

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145219.5 + phase: 0
(130 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
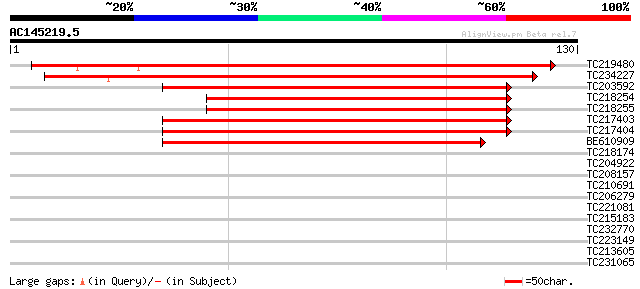
Sequences producing significant alignments: (bits) Value
TC219480 similar to UP|Q6UEI6 (Q6UEI6) Early flowering 4, partia... 161 9e-41
TC234227 weakly similar to UP|Q6UEI6 (Q6UEI6) Early flowering 4,... 105 8e-24
TC203592 87 2e-18
TC218254 84 2e-17
TC218255 82 5e-17
TC217403 79 5e-16
TC217404 79 5e-16
BE610909 77 3e-15
TC218174 similar to UP|Q41102 (Q41102) Phaseolin G-box binding p... 28 0.93
TC204922 similar to GB|CAC04249.1|9886727|PSA297606 PPF-1 protei... 27 2.7
TC208157 similar to UP|Q9FNV6 (Q9FNV6) Hypoxanthine-guanine phos... 27 2.7
TC210691 27 3.5
TC206279 similar to UP|Q8S976 (Q8S976) Auxin response factor 10 ... 27 3.5
TC221081 homologue to UP|Q9LR10 (Q9LR10) F10A5.10, partial (29%) 26 4.6
TC215183 26 4.6
TC232770 homologue to UP|Q6QUQ3 (Q6QUQ3) Auxin and ethylene resp... 26 6.0
TC223149 similar to UP|Q9LFT7 (Q9LFT7) Protein serine/threonine ... 26 6.0
TC213605 weakly similar to UP|RN12_HUMAN (Q9NVW2) RING finger pr... 26 6.0
TC231065 25 7.9
>TC219480 similar to UP|Q6UEI6 (Q6UEI6) Early flowering 4, partial (42%)
Length = 610
Score = 161 bits (407), Expect = 9e-41
Identities = 86/130 (66%), Positives = 98/130 (75%), Gaps = 10/130 (7%)
Frame = -3
Query: 6 SPSSAMEDT--NHRSKKNQRRRRPS--------ATPTNNDYSDVEDAEDGDPEAWQTLNK 55
S SSA E T + RSKK+ RR R +T T +VE+AEDGDPEAW TLNK
Sbjct: 539 SSSSAAESTMEDRRSKKHHRRHRHHPATTTTTLSTTTGESDGEVEEAEDGDPEAWATLNK 360
Query: 56 SFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVASLYSDLNSDFT 115
FRQVQSVLDRNR +IQQVNENQQSRM DNMVKNV LIQELNGNISKV SLYSDLN++FT
Sbjct: 359 GFRQVQSVLDRNRLLIQQVNENQQSRMHDNMVKNVSLIQELNGNISKVVSLYSDLNTNFT 180
Query: 116 NICHQQQRSS 125
N+C Q+ ++S
Sbjct: 179 NVCQQRSKNS 150
>TC234227 weakly similar to UP|Q6UEI6 (Q6UEI6) Early flowering 4, partial
(42%)
Length = 424
Score = 105 bits (261), Expect = 8e-24
Identities = 53/115 (46%), Positives = 77/115 (66%), Gaps = 2/115 (1%)
Frame = +2
Query: 9 SAMEDTNHRSKKN--QRRRRPSATPTNNDYSDVEDAEDGDPEAWQTLNKSFRQVQSVLDR 66
+AM++T +K + R R + T + +D E+ D EAW L +SF Q Q+VLD
Sbjct: 44 AAMDETAVTTKPDWLNNRARVAGERTEEFFCGGDDEEECDDEAWDMLTRSFWQAQTVLDE 223
Query: 67 NRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVASLYSDLNSDFTNICHQQ 121
NRA+I +VN N +S++PDNM KNVGLI +++GNISKV S+YSD++ +F NI Q+
Sbjct: 224 NRALIDEVNSNHESKIPDNMAKNVGLITQIHGNISKVRSIYSDVSVNFPNIIRQR 388
>TC203592
Length = 729
