

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC145024.2 - phase: 0
(355 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
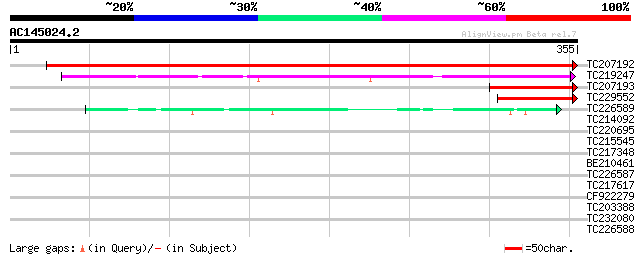
Score E
Sequences producing significant alignments: (bits) Value
TC207192 533 e-152
TC219247 134 8e-32
TC207193 106 2e-23
TC229552 homologue to UP|Q9SGQ0 (Q9SGQ0) F3M18.4 (Transcription ... 96 3e-20
TC226589 42 6e-04
TC214092 similar to UP|Q9FJY6 (Q9FJY6) Apospory-associated prote... 40 0.002
TC220695 weakly similar to UP|Q8W1C6 (Q8W1C6) Ring-H2 zinc finge... 33 0.20
TC215545 homologue to GB|AAC49174.1|849136|VRU26709 vacuolar H+-... 31 0.75
TC217348 similar to UP|Q8W436 (Q8W436) PBng110, complete 30 1.3
BE210461 weakly similar to GP|12054507|em ribosomal protein L2 {... 30 1.3
TC226587 30 2.2
TC217617 similar to UP|Q9LYL2 (Q9LYL2) Alpha-galactosidase-like ... 29 3.7
CF922279 29 3.7
TC203388 homologue to UP|H2B_GOSHI (O22582) Histone H2B, complete 28 4.8
TC232080 homologue to UP|O64851 (O64851) Expressed protein (At2g... 28 4.8
TC226588 28 4.8
>TC207192
Length = 1307
Score = 533 bits (1372), Expect = e-152
Identities = 261/332 (78%), Positives = 290/332 (86%)
Frame = +3
Query: 24 TTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKD 83
T PL T E LN+KFGRKGIKFLES++ PIV+LTVRNGSSL LR+PDAHVTSYKPKV WKD
Sbjct: 27 TRPLPTAETLNEKFGRKGIKFLESDNTPIVDLTVRNGSSLRLRIPDAHVTSYKPKVNWKD 206
Query: 84 DGLEELLYTIPANETGLYKAKGGIGLVLNEVLQPGAKELLPSTLEWTVKDVDYDAIDALQ 143
DG +E+LYTIPA ETG YKAKGG+GLV+NEVLQPGAK LLPSTLEWTV DVD D+IDALQ
Sbjct: 207 DGFQEVLYTIPATETGPYKAKGGVGLVMNEVLQPGAKGLLPSTLEWTVNDVDSDSIDALQ 386
Query: 144 VELISTNRFFDMTYIVSLYPVSMATAVIVKNKSPKPVTLTNAILSHFRFKRRGGAAIKGL 203
VEL T+RFFD+TYIV+LYPVSMATAV+ KN PKP TLTNAILSHFRFK R G AIKGL
Sbjct: 387 VELSCTSRFFDITYIVTLYPVSMATAVVAKNIGPKPATLTNAILSHFRFKNRRGTAIKGL 566
Query: 204 QTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPEMKPGSWTQQGVPITLLENKMS 263
++CSY H PL SPFQILTPSEA SE R +SFG E E KPG+W QQ + ITLLENKMS
Sbjct: 567 RSCSYIPHAPLSSPFQILTPSEATISEPPRWLSFGNETEAKPGTWGQQALSITLLENKMS 746
Query: 264 RVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSMSDKYGRDYF 323
RV+AAPPKER KAFYNTPPSKYE IDQGR + +RVIRMGFEDIY+SSPGS+S+KYG+DYF
Sbjct: 747 RVYAAPPKERLKAFYNTPPSKYETIDQGRGLCFRVIRMGFEDIYLSSPGSLSEKYGKDYF 926
Query: 324 VCTGPASILEPVTVNPGEEFRGAQVIEHDNLS 355
