

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145202.10 + phase: 0
(235 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
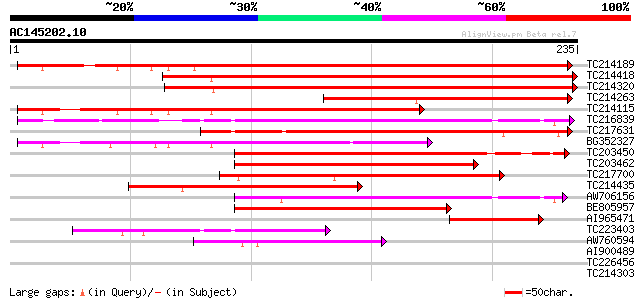
Score E
Sequences producing significant alignments: (bits) Value
TC214189 homologue to UP|Q8L949 (Q8L949) Plastid protein, partia... 333 5e-92
TC214418 homologue to UP|Q8L949 (Q8L949) Plastid protein, partia... 301 2e-82
TC214320 similar to UP|O22793 (O22793) Plastid protein (At2g3343... 287 3e-78
TC214263 homologue to UP|Q8L949 (Q8L949) Plastid protein, partia... 183 7e-47
TC214115 similar to UP|O22793 (O22793) Plastid protein (At2g3343... 180 4e-46
TC216839 152 1e-37
TC217631 similar to UP|WAIP_HUMAN (O43516) Wiskott-Aldrich syndr... 141 2e-34
BG352327 similar to PIR|D84745|D847 plastid protein [imported] -... 140 5e-34
TC203450 homologue to UP|DAG_ANTMA (Q38732) DAG protein, chlorop... 139 1e-33
TC203462 homologue to UP|DAG_ANTMA (Q38732) DAG protein, chlorop... 123 6e-29
TC217700 similar to PIR|T04886|T04886 DAG protein homolog F18F4.... 115 2e-26
TC214435 similar to UP|O22793 (O22793) Plastid protein (At2g3343... 108 2e-24
AW706156 102 2e-22
BE805957 90 8e-19
AI965471 homologue to GP|21617909|gb| plastid protein {Arabidops... 71 5e-13
TC223403 62 2e-10
AW760594 59 2e-09
AI900489 40 0.001
TC226456 39 0.002
TC214303 similar to UP|Q9T0K5 (Q9T0K5) Extensin-like protein, pa... 38 0.003
>TC214189 homologue to UP|Q8L949 (Q8L949) Plastid protein, partial (71%)
Length = 969
Score = 333 bits (853), Expect = 5e-92
Identities = 172/242 (71%), Positives = 198/242 (81%), Gaps = 12/242 (4%)
Frame = +2
Query: 4 RAIASSFKR---RLFSTTTSTSTSTSTYTTTVAGALSKPSSLT--SPRFIIPMSQTIFQ- 57
RA++SS R RLFSTTT+T+T+T+ + T L KPSSL+ + RF+ P+S I
Sbjct: 56 RAVSSSCARLTMRLFSTTTTTTTTTAAFAVT----LPKPSSLSVSTRRFLFPLSHAIVPP 223
Query: 58 SLHFGGI----HRAGYCNYSPL--VPGSNTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPG 111
S GI +RAG YSPL S++SFSDRPP EMAPLFPGCDYNHWLI++D PG
Sbjct: 224 STRVAGIRCRVNRAGDSAYSPLNSASSSSSSFSDRPPTEMAPLFPGCDYNHWLIVMDHPG 403
Query: 112 GEGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGV 171
GEGATKQQMIDCY++TLA+VLGSEEEAKKKIYNVSCERYFGFGCE+DEETSNKLEG+PGV
Sbjct: 404 GEGATKQQMIDCYIQTLAKVLGSEEEAKKKIYNVSCERYFGFGCEIDEETSNKLEGLPGV 583
Query: 172 LFVLPDSYVDPEHQDYGAELFVNGEIVQRSPERQRRVEPQAQRGDSRPRYHDRTKYVRRR 231
LFVLPDSYVDPE++DYGAELFVNGEIVQRSPERQRRVEPQ QR RPRY+DRT+YVRRR
Sbjct: 584 LFVLPDSYVDPENKDYGAELFVNGEIVQRSPERQRRVEPQPQRHQDRPRYNDRTRYVRRR 763
Query: 232 DN 233
+N
Sbjct: 764 EN 769
>TC214418 homologue to UP|Q8L949 (Q8L949) Plastid protein, partial (74%)
Length = 1130
Score = 301 bits (771), Expect = 2e-82
