

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145218.1 - phase: 0
(63 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
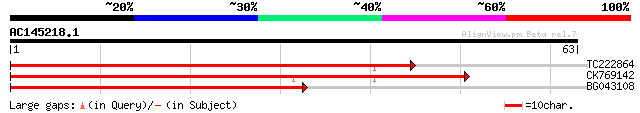
Sequences producing significant alignments: (bits) Value
TC222864 similar to GB|AAO64841.1|29028924|BT005906 At5g63450 {A... 55 5e-09
CK769142 54 2e-08
BG043108 49 7e-07
>TC222864 similar to GB|AAO64841.1|29028924|BT005906 At5g63450 {Arabidopsis
thaliana;} , partial (11%)
Length = 570
Score = 55.5 bits (132), Expect = 5e-09
Identities = 29/49 (59%), Positives = 36/49 (73%), Gaps = 4/49 (8%)
Frame = +1
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFSFARDEK----IFDLRD 45
MASLEL SV +R+YA+E++NEEIH RLIP M S AR E + DL+D
Sbjct: 424 MASLELGSVAIRTYAMELVNEEIHARLIPIMESTARGELNSVCVLDLQD 570
>CK769142
Length = 753
Score = 53.5 bits (127), Expect = 2e-08
Identities = 31/57 (54%), Positives = 39/57 (68%), Gaps = 6/57 (10%)
Frame = +2
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPF-MFSFARDEK-----IFDLRDIMRSSS 51
MASLEL SV +R+ A+E++NEEIH RLIPF M S DE + DL+DI+R S
Sbjct: 488 MASLELGSVAIRTNAMELVNEEIHARLIPFIMGSVTHDELNDSVCVLDLQDILRRFS 658
>BG043108
Length = 451
Score = 48.5 bits (114), Expect = 7e-07
Identities = 22/33 (66%), Positives = 28/33 (84%)
Frame = +2
Query: 1 MASLELTSVTVRSYALEIINEEIHTRLIPFMFS 33
MASLEL V +R+YA+E++N+EIHTRLIP M S
Sbjct: 353 MASLELCGVAMRTYAMELVNKEIHTRLIPIMES 451
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.135 0.352
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 1,802,031
Number of Sequences: 63676
Number of extensions: 13559
Number of successful extensions: 88
Number of sequences better than 10.0: 6
Number of HSP's better than 10.0 without gapping: 88
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 88
length of query: 63
length of database: 12,639,632
effective HSP length: 39
effective length of query: 24
effective length of database: 10,156,268
effective search space: 243750432
effective search space used: 243750432
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 40 (21.6 bits)
S2: 52 (24.6 bits)
Medicago: description of AC145218.1