Score = 87.0 bits (214), Expect = 2e-18
Identities = 39/80 (48%), Positives = 59/80 (73%)
Frame = +3
Query: 36 YSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQE 95
+S + + D + QT KSF QVQ++LD+NR +I ++N+N +S++PDN+ +NVGLI+E
Sbjct: 201 FSGLGNGTQIDSKILQTFKKSFVQVQNILDQNRVLINEINQNHESKVPDNLSRNVGLIRE 380
Query: 96 LNGNISKVASLYSDLNSDFT 115
LN NI +V LY+DL+S FT
Sbjct: 381 LNNNIRRVVDLYADLSSSFT 440
>TC218254
Length = 887
Score = 83.6 bits (205), Expect = 2e-17
Identities = 36/70 (51%), Positives = 53/70 (75%)
Frame = +1
Query: 46 DPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVAS 105
D + QT K+F QVQ + D+NR +I ++N+N +S++PDN+ +NVGLI+ELN NI +V
Sbjct: 271 DAKIMQTFKKNFVQVQDIFDQNRLLINEINQNHESKVPDNLTRNVGLIRELNNNIRRVYD 450
Query: 106 LYSDLNSDFT 115
LY+DL+S FT
Sbjct: 451 LYADLSSSFT 480
>TC218255
Length = 737
Score = 82.4 bits (202), Expect = 5e-17
Identities = 36/70 (51%), Positives = 52/70 (73%)
Frame = +1
Query: 46 DPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVAS 105
D + QT K+F QVQ + D+NR +I ++N+N +S++PDN+ +NVGLI+ELN NI +V
Sbjct: 274 DAKIMQTFKKNFVQVQDIFDQNRLLINEINQNHESKVPDNLTRNVGLIRELNNNIRRVYD 453
Query: 106 LYSDLNSDFT 115
LY DL+S FT
Sbjct: 454 LYVDLSSSFT 483
>TC217403
Length = 943
Score = 79.3 bits (194), Expect = 5e-16
Identities = 34/80 (42%), Positives = 55/80 (68%)
Frame = +2
Query: 36 YSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQE 95
+ ++ ++ D Q KS Q Q +L++NR +I ++N+N +S+MPDN+ +NVGLI+E
Sbjct: 137 FGELGNSTQVDSRLLQVFQKSLLQAQDILNQNRLLINEINQNHESKMPDNLSRNVGLIRE 316
Query: 96 LNGNISKVASLYSDLNSDFT 115
LN NI +V LY+DL++ FT
Sbjct: 317 LNSNIRRVVDLYADLSNSFT 376
>TC217404
Length = 618
Score = 79.3 bits (194), Expect = 5e-16
Identities = 34/80 (42%), Positives = 55/80 (68%)
Frame = +2
Query: 36 YSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQE 95
+ ++ ++ D Q KS Q Q +L++NR +I ++N+N +S+MPDN+ +NVGLI+E
Sbjct: 371 FGELGNSTQVDSRLLQVFQKSLLQAQDILNQNRLLINEINQNHESKMPDNLSRNVGLIRE 550
Query: 96 LNGNISKVASLYSDLNSDFT 115
LN NI +V LY+DL++ FT
Sbjct: 551 LNSNIRRVVDLYADLSNSFT 610
>BE610909
Length = 407
Score = 76.6 bits (187), Expect = 3e-15
Identities = 34/74 (45%), Positives = 53/74 (70%)
Frame = +1
Query: 36 YSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQE 95
+S + + D + T KSF QVQ++LD+NR +I ++N+N +S++PDN+ +NVGLI+E
Sbjct: 184 FSGLGNGTQIDSKILLTFKKSFVQVQNILDQNRVLINEINQNHESKVPDNLSRNVGLIRE 363
Query: 96 LNGNISKVASLYSD 109
LN NI +V LY+D
Sbjct: 364 LNNNIRRVVDLYAD 405
>TC218174 similar to UP|Q41102 (Q41102) Phaseolin G-box binding protein PG2