Identities = 141/173 (81%), Positives = 159/173 (91%), Gaps = 1/173 (0%)
Frame = +3
Query: 64 IHRAGYCNYSPLVPGSNTS-FSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQMID 122
++RAG YSPL GS++S FSDRPP EMAPLFPGCDYNHWLI+++ PGGEGA KQQMID
Sbjct: 39 VNRAGDSAYSPLNSGSSSSSFSDRPPTEMAPLFPGCDYNHWLIVMENPGGEGANKQQMID 218
Query: 123 CYVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDP 182
CY++TLA+VLGSEEEAKKKIYNVSCERYFGFGCE+DEETSNKLEG+PGVLFVLPDSYVDP
Sbjct: 219 CYIQTLAKVLGSEEEAKKKIYNVSCERYFGFGCEIDEETSNKLEGLPGVLFVLPDSYVDP 398
Query: 183 EHQDYGAELFVNGEIVQRSPERQRRVEPQAQRGDSRPRYHDRTKYVRRRDNQR 235
E++DYGAELFVNGEIVQRSPERQRRVEPQ QR RPRY+DRT+YVRRR+N +
Sbjct: 399 ENKDYGAELFVNGEIVQRSPERQRRVEPQPQRHQDRPRYNDRTRYVRRRENMQ 557
>TC214320 similar to UP|O22793 (O22793) Plastid protein (At2g33430/F4P9.20),
partial (68%)
Length = 1045
Score = 287 bits (735), Expect = 3e-78
Identities = 135/175 (77%), Positives = 152/175 (86%), Gaps = 4/175 (2%)
Frame = +3
Query: 65 HRAGYCNYSPLVPGSNTSF----SDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQM 120
+R G YSPL G TS ++RPP +MAPLFPGCDY+HWL++I KPGGEGATKQQM
Sbjct: 255 NRPGDSAYSPLNTGKPTSILSSITERPPTDMAPLFPGCDYHHWLVVIHKPGGEGATKQQM 434
Query: 121 IDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYV 180
IDCYV+TLA+VLGSEEEAKKKIYNVSCERYF FGC++DEETSNKLEG+PGVLFVLPDSYV
Sbjct: 435 IDCYVQTLAKVLGSEEEAKKKIYNVSCERYFAFGCDIDEETSNKLEGLPGVLFVLPDSYV 614
Query: 181 DPEHQDYGAELFVNGEIVQRSPERQRRVEPQAQRGDSRPRYHDRTKYVRRRDNQR 235
DPEH+DYG ELFVNGEIVQR PERQRRVEPQ QR RPRY+DRT+YVRRR+N +
Sbjct: 615 DPEHKDYGGELFVNGEIVQRPPERQRRVEPQPQRQQDRPRYNDRTRYVRRRENTK 779
>TC214263 homologue to UP|Q8L949 (Q8L949) Plastid protein, partial (47%)
Length = 581
Score = 183 bits (464), Expect = 7e-47
Identities = 92/125 (73%), Positives = 99/125 (78%), Gaps = 22/125 (17%)
Frame = +2
Query: 131 VLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEG----------------------I 168
VLGSEEEAKKKIYNVSCERYFGFGCE+DEETSNKLEG +
Sbjct: 2 VLGSEEEAKKKIYNVSCERYFGFGCEIDEETSNKLEGMFSEVEVFEISFLCSLFFICVGL 181
Query: 169 PGVLFVLPDSYVDPEHQDYGAELFVNGEIVQRSPERQRRVEPQAQRGDSRPRYHDRTKYV 228
PGVLFVLPDSYVDPE++DYGAELFVNGEIVQRSPERQRRVEPQ QR RPRY+DRT+YV
Sbjct: 182 PGVLFVLPDSYVDPENKDYGAELFVNGEIVQRSPERQRRVEPQPQRHQDRPRYNDRTRYV 361
Query: 229 RRRDN 233
RRR+N
Sbjct: 362 RRREN 376
>TC214115 similar to UP|O22793 (O22793) Plastid protein (At2g33430/F4P9.20),
partial (46%)
Length = 557
Score = 180 bits (457), Expect = 4e-46
Identities = 103/180 (57%), Positives = 123/180 (68%), Gaps = 11/180 (6%)
Frame = +3
Query: 4 RAIASSFKR---RLFSTTTSTSTSTSTYTTTVAGALSKPSSLT--SPRFIIPMSQTIFQ- 57
R I+SS R RLFSTTT+ T A L KPSSL+ S RF+ P+S I
Sbjct: 42 RVISSSCARLAMRLFSTTTTA--------TAFAVTLPKPSSLSLSSRRFLFPLSHAIVPP 197
