

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
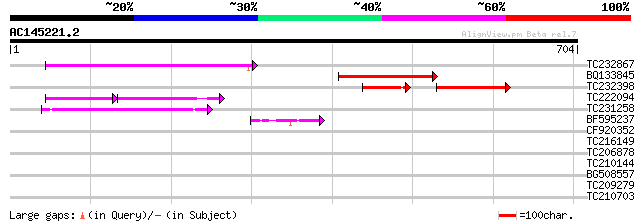
Query= AC145221.2 - phase: 0
(704 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial ... 176 2e-44
BQ133845 140 3e-33
TC232398 96 3e-32
TC222094 73 1e-25
TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kin... 106 4e-23
BF595237 48 1e-05
CF920352 33 0.33
TC216149 weakly similar to UP|MNN4_YEAST (P36044) MNN4 protein, ... 30 4.8
TC206878 similar to UP|Q6K8D7 (Q6K8D7) DNAJ heat shock N-termina... 30 4.8
TC210144 similar to UP|Q8VZI2 (Q8VZI2) AT4g33700/T16L1_190, part... 29 6.2
BG508557 weakly similar to GP|9294379|dbj glucan synthase-like p... 29 6.2
TC209279 weakly similar to UP|Q99738 (Q99738) Pinin, partial (3%) 29 6.2
TC210703 similar to UP|Q96397 (Q96397) LRG5 protein, partial (4%) 29 8.1
>TC232867 similar to UP|Q9MA68 (Q9MA68) F2J6.13 protein, partial (3%)
Length = 1144
Score = 176 bits (447), Expect = 2e-44
Identities = 96/266 (36%), Positives = 152/266 (57%), Gaps = 3/266 (1%)
Frame = -3
Query: 45 IQLVSNHPFAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLFQHSLIGKAREWYLD 104
IQL+ ++ F GL +EDPY HL+ + + N++ + +A L L SL G+A+ W
Sbjct: 806 IQLIQSNFFHGLPNEDPYAHLATYIEICNTIRLAGVPADAIRLSLLSFSLSGEAKRWLHS 627
Query: 105 QSPDVLKDWNALEKAFQERFFPDHKHMDTKTAIAMFAQGADESLCDAWERYKSLLRKCPK 164
+ LK W+ + + F +++FP+ K + K AI+ F Q DESL +A ER++ LLRK P
Sbjct: 626 FKGNSLKSWDEVVEKFLKKYFPESKTAEGKAAISSFHQFPDESLSEALERFRGLLRKTPT 447
Query: 165 HGFDDATQIHMFRGGLQPEPKLILDAAAHGSLMALSPREAVDIINKMSLNDRAAKHNRSG 224
HGF + Q+++F L+PE K ++DA+ G + +P EA+D+I M+ +D A +R+
Sbjct: 446 HGFSEPIQLNIFIDELRPESKQLVDASVGGKIKMKTPDEAMDLIESMAASDIAILRDRAH 267
Query: 225 TQRKSGILELGSNDANLAQTKLLTQQMEEMMKQMKEMPKLVKEQIQKEQRHQQVHFCEMC 284
K +LEL S D LAQ KLL++Q+E + K + ++P + QV C +
Sbjct: 266 IPTKKSLLELTSQDTLLAQNKLLSKQLETLTKTLSKLPTQLHSAQTSHSSILQVTGCTIF 87
Query: 285 SGDHPTGYCPP---QLEEEVNYVGNQ 307
H +G C P Q+ EVNY+GN+
Sbjct: 86 GEAHESGCCIPNEEQIAHEVNYMGNR 9
>BQ133845
Length = 389
Score = 140 bits (352), Expect = 3e-33
Identities = 67/123 (54%), Positives = 92/123 (74%)
Frame = +2
Query: 409 VPAHKLPYPKAQSKKDLEKHFKRFLDIFKRLEINIPFSEALEQMPTYAKFMKEILSKKHR 468
+ ++PYP S KD E+H +FLDIFK+LEI +PF EAL+QMP YA F+K++L+KK+
Sbjct: 20 IECKEVPYPLVPS*KDKEQHLAKFLDIFKKLEITLPFEEALQQMPLYANFLKDMLTKKNW 199