(Fragment), partial (49%)
Length = 1345
Score = 28.5 bits (62), Expect = 0.93
Identities = 15/48 (31%), Positives = 28/48 (58%)
Frame = -1
Query: 22 QRRRRPSATPTNNDYSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRA 69
+RRRR S + + + + +E+GD + Q+L ++ Q S+L R R+
Sbjct: 349 RRRRRFSCSRRS*ESRRIPASENGDSDCSQSLGRNHTQKSSILWRRRS 206
>TC204922 similar to GB|CAC04249.1|9886727|PSA297606 PPF-1 protein {Pisum
sativum;} , partial (43%)
Length = 993
Score = 26.9 bits (58), Expect = 2.7
Identities = 11/32 (34%), Positives = 19/32 (59%)
Frame = +2
Query: 94 QELNGNISKVASLYSDLNSDFTNICHQQQRSS 125
+E+NG + K+ + L S TNIC+ + S+
Sbjct: 71 EEINGKLRKLILYFKSLYSIITNICYAGRTST 166
>TC208157 similar to UP|Q9FNV6 (Q9FNV6) Hypoxanthine-guanine
phosphoribosyltransferase 1 (Fragment), partial (77%)
Length = 863
Score = 26.9 bits (58), Expect = 2.7
Identities = 19/69 (27%), Positives = 34/69 (48%), Gaps = 8/69 (11%)
Frame = -3
Query: 8 SSAMEDTNHRSKKNQRRRRPSATPTNN-------DYSDVEDAEDGD-PEAWQTLNKSFRQ 59
S+AME + R++ ++R P ATPT S++ A E W +L++
Sbjct: 207 STAMERSILRTRSAKKRNAPVATPTTTGAGEWPPKSSEISAASSKTRREIWSSLHRILSI 28
Query: 60 VQSVLDRNR 68
+ +++RNR
Sbjct: 27 WECMVNRNR 1
>TC210691
Length = 811
Score = 26.6 bits (57), Expect = 3.5
Identities = 26/111 (23%), Positives = 48/111 (42%)
Frame = +1
Query: 18 SKKNQRRRRPSATPTNNDYSDVEDAEDGDPEAWQTLNKSFRQVQSVLDRNRAIIQQVNEN 77
+K + R + P + + SD+ D G + S Q+ S L +N+ +Q N
Sbjct: 247 NKFSSRHQYPHGSSIHPHDSDLSDNMYGQQQI------SRFQILSSLQKNQRFMQIEN-- 402
Query: 78 QQSRMPDNMVKNVGLIQELNGNISKVASLYSDLNSDFTNICHQQQRSSHNR 128
+P + + E +SKVA+ + S+ ++ +Q R+ HNR
Sbjct: 403 ----LPPLKLAPPAVYHE---KVSKVANFLTGETSNSSSAIEKQNRNKHNR 534
>TC206279 similar to UP|Q8S976 (Q8S976) Auxin response factor 10 (Fragment),
partial (12%)
Length = 835
Score = 26.6 bits (57), Expect = 3.5
Identities = 16/62 (25%), Positives = 31/62 (49%), Gaps = 12/62 (19%)
Frame = -3
Query: 8 SSAMEDTNHRSKKNQRRRRPSAT------------PTNNDYSDVEDAEDGDPEAWQTLNK 55
+S M ++++ ++++ R RP+++ PT + S VED G+ E W +L
Sbjct: 194 ASLMYNSSYEPRQDKSRVRPTSSDSMKTLQ*LVSEPTCQENSVVEDDFPGETEDWVSLRN 15
Query: 56 SF 57
F
Sbjct: 14 PF 9
>TC221081 homologue to UP|Q9LR10 (Q9LR10) F10A5.10, partial (29%)
Length = 633
Score = 26.2 bits (56), Expect = 4.6
Identities = 16/51 (31%), Positives = 27/51 (52%), Gaps = 4/51 (7%)
Frame = +3
Query: 72 QQVNENQQSRMPDNMVKNVGLIQELNGNI----SKVASLYSDLNSDFTNIC 118
QQ + Q+ + +KN+ +++ G+ SKV S YS L S ++IC
Sbjct: 81 QQQQQKQKPASSWDQIKNLLTCKQIEGSRVHDPSKVVSGYSKLGSSCSSIC 233
>TC215183
Length = 2081