Query: 58 SLHFGGI----HRAGYCNYSPLVPGSNTS-FSDRPPAEMAPLFPGCDYNHWLIIIDKPGG 112
S GI +RAG YSPL GS++S FSDRPP EMAPLFPGCDYNHWLI+++ PGG
Sbjct: 198 STRVAGIRCRVNRAGDSAYSPLNSGSSSSSFSDRPPTEMAPLFPGCDYNHWLIVMENPGG 377
Query: 113 EGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVL 172
EGA KQQMIDCY++TLA+VLGSE+EAK+K YN C+ Y GFGCE+DEET+ G+ VL
Sbjct: 378 EGANKQQMIDCYIQTLAKVLGSEDEAKRKTYNSFCDTYCGFGCEIDEETTK*HGGLADVL 557
>TC216839
Length = 1055
Score = 152 bits (385), Expect = 1e-37
Identities = 101/234 (43%), Positives = 135/234 (57%), Gaps = 3/234 (1%)
Frame = +1
Query: 4 RAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALSKPSSLTSPRFIIPMSQTIFQSLHFGG 63
R +AS+ R L +S+S S S + A AL P+ T+P QS G
Sbjct: 40 RTLASTLSRAL----SSSSASASRFRFAFAFALL-PAKQTAPNPHWASFAVRTQSSGSG- 201
Query: 64 IHRAGYCNYSPLVPGSNTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQMIDC 123
YSPL S ++S+RPP E L GCDY HWLI+++ P ++ M++
Sbjct: 202 --------YSPLNDPS-PNWSNRPPKETI-LLDGCDYEHWLIVMEFPDNPKPSEDHMVNA 351
Query: 124 YVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDPE 183
YVKTLAQVLGSEE+AK KIY+VS Y GFG + EE S K++ +PGVL+VLPDSY+D
Sbjct: 352 YVKTLAQVLGSEEDAKNKIYSVSTSTYTGFGALISEELSYKVKELPGVLWVLPDSYLDVP 531
Query: 184 HQDYGAELFVNGEIVQRSPERQRRVEPQAQRGDSRPRYHDR---TKYVRRRDNQ 234
++DYG +LFV+G+++ R + R + Q R RPR HDR T V RRD Q
Sbjct: 532 NKDYGGDLFVDGKVIPR--PQYRYSDRQPSRSRPRPR-HDRRRETMQVERRDQQ 684
>TC217631 similar to UP|WAIP_HUMAN (O43516) Wiskott-Aldrich syndrome protein
interacting protein (WASP interacting protein) (PRPL-2
protein), partial (7%)
Length = 1440
Score = 141 bits (356), Expect = 2e-34
Identities = 72/161 (44%), Positives = 106/161 (65%), Gaps = 7/161 (4%)
Frame = +1
Query: 80 NTSFSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSEEEAK 139
N ++S+RPP E L GCD+ HWL++++KP G+ T+ +ID Y+KTLA+V+GSEEEA+
Sbjct: 301 NPNWSNRPPKETI-LLDGCDFEHWLVVMEKPEGD-PTRDDIIDSYIKTLAKVIGSEEEAR 474
Query: 140 KKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAELFVNGEIVQ 199
KIY+VS YF FG + EE S K++ +PGV +VLPDSY++ + +DYG E F+NG+
Sbjct: 475 MKIYSVSTRHYFAFGALVSEELSYKIKELPGVRWVLPDSYLNVKEKDYGGEPFINGQAAP 654
Query: 200 RSPE------RQRRVEPQAQRGDSRPRYHDRTK-YVRRRDN 233
P+ R + R + RPR DR++ + RRR+N
Sbjct: 655 YDPKYHEEWVRNNARANERNRRNDRPRNADRSRNFERRREN 777
>BG352327 similar to PIR|D84745|D847 plastid protein [imported] - Arabidopsis
thaliana, partial (23%)
Length = 526
Score = 140 bits (353), Expect = 5e-34
Identities = 87/183 (47%), Positives = 110/183 (59%), Gaps = 11/183 (6%)
Frame = +2
Query: 4 RAIASSFKR---RLFSTTTSTSTSTSTYTTTVAGALSKPS--SLTSPRFIIPMSQTIFQS 58
R I+SS R RLFSTTT+ T AG L KPS SL+S RF+ P+ I