Query: 469 YKEDETIQLDANCSAIIQRRLPKKEKDPGRVTLPVAIGKVNVGKALIDLGSSINLIPYSV 528
Y + I ++ NCSA+IQR LP DPG VT+P +IG+V VGKALIDLG+SINL+P S+
Sbjct: 200 YIHSDKIVVEGNCSAVIQRILPP*HTDPGFVTMPCSIGEVAVGKALIDLGASINLMPLSM 379
Query: 529 VKR 531
++
Sbjct: 380 CRQ 388
>TC232398
Length = 1054
Score = 95.9 bits (237), Expect(2) = 3e-32
Identities = 44/92 (47%), Positives = 67/92 (72%)
Frame = +1
Query: 530 KRVGGLDMKLTRMTLQLADKSVARPMGIAEDVLVKVDKFVFPVDFVVMDIEEDDDVPLIL 589
+R+G ++ TRMTLQLAD S+ R G+ ED+LVKV + +F VDFV+MDIEED ++ LIL
Sbjct: 172 RRIGNQKIEPTRMTLQLADHSITRSFGVVEDILVKVHQLIFLVDFVIMDIEEDAEIRLIL 351
Query: 590 GRTFMLAARMMIDYDDGLMKVRLDDEEINFNL 621
G FM+ A+ ++D G +++ ++D++ FNL
Sbjct: 352 GWPFMVTAKCVVDMGKGNLEMSVEDQKATFNL 447
Score = 61.6 bits (148), Expect(2) = 3e-32
Identities = 32/59 (54%), Positives = 40/59 (67%)
Frame = +2
Query: 439 LEINIPFSEALEQMPTYAKFMKEILSKKHRYKEDETIQLDANCSAIIQRRLPKKEKDPG 497
LEI IPF E ++QMP Y KF+K+IL KK +Y ETI + C A+IQ +LP K KD G
Sbjct: 2 LEITIPFGERIQQMPLYKKFLKDILIKKGKYINSETIVVGEYCRALIQ-KLPPKFKDLG 175
>TC222094
Length = 984
Score = 73.2 bits (178), Expect(2) = 1e-25
Identities = 44/134 (32%), Positives = 67/134 (49%), Gaps = 1/134 (0%)
Frame = -3
Query: 134 KTAIAMFAQGADESLCDAWERYKSLLRKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAH 193
K I+ F Q ESL +A +R+ LL K P HGF + Q+++F G+QP K +LDA+A
Sbjct: 478 KVEISSFHQHPHESLSEALDRFHGLLWKTPTHGFSEPVQLNIFIDGMQPHSKQLLDASAG 299
Query: 194 GSLMALSPREAVDIINKMSLNDRA-AKHNRSGTQRKSGILELGSNDANLAQTKLLTQQME 252
G + +P EA+++I M+ ND + Q +S LE +LL Q M+
Sbjct: 298 GKIKLKTPEEAIELIENMAANDYVILRDQEPSPQEESTRLE-----------ELLVQFMQ 152
Query: 253 EMMKQMKEMPKLVK 266
E K ++
Sbjct: 151 ETRSHQKSTDAAIR 110
Score = 62.0 bits (149), Expect(2) = 1e-25
Identities = 32/90 (35%), Positives = 49/90 (53%)
Frame = -2
Query: 45 IQLVSNHPFAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLFQHSLIGKAREWYLD 104
IQL+ + F GL EDPY HL+ + + N++ E+A L LF SL G+AR W
Sbjct: 746 IQLIQGNLFYGLPTEDPYAHLATYIDICNTVKIVGVPEDAIHLDLFCFSLAGEARTWLRS 567
Query: 105 QSPDVLKDWNALEKAFQERFFPDHKHMDTK 134
+ L+ WN + F +++FP+ K + K
Sbjct: 566 FKGNNLRTWNEXXEKFLKKYFPESKTIXXK 477
>TC231258 similar to UP|Q886T6 (Q886T6) Sensory box histidine kinase/response
regulator, partial (3%)
Length = 1374
Score = 106 bits (264), Expect = 4e-23
Identities = 62/212 (29%), Positives = 112/212 (52%)
Frame = +2