Score = 26.2 bits (56), Expect = 4.6
Identities = 14/60 (23%), Positives = 29/60 (48%)
Frame = +1
Query: 46 DPEAWQTLNKSFRQVQSVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQELNGNISKVAS 105
+PE + L + Q+Q ++DRN + +N+ R +N +++E+ N + S
Sbjct: 676 NPEIVRNLIMNNPQMQELMDRNPELAHILNDPSTLRQTLEATRNPEIMREMMRNTDRAMS 855
>TC232770 homologue to UP|Q6QUQ3 (Q6QUQ3) Auxin and ethylene responsive
GH3-like protein, partial (35%)
Length = 744
Score = 25.8 bits (55), Expect = 6.0
Identities = 10/26 (38%), Positives = 15/26 (57%)
Frame = +3
Query: 16 HRSKKNQRRRRPSATPTNNDYSDVED 41
HR +QRRRRP +P + ++ D
Sbjct: 186 HRRNDSQRRRRPGESPGGDSHTQRPD 263
>TC223149 similar to UP|Q9LFT7 (Q9LFT7) Protein serine/threonine kinase-like
protein, partial (17%)
Length = 663
Score = 25.8 bits (55), Expect = 6.0
Identities = 13/26 (50%), Positives = 16/26 (61%)
Frame = +1
Query: 3 TLYSPSSAMEDTNHRSKKNQRRRRPS 28
TLY+ ++ TN R KN RRRR S
Sbjct: 400 TLYASNTRGPSTNVRGCKNARRRRVS 477
>TC213605 weakly similar to UP|RN12_HUMAN (Q9NVW2) RING finger protein 12
(LIM domain interacting RING finger protein) (RING
finger LIM domain-binding protein) (R-LIM) (NY-REN-43
antigen), partial (6%)
Length = 704
Score = 25.8 bits (55), Expect = 6.0
Identities = 15/49 (30%), Positives = 24/49 (48%), Gaps = 2/49 (4%)
Frame = -3
Query: 11 MEDTNHRSKKNQRRRRPSATPT--NNDYSDVEDAEDGDPEAWQTLNKSF 57
MED +H KK+ ++ S + ++ D G E WQ++NK F
Sbjct: 312 MEDDDHAEKKSYPKKTKSKKRKG*SAEFPQTRDG-CGQIECWQSINKFF 169
>TC231065
Length = 1117
Score = 25.4 bits (54), Expect = 7.9
Identities = 10/34 (29%), Positives = 21/34 (61%)
Frame = +3
Query: 62 SVLDRNRAIIQQVNENQQSRMPDNMVKNVGLIQE 95
+++DR + NEN++ R+ +NM+K + + E
Sbjct: 351 ALMDRLLTLNNCTNENEKERVKENMLKLIDIYYE 452
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.309 0.123 0.340
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 4,668,513
Number of Sequences: 63676
Number of extensions: 44151
Number of successful extensions: 303
Number of sequences better than 10.0: 41
Number of HSP's better than 10.0 without gapping: 302
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 302
length of query: 130
length of database: 12,639,632
effective HSP length: 86
effective length of query: 44
effective length of database: 7,163,496
effective search space: 315193824
effective search space used: 315193824
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.1 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 42 (21.7 bits)
S2: 53 (25.0 bits)
Medicago: description of AC145219.5