Sbjct: 5 RVISSSCARLAMRLFSTTTTA--------TAFAGTLPKPSLLSLSSRRFLFPLFHAIVPP 160
Query: 59 L-HFGGI----HRAGYCNYSPLVPGSNTS-FSDRPPAEMAPLFPGCDYNHWLIIIDKPGG 112
GI +RAG YSP+ GS++S FSD PP EM P+FPGCDYNHWLI+++ PG
Sbjct: 161 FTRVAGIRCRVNRAGDSGYSPVNSGSSSSSFSDPPPTEMGPVFPGCDYNHWLIVMENPGW 340
Query: 113 EGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVL 172
EGA +QQMIDC+++TL + ++ YNVSCER CE+ ++ N LEG GVL
Sbjct: 341 EGANQQQMIDCFIQTLTKFSA*RRN*EEN-YNVSCERTLDSACEMMKKLLNSLEGWLGVL 517
Query: 173 FVL 175
F L
Sbjct: 518 FGL 526
>TC203450 homologue to UP|DAG_ANTMA (Q38732) DAG protein, chloroplast
precursor, partial (53%)
Length = 1196
Score = 139 bits (349), Expect = 1e-33
Identities = 69/139 (49%), Positives = 94/139 (66%)
Frame = +3
Query: 94 LFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGF 153
+ PGCDYNHWLI+++ P T++QMID Y+ TLA VLGS EEAKK +Y S Y GF
Sbjct: 288 MLPGCDYNHWLIVMEFPKDPAPTREQMIDTYLDTLATVLGSMEEAKKNMYAFSTTTYTGF 467
Query: 154 GCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAELFVNGEIVQRSPERQRRVEPQAQ 213
C +DE TS K +G+PGVL+VLPDSY+D +++DYG + ++NGEI+ P + +P+
Sbjct: 468 QCTVDEATSEKFKGLPGVLWVLPDSYIDVKNKDYGGDKYINGEII---PCKYPTYQPKR- 635
Query: 214 RGDSRPRYHDRTKYVRRRD 232
S P+ R +Y RRRD
Sbjct: 636 ---SAPKNESR-RYERRRD 680
>TC203462 homologue to UP|DAG_ANTMA (Q38732) DAG protein, chloroplast
precursor, partial (53%)
Length = 708
Score = 123 bits (309), Expect = 6e-29
Identities = 55/101 (54%), Positives = 73/101 (71%)
Frame = +1
Query: 94 LFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGF 153
+ PGCDYNHWLI+++ P T++QMID Y+ TLA VLGS EEAKK +Y S Y GF
Sbjct: 265 MLPGCDYNHWLIVMEFPKDPAPTREQMIDTYLDTLATVLGSMEEAKKNMYAFSTTTYTGF 444
Query: 154 GCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAELFVN 194
C +DE TS K +G+PGVL+VLPDSY+D +++DYG ++
Sbjct: 445 QCTVDEATSEKFKGLPGVLWVLPDSYIDVKNKDYGGFFLIS 567
>TC217700 similar to PIR|T04886|T04886 DAG protein homolog F18F4.120 -
Arabidopsis thaliana {Arabidopsis thaliana;} , partial
(28%)
Length = 760
Score = 115 bits (287), Expect = 2e-26
Identities = 57/122 (46%), Positives = 77/122 (62%), Gaps = 4/122 (3%)
Frame = +2
Query: 88 PAEMAP---LFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLG-SEEEAKKKIY 143
P E+ P LF GCDYNHWLI+++ + + M+ Y +T A+ L S EEAKKK+Y
Sbjct: 197 PDEIGPDTILFEGCDYNHWLIVMEFKDNNKPSPEDMVRVYEETCAKGLNISLEEAKKKMY 376
Query: 144 NVSCERYFGFGCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAELFVNGEIVQRSPE 203
S Y GF + EE S K EG+PGV+FVLPDSY+DP ++ YG + ++NG I+ R P
Sbjct: 377 ACSTTTYIGFQAVMTEEESKKFEGLPGVIFVLPDSYIDPVNKQYGGDQYINGTIIPRPPP 556
Query: 204 RQ 205
Q
Sbjct: 557 VQ 562
>TC214435 similar to UP|O22793 (O22793) Plastid protein (At2g33430/F4P9.20),