+CTGPASIL PVTVNPGEE+RGAQVIEHDNL+
Sbjct: 927 ICTGPASILVPVTVNPGEEWRGAQVIEHDNLT 1022
>TC219247
Length = 1272
Score = 134 bits (336), Expect = 8e-32
Identities = 95/329 (28%), Positives = 155/329 (46%), Gaps = 7/329 (2%)
Frame = +1
Query: 33 LNDKFGRKGIKFLESNSIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYT 92
L ++F G+ F I ++ ++NGS +++ LP +TSYK + W LE L
Sbjct: 157 LQEEFSGHGVTFEGVEDSCIAKMELKNGSIVTMMLPSGLITSYKAPM-WHGGKLELLHTN 333
Query: 93 IPANETGLYKAKGGIGLVLNEVLQPGAKELLPSTLEWTVKDVDYDAIDALQVELISTNRF 152
+ E G +GG+ L N G E S W + + +A +++QVEL TNR
Sbjct: 334 VSEGEYGDAIIQGGVSLNFNFQTDDG--EFSWSPTNWVLHKIQGNANESIQVEL--TNRT 501
Query: 153 FD----MTYIVSLYPVSMATAVIVKNKSPKPVTLTNAILSHFRFKRRGGAAIKGLQTCSY 208
D + YIV+L ++ + V + NK PV +T +ILSH GL+ +Y
Sbjct: 502 SDDKIGLKYIVTLEKDALNSEVEISNKKSLPVQMTGSILSHLTVSSPEATYALGLERSNY 681
Query: 209 CSHPPLDSPFQILTPS---EAMKSESQRLISFGAEPEMKPGSWTQQGVPITLLENKMSRV 265
CS P +S F + P E ++L + GS Q + + ++
Sbjct: 682 CSKPLFESEFMLSPPDGQEEGFGKIVEQLFPQWGTKDQNNGSEGSQSEEMDEEIDNYKQL 861
Query: 266 FAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSMSDKYGRDYFVC 325
E+ Y P + +ID+GR V R GF+++Y+ SPGS + Y + ++C
Sbjct: 862 -----SEKLSHVYTDVPRNFTVIDRGRRNSVSVGRTGFDEMYLFSPGSRVEIYSKYSYIC 1026
Query: 326 TGPASILEPVTVNPGEEFRGAQVIEHDNL 354
G A+IL+P+ ++P + +RG Q I + NL
Sbjct: 1027VGQAAILKPIILSPEDVWRGGQYIHNPNL 1113
>TC207193
Length = 484
Score = 106 bits (264), Expect = 2e-23
Identities = 47/55 (85%), Positives = 54/55 (97%)
Frame = -3
Query: 301 MGFEDIYVSSPGSMSDKYGRDYFVCTGPASILEPVTVNPGEEFRGAQVIEHDNLS 355
MGFEDIY+SSPGS+S+KYG+DYF+CTGPASIL PVTVNPGEE+RGAQVIEHDNL+
Sbjct: 482 MGFEDIYLSSPGSLSEKYGKDYFICTGPASILVPVTVNPGEEWRGAQVIEHDNLT 318
>TC229552 homologue to UP|Q9SGQ0 (Q9SGQ0) F3M18.4 (Transcription factor
AtVOZ1), partial (27%)
Length = 1520
Score = 95.5 bits (236), Expect = 3e-20
Identities = 42/50 (84%), Positives = 49/50 (98%)
Frame = +2
Query: 306 IYVSSPGSMSDKYGRDYFVCTGPASILEPVTVNPGEEFRGAQVIEHDNLS 355
IY+SSPGS+S+KYG+DYF+CTGPASIL PVTVNPGEE+RGAQVIEHDNL+
Sbjct: 1082 IYLSSPGSLSEKYGKDYFICTGPASILVPVTVNPGEEWRGAQVIEHDNLT 1231
>TC226589
Length = 1426
Score = 41.6 bits (96), Expect = 6e-04
Identities = 70/311 (22%), Positives = 119/311 (37%), Gaps = 13/311 (4%)
Frame = +2
Query: 48 NSIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLYTIPANETGLYKAKGGI 107
N + V L GSS + L AHVTS WK+D EELL+ +++
Sbjct: 77 NGLDKVILRDARGSSAEVYLYGAHVTS------WKNDHAEELLFL--SSKFSNLGPLDSH 232
Query: 108 GLVLNE--VLQPGAKELLPSTLEWTVKDVDYDAIDALQVELISTNRFFDMTYIVSLYP-- 163
G N + L +T D+ + + ++ F+ V+L
Sbjct: 233 GFARNRFWTIDDSPPPFLTNTPSKAFVDL---ILKPSEDDIKIWPHSFEFRLRVALGSGG 403
Query: 164 -VSMATAVIVKNKSPKPVTLTNAILSHFRFKRRGGAAIKGLQTCSYCSHPPLDSPFQILT 222
+ M + + N KP + T A ++F ++GL+T Y +
Sbjct: 404 DLMMTSRIRNTNIDGKPFSFTFANHTYFSVSDISEVRVEGLETLDYLDNL---------- 553
Query: 223 PSEAMKSESQRLISFGAEPEMKPGSWTQQGVPITLLENKMSRVFAAPPKERTKAFYNTPP 282
K +T+QG +T E++ R+ Y + P
Sbjct: 554 --------------------QKRERFTEQGDALTF-ESEFDRI------------YLSTP 634
Query: 283 SKYEIIDQGREIFYRVIRMGFEDIYVSSPG-----SMSDKYGRD---YFVCTGPASILEP 334
+K I+D ++ + + G D V +P +MSD +G D Y +C A+I +P
Sbjct: 635 TKIAILDHEKKRTIVLRKDGLPDAVVWNPWDKKAKAMSD-FGDDEYKYMLCVEAAAIEKP 811
Query: 335 VTVNPGEEFRG 345
+T+ PGEE++G
Sbjct: 812 ITLKPGEEWKG 844
>TC214092 similar to UP|Q9FJY6 (Q9FJY6) Apospory-associated protein C-like
protein, partial (89%)
Length = 1287
Score = 39.7 bits (91), Expect = 0.002
Identities = 52/218 (23%), Positives = 92/218 (41%), Gaps = 12/218 (5%)
Frame = +1
Query: 8 SLPKLNLIKASSAATTTTPLSTPEALNDKFGRKGIKFLESN-SIPIVELTVRNGSSLSLR 66
+ P LN + + A+ + ST G++ E ++P + LT GS +
Sbjct: 199 TFPNLNRCRGVAYASLSNKASTTSL--------GVRVTEGEGNLPKLVLTSPAGSEAEIY 354
Query: 67 LPDAHVTSYKPKVFWKDDGLEELLYTIP-ANETGLYKAKGGIGLVLNEVLQPGAKEL--L 123
L +TS WK ++LL+ P A G GG+ + PG +
Sbjct: 355 LFGGCITS------WKVPSGKDLLFVRPDAVFNGNKPISGGVPHCFPQ-FGPGPIQQHGF 513
Query: 124 PSTLEWTVKD---VDYDAIDALQVELISTNRF-----FDMTYIVSLYPVSMATAVIVKNK 175
++WTV D + + + L+++ +R F + V+L S+AT + VKN
Sbjct: 514 ARNMDWTVVDSENTEGNPVVTLELKDAPYSRAMWDFSFHALFKVTLNAKSLATELTVKNT 693
Query: 176 SPKPVTLTNAILSHFRFKRRGGAAIKGLQTCSYCSHPP 213
K + + A+ ++FR A++KGL+ C + P
Sbjct: 694 DNKAFSFSTALHTYFR-ASASNASVKGLKGCKTLNKDP 804
>TC220695 weakly similar to UP|Q8W1C6 (Q8W1C6) Ring-H2 zinc finger protein,
partial (15%)
Length = 556
Score = 33.1 bits (74), Expect = 0.20
Identities = 31/106 (29%), Positives = 45/106 (42%), Gaps = 3/106 (2%)
Frame = +3
Query: 179 PVTLTNAILSHFRFKRRGGA-AIKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISF 237
P+ T +IL +RF R A A TC++ PP +P PS + ES L S
Sbjct: 183 PLNWTCSILCFWRFSCRASA*APFSSSTCAFSGTPPTTTPTPRYRPSLSPTPESP-LPSS 359
