQF*CHFHNDWQLC 299
Score = 47.4 bits (111), Expect(2) = 4e-27
Identities = 19/26 (73%), Positives = 22/26 (84%)
Frame = +1
Query: 199 PLQGIGRYAKTIHQQGCSNVACRDDK 224
PLQGIG Y KTI++ GC+NVAC DDK
Sbjct: 325 PLQGIGTYTKTIYEHGCANVACTDDK 402
>BE347480 similar to GP|19879972|gb| beta-D-xylosidase {Prunus persica},
partial (17%)
Length = 473
Score = 114 bits (285), Expect = 1e-25
Identities = 51/64 (79%), Positives = 57/64 (88%), Gaps = 1/64 (1%)
Frame = +3
Query: 328 PGGKLPVTWYPQEYLKNLAMTNMAMRPSKI-GYPGRTYRFYKGPVVYPFGHGLTYTHFVH 386
PGGKLP+TWYPQ Y+KNL MTNMAMR S+ GYPGRTYRFY GPVVYPFG+GL+YTHFVH
Sbjct: 282 PGGKLPMTWYPQGYIKNLPMTNMAMRASRSKGYPGRTYRFYNGPVVYPFGYGLSYTHFVH 461
Query: 387 ELSS 390
L+S
Sbjct: 462 TLTS 473
Score = 41.6 bits (96), Expect = 9e-04
Identities = 18/20 (90%), Positives = 20/20 (100%)
Frame = +2
Query: 310 PGQAGGAAIADILFGTASPG 329
PGQAGGAAIADILFGT++PG
Sbjct: 2 PGQAGGAAIADILFGTSNPG 61
>TC228394 similar to UP|Q8VZG5 (Q8VZG5) At1g78060/F28K19_32, partial (14%)
Length = 659
Score = 86.3 bits (212), Expect = 3e-17
Identities = 55/173 (31%), Positives = 87/173 (49%), Gaps = 11/173 (6%)
Frame = +2
Query: 358 GYPGRTYRFYKGPVVYPFGHGLTYTHFVHELSSAPTVVSVPVHGHRHGNN-----TNISN 412
GYPGRTYRFY GP VY FG+GL+YT + ++L S H H N T ++
Sbjct: 23 GYPGRTYRFYTGPKVYEFGYGLSYTKYSYKLLSLS-------HNTLHINQSSTHLTTQNS 181
Query: 413 KAIR------VTHARCGKLSIALHVDVKNVGSRDGTHTLLVFSAPPNGGNHWVPQKSLVA 466
+ IR + C + +++ + V N G+ G H +L+F N+ P K LV
Sbjct: 182 ETIRYKLVSELAEETCQTMLLSIALGVTNHGNMAGKHPVLLFVRQGKVRNNGNPVKQLVG 361
Query: 467 FEKVHVPAKTKQRVRVNIHVCKLLSVVDKSGIRRIPMGEHSLHIGDVKHSVSL 519
F+ V + A +V + C+ LSV +++G I G + L +GD ++ + +
Sbjct: 362 FQSVKLNAGETVQVGFELSPCEHLSVANEAGSMVIEEGSYLLLVGDQEYPIEI 520
>BE800657 weakly similar to GP|4100433|gb|A beta-glucosidase {Glycine max},
partial (19%)
Length = 220
Score = 82.0 bits (201), Expect = 6e-16
Identities = 37/73 (50%), Positives = 50/73 (67%), Gaps = 1/73 (1%)
Frame = +2
Query: 328 PGGKLPVTWYPQEYLKNLAMTNMAMRPS-KIGYPGRTYRFYKGPVVYPFGHGLTYTHFVH 386
PGG+LP+TWYPQ YL + +TN+ M P+ YP +TYRFYK PV++PF H L+Y+ F
Sbjct: 2 PGGRLPMTWYPQRYLAKVPITNIDMHPNPSTRYP*KTYRFYK*PVIFPFKHKLSYSRFSQ 181
Query: 387 ELSSAPTVVSVPV 399
LS P VS+ +
Sbjct: 182 ILSLTPKQVSIQI 220
>AI496179 weakly similar to GP|14194121|gb| At1g02640/T14P4_11 {Arabidopsis
thaliana}, partial (3%)
Length = 139
Score = 58.5 bits (140), Expect(2) = 2e-12