partial (25%)
Length = 420
Score = 108 bits (271), Expect = 2e-24
Identities = 54/105 (51%), Positives = 68/105 (64%), Gaps = 8/105 (7%)
Frame = +1
Query: 50 PMSQTIFQSLHFGGIHRAGYC--------NYSPLVPGSNTSFSDRPPAEMAPLFPGCDYN 101
P S +++ + H HR + S V G + + +PLFPGCDYN
Sbjct: 106 PRSPSLYPNPHHSPSHRVXFLFPLSHAIVPPSTRVAGIRCRVNRAGDSAYSPLFPGCDYN 285
Query: 102 HWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVS 146
HWLI+++ PGGEGA KQQMIDCY++TLA+VLGSEEEAKKKIYNVS
Sbjct: 286 HWLIVMENPGGEGANKQQMIDCYIQTLAKVLGSEEEAKKKIYNVS 420
Score = 27.3 bits (59), Expect = 6.2
Identities = 26/62 (41%), Positives = 30/62 (47%), Gaps = 5/62 (8%)
Frame = +3
Query: 4 RAIASSFKR---RLFSTTTSTSTSTSTYTTTVAGALSKPS--SLTSPRFIIPMSQTIFQS 58
R I+SS R RLFSTTT+ T A L KPS SL+S R +P S
Sbjct: 42 RVISSSCARLAMRLFSTTTTA--------TAFAVTLPKPSSLSLSSRRIPLPSLPCHRPS 197
Query: 59 LH 60
LH
Sbjct: 198LH 203
>AW706156
Length = 424
Score = 102 bits (253), Expect = 2e-22
Identities = 63/142 (44%), Positives = 82/142 (57%), Gaps = 4/142 (2%)
Frame = +2
Query: 94 LFPGCDYNHWLIIIDKPG-GEGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFG 152
L GCDY HWLI+++ P + + YVKTLAQVLG EEEAKKKIY S G
Sbjct: 8 LLNGCDY*HWLIVMESPR*PYTLLGHKWLTSYVKTLAQVLGCEEEAKKKIYADSTSTSSG 187
Query: 153 FGCELDEETSNKLEGIPGVLFVLPDSYVDPEHQDYGAELFVNGEIVQRSPERQRRVEPQA 212
FG + +E S K+ +PGV +VLPDS +D + DYG + +G+++ R + R + Q
Sbjct: 188 FGALISDELSYKVTELPGVFWVLPDSCLDDTNYDYGGD*CGDGKVIPR--PQYRYSDRQP 361
Query: 213 QRGDSRPRYHDR---TKYVRRR 231
R RPR HDR T V RR
Sbjct: 362 SRSRPRPR-HDRQRQTMQVERR 424
>BE805957
Length = 426
Score = 90.1 bits (222), Expect = 8e-19
Identities = 42/90 (46%), Positives = 62/90 (68%)
Frame = +3
Query: 94 LFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGSEEEAKKKIYNVSCERYFGF 153
LFP + HWL+ +DKPG E TK Q++D Y + L +V+G+E++A+ IY+VS + FGF
Sbjct: 156 LFPAGNSKHWLVKMDKPGVEAVTKAQIVDYYAQILTKVMGNEKDAQMCIYHVSWKTNFGF 335
Query: 154 GCELDEETSNKLEGIPGVLFVLPDSYVDPE 183
CELDE+ + +L G+ GVL V PD+ + E
Sbjct: 336 CCELDEDCAQELAGVLGVLSVQPDNNFESE 425
Score = 27.7 bits (60), Expect = 4.7
Identities = 13/32 (40%), Positives = 19/32 (58%)
Frame = +3
Query: 156 ELDEETSNKLEGIPGVLFVLPDSYVDPEHQDY 187
++DEE S +L +P VL V PD + +DY
Sbjct: 3 DIDEEISAQLASLPEVLLVRPDLEFNSLKKDY 98
>AI965471 homologue to GP|21617909|gb| plastid protein {Arabidopsis
thaliana}, partial (17%)
Length = 421
Score = 70.9 bits (172), Expect = 5e-13
Identities = 33/39 (84%), Positives = 35/39 (89%)
Frame = +1
Query: 183 EHQDYGAELFVNGEIVQRSPERQRRVEPQAQRGDSRPRY 221
E++DYGAELFVNGEIVQRSPERQRRVEPQ QR RPRY
Sbjct: 304 ENKDYGAELFVNGEIVQRSPERQRRVEPQPQRHQDRPRY 420
>TC223403
Length = 424
Score = 62.0 bits (149), Expect = 2e-10
