

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145157.11 + phase: 0
(74 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
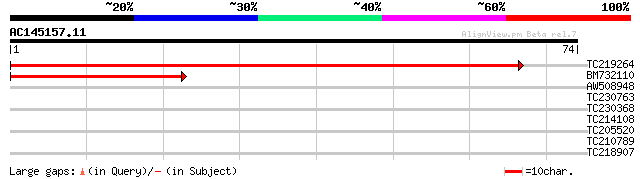
Score E
Sequences producing significant alignments: (bits) Value
TC219264 similar to GB|AAO63372.1|28950897|BT005308 At1g30825 {A... 128 5e-31
BM732110 44 2e-05
AW508948 similar to GP|13605793|gb| At1g50300/F14I3_23 {Arabidop... 27 2.5
TC230763 26 3.3
TC230368 26 4.3
TC214108 similar to UP|RK29_ARATH (Q9FJP3) 50S ribosomal protein... 26 4.3
TC205520 ferredoxin:sulfite reductase precursor [Glycine max] 26 4.3
TC210789 similar to UP|Q9SFE9 (Q9SFE9) T26F17.9 (GONST5 Golgi Nu... 25 5.7
TC218907 weakly similar to UP|Q94AW2 (Q94AW2) AT5g39590/MIJ24_60... 25 9.7
>TC219264 similar to GB|AAO63372.1|28950897|BT005308 At1g30825 {Arabidopsis
thaliana;} , partial (43%)
Length = 615
Score = 128 bits (322), Expect = 5e-31
Identities = 62/67 (92%), Positives = 66/67 (97%)
Frame = +1
Query: 1 MKNPNIFLLSVSLPTPSSETIFVCGLPFGAIEAIKAAYGSLVQILDPPRDGFNLTLKINL 60
MKNP++ LLSVSLPTPSSETIFVCGLPFGAIEAIKAAYG+LVQILDPPRDGFNLTLKINL
Sbjct: 307 MKNPHVLLLSVSLPTPSSETIFVCGLPFGAIEAIKAAYGNLVQILDPPRDGFNLTLKINL 486
Query: 61 SKVPANQ 67
SK+PANQ
Sbjct: 487 SKLPANQ 507
>BM732110
Length = 421
Score = 43.9 bits (102), Expect = 2e-05
Identities = 20/23 (86%), Positives = 22/23 (94%)
Frame = +2
Query: 1 MKNPNIFLLSVSLPTPSSETIFV 23
MKNP++ LLSVSLPTPSSETIFV
Sbjct: 353 MKNPHVLLLSVSLPTPSSETIFV 421
>AW508948 similar to GP|13605793|gb| At1g50300/F14I3_23 {Arabidopsis
thaliana}, partial (24%)
Length = 439
Score = 26.6 bits (57), Expect = 2.5
Identities = 9/30 (30%), Positives = 19/30 (63%)
Frame = +3
Query: 16 PSSETIFVCGLPFGAIEAIKAAYGSLVQIL 45
PS+ +++VC LP+G + + A Y + ++
Sbjct: 168 PSNGSVYVCNLPYGTDDNMLAEYFGTIGLI 257
>TC230763
Length = 1058
Score = 26.2 bits (56), Expect = 3.3
Identities = 12/26 (46%), Positives = 16/26 (61%), Gaps = 2/26 (7%)
Frame = -1
Query: 4 PNIFLL--SVSLPTPSSETIFVCGLP 27
PN+FLL + + PTPS F+C P
Sbjct: 554 PNLFLL*DAFADPTPSFRFFFICAAP 477
>TC230368
Length = 895
Score = 25.8 bits (55), Expect = 4.3
Identities = 9/15 (60%), Positives = 12/15 (80%)
Frame = +3
Query: 13 LPTPSSETIFVCGLP 27
LP+PSS T ++C LP
Sbjct: 384 LPSPSSGTTYICSLP 428
>TC214108 similar to UP|RK29_ARATH (Q9FJP3) 50S ribosomal protein L29,
chloroplast precursor, partial (68%)
Length = 1202
Score = 25.8 bits (55), Expect = 4.3
Identities = 13/34 (38%), Positives = 21/34 (61%)
Frame = +1
Query: 8 LLSVSLPTPSSETIFVCGLPFGAIEAIKAAYGSL 41
+LS+S+ +PSS TI PF I+ ++A S+
Sbjct: 85 MLSLSVASPSSSTISFLPKPFNGIQLRRSATCSI 186
>TC205520 ferredoxin:sulfite reductase precursor [Glycine max]
Length = 2633
Score = 25.8 bits (55), Expect = 4.3
Identities = 10/23 (43%), Positives = 14/23 (60%)
Frame = -2
Query: 2 KNPNIFLLSVSLPTPSSETIFVC 24
KNPNI L+ LP ++ +F C
Sbjct: 2281 KNPNISLVEPELPHSNNRVVFSC 2213
>TC210789 similar to UP|Q9SFE9 (Q9SFE9) T26F17.9 (GONST5 Golgi Nucleotide
sugar transporter), partial (33%)
Length = 502
Score = 25.4 bits (54), Expect = 5.7
Identities = 9/22 (40%), Positives = 16/22 (71%)
Frame = -3
Query: 5 NIFLLSVSLPTPSSETIFVCGL 26
N++ + +SLP ++ET F+C L
Sbjct: 473 NLYTI*LSLPLQTAETSFICSL 408
>TC218907 weakly similar to UP|Q94AW2 (Q94AW2) AT5g39590/MIJ24_60, partial
(40%)
Length = 991
Score = 24.6 bits (52), Expect = 9.7
Identities = 11/25 (44%), Positives = 16/25 (64%)
Frame = -3
Query: 4 PNIFLLSVSLPTPSSETIFVCGLPF 28
P + LLS++L + E I + GLPF
Sbjct: 878 PRLCLLSIALSNATLEMIQLSGLPF 804
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.321 0.141 0.409
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,345,141
Number of Sequences: 63676
Number of extensions: 39544
Number of successful extensions: 181
Number of sequences better than 10.0: 18
Number of HSP's better than 10.0 without gapping: 181
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 181
length of query: 74
length of database: 12,639,632
effective HSP length: 50
effective length of query: 24
effective length of database: 9,455,832
effective search space: 226939968
effective search space used: 226939968
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.4 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.8 bits)
S2: 52 (24.6 bits)
Medicago: description of AC145157.11