Query: 40 LKSGTIQLVSNHPFAGLDHEDPYQHLSVFYALANSLGASEDEEEAGFLRLFQHSLIGKAR 99
LK+G I L+ F GL EDP++HL F+ + +++ + +E+ FL+ F HSL G A+
Sbjct: 521 LKTGLIHLLPK--FHGLVGEDPHKHLKEFHIVCSTMKPPDVQEDHIFLKAFPHSLEGVAK 694
Query: 100 EWYLDQSPDVLKDWNALEKAFQERFFPDHKHMDTKTAIAMFAQGADESLCDAWERYKSLL 159
+W +P + +W+ L++ F E+FFP + + I+ Q + ESL + WER+K
Sbjct: 695 DWLYYLAPRSIFNWDDLKRVFLEKFFPASRTTTIRKDISGIRQLSRESLYEYWERFKKSC 874
Query: 160 RKCPKHGFDDATQIHMFRGGLQPEPKLILDAAAHGSLMALSPREAVDIINKMSLNDRAAK 219
CP H + F L + ++DAA+ G+L ++P EA ++I KM+ N +
Sbjct: 875 ASCPHHQISKQLLL*YFYEELSNMKRSMIDAASGGALGDMTPTEARNLIKKMASNSQQFS 1054
Query: 220 HNRSGTQRKSGILELGSNDANLAQTKLLTQQM 251
+ G+ E+ ++ ++ + K L + +
Sbjct: 1055ARNDAIVLR-GVHEVATDSSSSTENKSLRENL 1147
>BF595237
Length = 415
Score = 48.1 bits (113), Expect = 1e-05
Identities = 34/98 (34%), Positives = 47/98 (47%), Gaps = 7/98 (7%)
Frame = +3
Query: 300 EVNYVGNQQRQGQFQGQYSGNANQQRFPNNNNFQRGNNFRQ--PWKSNAG-----PSNQQ 352
EVNY+G Q G FQG G F +G NF Q WK++ G Q
Sbjct: 33 EVNYMGAQNHHG-FQGYNQGGPP--------GFNQGRNFTQG*SWKNHPGNQFNKEQRSQ 185
Query: 353 PFNNNFQQYPPQQDRTQKIEDTLNQFMQLTMANQQNNR 390
P N+ Q ++T K+E+TLNQFM + M+N + +
Sbjct: 186 PIQNS-NQVVNLYEKTSKLEETLNQFM*MFMSNYMSTK 296
>CF920352
Length = 571
Score = 33.5 bits (75), Expect = 0.33
Identities = 25/72 (34%), Positives = 37/72 (50%)
Frame = -2
Query: 317 YSGNANQQRFPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYPPQQDRTQKIEDTLN 376
Y NQQ+ N+ GN F + G S+ +P Y DRT K+E+TL
Sbjct: 513 YGQQYNQQQEQ*RNH--PGNQFN----GDQGGSSIRPQQQGSSLY----DRTTKLEETLA 364
Query: 377 QFMQLTMANQQN 388
QFM ++M+NQ++
Sbjct: 363 QFM*VSMSNQKS 328
>TC216149 weakly similar to UP|MNN4_YEAST (P36044) MNN4 protein, partial (4%)
Length = 2040
Score = 29.6 bits (65), Expect = 4.8
Identities = 23/44 (52%), Positives = 28/44 (63%), Gaps = 2/44 (4%)
Frame = +3
Query: 662 TDALSVLSAEEEKEIEECLKELETLKEIPPKK--AKVEELKKEE 703
TD LSV A+E ++ EE KE + KE P K+ AK EE KKEE
Sbjct: 1203 TDILSVGPAKEPEKKEEPKKEEK--KEEPKKEGEAKKEEEKKEE 1328
>TC206878 similar to UP|Q6K8D7 (Q6K8D7) DNAJ heat shock N-terminal
domain-containing protein-like, partial (28%)
Length = 426
Score = 29.6 bits (65), Expect = 4.8
Identities = 20/80 (25%), Positives = 34/80 (42%)
Frame = +3
Query: 363 PQQDRTQKIEDTLNQFMQLTMANQQNNRIHKESDVEKDKKKEVGKNVPAHKLPYPKAQSK 422
P+ T+K LN + + ++ + + E D +D +P H LP P + K