Identities = 43/113 (38%), Positives = 59/113 (52%), Gaps = 6/113 (5%)
Frame = +2
Query: 27 TYTTTVAGALSKPSSLTSP-RFIIPMSQT-----IFQSLHFGGIHRAGYCNYSPLVPGSN 80
T +AGALS S+ S RF + + I F ++ YSPL S
Sbjct: 17 TLAYRLAGALSSSSASASRCRFALALHHAKQTVPIPHPASFAVRTQSSGSGYSPLNDPS- 193
Query: 81 TSFSDRPPAEMAPLFPGCDYNHWLIIIDKPGGEGATKQQMIDCYVKTLAQVLG 133
++S+RPP E L GCDY HWLI+++ P ++ M++ YVKTLAQVLG
Sbjct: 194 PNWSNRPPKETI-LLDGCDYEHWLIVMEFPDNPKPSEDHMVNSYVKTLAQVLG 349
>AW760594
Length = 424
Score = 58.5 bits (140), Expect = 2e-09
Identities = 34/82 (41%), Positives = 47/82 (56%), Gaps = 2/82 (2%)
Frame = +1
Query: 77 PGSNTSFSDRPPAEMAPLF-PGCDYN-HWLIIIDKPGGEGATKQQMIDCYVKTLAQVLGS 134
P N S + A P F P N HW++++D P +K Q+ID YVKTL VLGS
Sbjct: 178 PCCNWSITRVAAATTYPSFNPSTTQNPHWMVLMDTPPQGVNSKPQVIDYYVKTLQTVLGS 357
Query: 135 EEEAKKKIYNVSCERYFGFGCE 156
E++A+ IY+ S +FGF C+
Sbjct: 358 EKDAQMCIYDASWNTHFGFCCD 423
>AI900489
Length = 109
Score = 39.7 bits (91), Expect = 0.001
Identities = 19/36 (52%), Positives = 23/36 (63%)
Frame = +2
Query: 167 GIPGVLFVLPDSYVDPEHQDYGAELFVNGEIVQRSP 202
G P LF L DS+VDP+++ YGA FVNG R P
Sbjct: 2 GSPEALFGLSDSFVDPKNKVYGAYFFVNGADTFRDP 109
>TC226456
Length = 788
Score = 38.9 bits (89), Expect = 0.002
Identities = 20/53 (37%), Positives = 30/53 (55%)
Frame = +3
Query: 124 YVKTLAQVLGSEEEAKKKIYNVSCERYFGFGCELDEETSNKLEGIPGVLFVLP 176
+++TL VLGSEE AK+ + GF +L E ++ +PGVL V+P
Sbjct: 309 HIRTLTSVLGSEEAAKEALLYSYKSAASGFSAKLTPEQVEQISKLPGVLQVVP 467
>TC214303 similar to UP|Q9T0K5 (Q9T0K5) Extensin-like protein, partial
(18%)
Length = 1014
Score = 38.1 bits (87), Expect = 0.003
Identities = 24/68 (35%), Positives = 38/68 (55%), Gaps = 12/68 (17%)
Frame = +1
Query: 2 TTRAIASSFKRRLFSTTTSTSTSTSTYTTTVAGALS-------KPSSLTSPRFII----- 49
TT A+ +F+TTT+T +ST+T+TTT +LS SSL+SP +
Sbjct: 13 TTSVFAACTASNVFATTTATVSSTTTFTTTTTTSLSLFITATTSSSSLSSPTSLFTTTAT 192
Query: 50 PMSQTIFQ 57
P+S T+++
Sbjct: 193PISTTLYR 216
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.317 0.134 0.399
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 10,170,537
Number of Sequences: 63676
Number of extensions: 148548
Number of successful extensions: 1344
Number of sequences better than 10.0: 79
Number of HSP's better than 10.0 without gapping: 1189
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1291
length of query: 235
length of database: 12,639,632
effective HSP length: 94
effective length of query: 141
effective length of database: 6,654,088
effective search space: 938226408
effective search space used: 938226408
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.7 bits)
S2: 57 (26.6 bits)
Medicago: description of AC145202.10


