

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
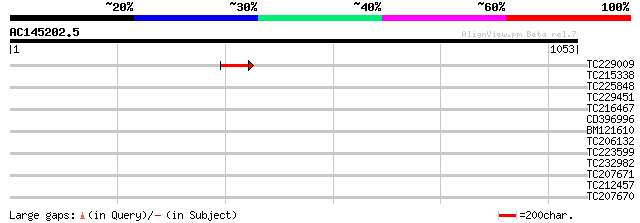
Query= AC145202.5 - phase: 0 /pseudo
(1053 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
TC229009 similar to UP|Q94BZ7 (Q94BZ7) AT3g10270/F14P13_13, part... 49 2e-05
TC215338 UP|UGDH_SOYBN (Q96558) UDP-glucose 6-dehydrogenase (UD... 33 0.50
TC225848 homologue to UP|SUS2_PEA (O24301) Sucrose synthase 2 (... 32 1.5
TC229451 similar to UP|Q9FH68 (Q9FH68) Arabidopsis thaliana geno... 31 3.3
TC216467 similar to UP|Q6RK07 (Q6RK07) UDP-glucose dehydrogenase... 31 3.3
CD396996 30 7.3
BM121610 30 7.3
TC206132 similar to GB|AAL11601.1|15983466|AF424607 At1g33990/F1... 30 7.3
TC223599 UP|Q9STY2 (Q9STY2) Hypthetical protein, partial (5%) 30 7.3
TC232982 29 9.5
TC207671 similar to UP|Q9FIL1 (Q9FIL1) Protein kinase-like prote... 29 9.5
TC212457 UP|Q39871 (Q39871) Maturation polypeptide, complete 29 9.5
TC207670 similar to UP|Q9FIL1 (Q9FIL1) Protein kinase-like prote... 29 9.5
>TC229009 similar to UP|Q94BZ7 (Q94BZ7) AT3g10270/F14P13_13, partial (30%)
Length = 1122
Score = 48.5 bits (114), Expect = 2e-05
Identities = 23/63 (36%), Positives = 40/63 (62%)
Frame = +1
Query: 391 RKLFKKKEIQNLMAILGLVRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFYSFW 450
++L+K +EIQNL+ LGL + LRY +++++ + D DGAH + LL+ FF+ +
Sbjct: 130 QQLYKNEEIQNLILGLGLGVKGEDFKKDALRYHKIIILTDADVDGAHIRTLLLTFFFRYQ 309
Query: 451 PSL 453
+L
Sbjct: 310 RAL 318
>TC215338 UP|UGDH_SOYBN (Q96558) UDP-glucose 6-dehydrogenase (UDP-Glc
dehydrogenase) (UDP-GlcDH) (UDPGDH) , complete
Length = 1780
Score = 33.5 bits (75), Expect = 0.50
Identities = 28/117 (23%), Positives = 47/117 (39%), Gaps = 11/117 (9%)
Frame = +3
Query: 409 VRNKKYSNAKCLRYGQLMVMANQDEDGAHFKGLLINFFY-----------SFWPSLLKVS 457
V+ YS R G + A+ G+ F+ ++N Y +W ++K++
Sbjct: 798 VQQVSYSVGTDSRIGPKFLNASVGFGGSCFQKDILNLVYICECNGLPEVAEYWKQVIKIN 977
Query: 458 KFMSVFTFPIIKVSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGL 514
+ + SM+N +KK L F +KK+ G+T I CQGL
Sbjct: 978 DYQKSRFVNRVVASMFNTVSNKKIAILGF-------AFKKDTGDTRETPAIDVCQGL 1127
>TC225848 homologue to UP|SUS2_PEA (O24301) Sucrose synthase 2 (Sucrose-UDP
glucosyltransferase 2) , partial (83%)
Length = 2487
Score = 32.0 bits (71), Expect = 1.5
Identities = 24/51 (47%), Positives = 30/51 (58%), Gaps = 10/51 (19%)
Frame = +2
Query: 829 GLIESFRQKGDY-EIVNFKV-----NVKQ----EEIATIEKDEELRKKFNL 869
GL+ESF + E+VN V +VK+ EEIA IEK EL KK+NL
Sbjct: 1379 GLVESFGKNSKLRELVNLVVVAGYIDVKKSSDREEIAEIEKMHELMKKYNL 1531
>TC229451 similar to UP|Q9FH68 (Q9FH68) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K16E1, partial (80%)
Length = 850
Score = 30.8 bits (68), Expect = 3.3
Identities = 21/48 (43%), Positives = 29/48 (59%), Gaps = 1/48 (2%)
Frame = +2
Query: 986 QSVSIEGATLGDYEYLSSLPFENFTLES-LKKLEAELDEKENELKTLE 1032
QS S E G+ E +L E TL+S +KKLE+E + K ++ KTLE
Sbjct: 494 QSRSFEDGKNGNSEEHKALNEEITTLKSKIKKLESECEAKGSQAKTLE 637
>TC216467 similar to UP|Q6RK07 (Q6RK07) UDP-glucose dehydrogenase, partial
(83%)
Length = 1485
Score = 30.8 bits (68), Expect = 3.3
Identities = 25/105 (23%), Positives = 44/105 (41%), Gaps = 11/105 (10%)
Frame = +3
Query: 421 RYGQLMVMANQDEDGAHFKGLLINFFY-----------SFWPSLLKVSKFMSVFTFPIIK 469
R G + A+ G+ F+ ++N Y ++W ++KV+ + + +
Sbjct: 528 RIGPKFLNASVGFGGSCFQKDILNLVYICECNGLPEVANYWKQVIKVNDYQKMRFVNRVV 707
Query: 470 VSMYNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGL 514
SM+N KK L F +KK+ G+T I C+GL