Sbjct: 6 PRHRETKKRSTLLNSISFVPLTHKLKHNMMWEWDDYQD-------TIPKHDLPQPDSHLK 164
Query: 423 KDLEKHFKRFLDIFKRLEIN 442
D R D +K LE++
Sbjct: 165 FDFFSALSRPKDYYKILEVD 224
>TC210144 similar to UP|Q8VZI2 (Q8VZI2) AT4g33700/T16L1_190, partial (54%)
Length = 1167
Score = 29.3 bits (64), Expect = 6.2
Identities = 15/57 (26%), Positives = 26/57 (45%), Gaps = 2/57 (3%)
Frame = +2
Query: 326 FPNNNNFQRGNNFRQPWKSNAGPSNQQPFNNNFQQYPPQQDRTQKI--EDTLNQFMQ 380
FPN+NN RG + + W N + N+ P +++ I ED + + +Q
Sbjct: 602 FPNSNNSNRGGSRSRKWSKNMYSDILEIDGNSLPSLPEKEEAVGIITMEDVIEELLQ 772
>BG508557 weakly similar to GP|9294379|dbj glucan synthase-like protein
{Arabidopsis thaliana}, partial (4%)
Length = 437
Score = 29.3 bits (64), Expect = 6.2
Identities = 16/43 (37%), Positives = 23/43 (53%), Gaps = 2/43 (4%)
Frame = -1
Query: 252 EEMMKQMKEMPKLVKEQIQKEQRHQQVHFC--EMCSGDHPTGY 292
EE M + P+ +Q Q ++ H+Q +FC E C GD P Y
Sbjct: 437 EEAMANLPS-PRGKLQQSQSKKGHKQENFCYAERCLGDRPNXY 312
>TC209279 weakly similar to UP|Q99738 (Q99738) Pinin, partial (3%)
Length = 1011
Score = 29.3 bits (64), Expect = 6.2
Identities = 22/51 (43%), Positives = 29/51 (56%)
Frame = +2
Query: 646 ETRKQLHKPTPLERVLTDALSVLSAEEEKEIEECLKELETLKEIPPKKAKV 696
ET QL + ERV A SV +AEEE I E + + E ++EI + AKV
Sbjct: 749 ETAAQLPEAARKERVKQPAQSVGAAEEE-AIAEGVDDEEEMEEIRARLAKV 898
>TC210703 similar to UP|Q96397 (Q96397) LRG5 protein, partial (4%)
Length = 491
Score = 28.9 bits (63), Expect = 8.1
Identities = 17/49 (34%), Positives = 27/49 (54%), Gaps = 4/49 (8%)
Frame = -2
Query: 655 TPLERVLTDALSVLSAEEEKEIEECLKELE----TLKEIPPKKAKVEEL 699
TPL + ++ AEEE +EE L E LK++PP + ++EE+
Sbjct: 220 TPLMNHIATPSNLPQAEEEAWVEETLDSSEFGDRRLKKLPPPQLELEEV 74
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.133 0.384
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 27,725,978
Number of Sequences: 63676
Number of extensions: 361997
Number of successful extensions: 1890
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 1850
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1881
length of query: 704
length of database: 12,639,632
effective HSP length: 104
effective length of query: 600
effective length of database: 6,017,328
effective search space: 3610396800
effective search space used: 3610396800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 62 (28.5 bits)
Medicago: description of AC145221.2