Identities = 27/31 (87%), Positives = 30/31 (96%)
Frame = +1
Query: 408 TNISNKAIRVTHARCGKLSIALHVDVKNVGS 438
+NI+NKAI+VTHARCGKLSI LHVDVKNVGS
Sbjct: 1 SNIANKAIKVTHARCGKLSINLHVDVKNVGS 93
Score = 32.0 bits (71), Expect(2) = 2e-12
Identities = 13/16 (81%), Positives = 14/16 (87%)
Frame = +2
Query: 460 PQKSLVAFEKVHVPAK 475
P K LVAFEKVH+PAK
Sbjct: 92 PHKQLVAFEKVHIPAK 139
>AW202195 similar to GP|19879972|gb| beta-D-xylosidase {Prunus persica},
partial (14%)
Length = 417
Score = 67.0 bits (162), Expect = 2e-11
Identities = 29/37 (78%), Positives = 34/37 (91%)
Frame = +1
Query: 293 TFAKNDPKVAGILWAGYPGQAGGAAIADILFGTASPG 329
T AKN+P++ ILWAGYPGQAGGAAIADILFGT++PG
Sbjct: 1 TSAKNNPRIQAILWAGYPGQAGGAAIADILFGTSNPG 111
Score = 51.6 bits (122), Expect = 9e-07
Identities = 24/33 (72%), Positives = 27/33 (81%), Gaps = 1/33 (3%)
Frame = +2
Query: 329 GGKLPVTWYPQEYLKNLAMTNMAMRPSKI-GYP 360
GGKLP+TWYPQ +KNL MTNMAMR S+ GYP
Sbjct: 317 GGKLPMTWYPQGSIKNLPMTNMAMRASRSKGYP 415
>AW349778
Length = 338
Score = 65.5 bits (158), Expect = 6e-11
Identities = 31/57 (54%), Positives = 40/57 (69%)
Frame = -2
Query: 437 GSRDGTHTLLVFSAPPNGGNHWVPQKSLVAFEKVHVPAKTKQRVRVNIHVCKLLSVV 493
GS DGTH+LL+F PP G W + LV+ K HVPA +K+R++VN+H CK LSVV
Sbjct: 334 GSMDGTHSLLIFCKPPQGA--W*QIQQLVSLAKQHVPAGSKERIKVNVHSCKQLSVV 170
>BE057692 similar to GP|10178060|dbj beta-xylosidase {Arabidopsis thaliana},
partial (5%)
Length = 225
Score = 54.3 bits (129), Expect = 1e-07
Identities = 21/45 (46%), Positives = 36/45 (79%)
Frame = +2
Query: 475 KTKQRVRVNIHVCKLLSVVDKSGIRRIPMGEHSLHIGDVKHSVSL 519
K++ +VR + +CK LSVVD+ G R++P+G+H LH+G++KH +S+
Sbjct: 8 KSEAQVRFKVDICKDLSVVDELGNRKVPLGQHLLHVGNLKHQLSV 142
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.337 0.146 0.508
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,984,326
Number of Sequences: 63676
Number of extensions: 480517
Number of successful extensions: 3662
Number of sequences better than 10.0: 58
Number of HSP's better than 10.0 without gapping: 3583
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 3631
length of query: 528
length of database: 12,639,632
effective HSP length: 102
effective length of query: 426
effective length of database: 6,144,680
effective search space: 2617633680
effective search space used: 2617633680
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 39 (21.7 bits)
S2: 61 (28.1 bits)
Medicago: description of AC145329.13