Sbjct: 708 SSMFNTVSGKKIAILGF-------AFKKDTGDTRETPAIDVCKGL 821
>CD396996
Length = 450
Score = 29.6 bits (65), Expect = 7.3
Identities = 14/28 (50%), Positives = 18/28 (64%), Gaps = 2/28 (7%)
Frame = -3
Query: 856 TIEKDEELRKKFNLTCTIS--TSNMYLF 881
TIEK +E+ K F +TCTI N+Y F
Sbjct: 106 TIEKSKEIGKLFQITCTIQYRLGNLYCF 23
>BM121610
Length = 742
Score = 29.6 bits (65), Expect = 7.3
Identities = 16/40 (40%), Positives = 21/40 (52%)
Frame = -1
Query: 735 HAMGTCFSSDIPKYHPYEIIKNVRRFLNNEEMVSMKPWYK 774
H M C + D +YH + I+ V L+ E VS KPW K
Sbjct: 205 HPMFECKTGD--QYHICQQIETVSPLLDQEGYVSWKPWLK 92
>TC206132 similar to GB|AAL11601.1|15983466|AF424607 At1g33990/F12G12_220
{Arabidopsis thaliana;} , partial (33%)
Length = 701
Score = 29.6 bits (65), Expect = 7.3
Identities = 16/40 (40%), Positives = 21/40 (52%)
Frame = -2
Query: 735 HAMGTCFSSDIPKYHPYEIIKNVRRFLNNEEMVSMKPWYK 774
H M C + D +YH + I+ V L+ E VS KPW K
Sbjct: 211 HPMFECKTGD--QYHICQQIETVSPLLDQEGYVSWKPWLK 98
>TC223599 UP|Q9STY2 (Q9STY2) Hypthetical protein, partial (5%)
Length = 518
Score = 29.6 bits (65), Expect = 7.3
Identities = 14/33 (42%), Positives = 18/33 (54%)
Frame = +2
Query: 839 DYEIVNFKVNVKQEEIATIEKDEELRKKFNLTC 871
DYE V N+K +E+A EK + RKK C
Sbjct: 254 DYEAVKLSQNMKCKEVAVGEKQKRRRKKKKTIC 352
>TC232982
Length = 589
Score = 29.3 bits (64), Expect = 9.5
Identities = 16/44 (36%), Positives = 25/44 (56%), Gaps = 3/44 (6%)
Frame = +3
Query: 579 FVNKELLFSSMKNLQRSIPSVVDGLTLGQRKTL---FCSFEMNL 619
F +KEL+F S + R PS DG+ + + + L +C+F NL
Sbjct: 330 FEDKELVFMSRDEIDRCSPSRKDGIDVLRERHLRYSYCAFLQNL 461
>TC207671 similar to UP|Q9FIL1 (Q9FIL1) Protein kinase-like protein
(AT5g59010/k19m22_210), partial (16%)
Length = 504
Score = 29.3 bits (64), Expect = 9.5
Identities = 15/31 (48%), Positives = 19/31 (60%)
Frame = +2
Query: 664 LLIPYGQFGTRDLGGKDHASSRKLYIKLNCV 694
LL+PY Q GTR G+ HASS + NC+
Sbjct: 164 LLLPYDQHGTRST-GRCHASSISIPHMANCI 253
>TC212457 UP|Q39871 (Q39871) Maturation polypeptide, complete
Length = 1741
Score = 29.3 bits (64), Expect = 9.5
Identities = 17/70 (24%), Positives = 31/70 (44%)
Frame = +1
Query: 473 YNASDSKKGKELSFDSMQQYEDWKKELGNTANDWEIKYCQGLSSAQEVREYFQDLGKHRA 532
Y A +K+GK+ + + M +Y+D+ E D L E+++ D K
Sbjct: 733 YAAEKTKEGKDATVNKMGEYKDYTAEKAKEGKD------TTLGKLGELKDTASDAAKRAV 894
Query: 533 YFVWDDKDET 542
++ K+ET
Sbjct: 895 GYLSGKKEET 924
>TC207670 similar to UP|Q9FIL1 (Q9FIL1) Protein kinase-like protein
(AT5g59010/k19m22_210), partial (35%)
Length = 800
Score = 29.3 bits (64), Expect = 9.5
Identities = 15/31 (48%), Positives = 19/31 (60%)
Frame = +2
Query: 664 LLIPYGQFGTRDLGGKDHASSRKLYIKLNCV 694
LL+PY Q GTR G+ HASS + NC+
Sbjct: 341 LLLPYDQHGTRST-GRCHASSISIPHMANCI 430
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.323 0.139 0.411
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 42,972,694
Number of Sequences: 63676
Number of extensions: 565529
Number of successful extensions: 2648
Number of sequences better than 10.0: 26
Number of HSP's better than 10.0 without gapping: 2626
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 2648
length of query: 1053
length of database: 12,639,632
effective HSP length: 107
effective length of query: 946
effective length of database: 5,826,300
effective search space: 5511679800
effective search space used: 5511679800
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.5 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 64 (29.3 bits)
Medicago: description of AC145202.5