Query: 238 GAEPEM--KPGSWTQQGVPITLLENKMSRVFAAPPKERTKAFYNTP 281
+ PE + SW + P ++ S F P T + +N P
Sbjct: 360 TSSPESPERTFSW-ETNAPSAWTRSEQSNRFGWFPVATTPSIWNAP 494
>TC215545 homologue to GB|AAC49174.1|849136|VRU26709 vacuolar H+-ATPase
subunit A {Vigna radiata;} , complete
Length = 2358
Score = 31.2 bits (69), Expect = 0.75
Identities = 22/76 (28%), Positives = 34/76 (43%), Gaps = 4/76 (5%)
Frame = +1
Query: 256 TLLENK-MSRVFAAPPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIY---VSSP 311
T+ EN M A PP K Y PP +Y I D E+ ++ ++ F + V +P
Sbjct: 523 TVFENSLMQHHIALPPDNMGKITYIAPPGQYSIKDTVLELEFQGVKKKFTMLQTWPVRTP 702
Query: 312 GSMSDKYGRDYFVCTG 327
++ K D + TG
Sbjct: 703 RPVASKLAADTPLLTG 750
>TC217348 similar to UP|Q8W436 (Q8W436) PBng110, complete
Length = 1456
Score = 30.4 bits (67), Expect = 1.3
Identities = 19/79 (24%), Positives = 34/79 (42%), Gaps = 1/79 (1%)
Frame = +3
Query: 202 GLQTCSYCSHPPLDSPFQIL-TPSEAMKSESQRLISFGAEPEMKPGSWTQQGVPITLLEN 260
G TC+ SHPP PFQ L +P +S ++ + G + ++ + + + +
Sbjct: 57 GFYTCNLLSHPPTLQPFQFLPSPKPHYRSSTRSCLLSGLDKRIEA*NGLRSTSTSGFVCD 236
Query: 261 KMSRVFAAPPKERTKAFYN 279
FA K +A +N
Sbjct: 237 IFLEAFAMEQKAEVEAVFN 293
>BE210461 weakly similar to GP|12054507|em ribosomal protein L2 {Glycine
max}, partial (56%)
Length = 505
Score = 30.4 bits (67), Expect = 1.3
Identities = 21/76 (27%), Positives = 34/76 (44%)
Frame = +3
Query: 2 ASFLSLSLPKLNLIKASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRNGS 61
+S S S KASS ++TTP++TP + + +S + + N S
Sbjct: 81 SSTASTSTSTTTTSKASSPTSSTTPITTPLSPK*HSTTPSTTRXKQSSSSPPKTSTPNSS 260
Query: 62 SLSLRLPDAHVTSYKP 77
S + R P +T+Y P
Sbjct: 261 STTTRKPPCIITTYYP 308
>TC226587
Length = 566
Score = 29.6 bits (65), Expect = 2.2
Identities = 19/44 (43%), Positives = 23/44 (52%)
Frame = +3
Query: 48 NSIPIVELTVRNGSSLSLRLPDAHVTSYKPKVFWKDDGLEELLY 91
N + V L GSS + L AHVTS WK+D EELL+
Sbjct: 201 NGLDKVILRDARGSSAEVYLYGAHVTS------WKNDHAEELLF 314
>TC217617 similar to UP|Q9LYL2 (Q9LYL2) Alpha-galactosidase-like protein,
partial (89%)
Length = 1691
Score = 28.9 bits (63), Expect = 3.7
Identities = 18/54 (33%), Positives = 25/54 (45%)
Frame = +3
Query: 269 PPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFEDIYVSSPGSMSDKYGRDY 322
PPKER PP + + G++IFY + G ED P +DK G +
Sbjct: 717 PPKERY------PPMRDALNATGQKIFYSLCEWGVED-----PALWADKVGNSW 845
>CF922279
Length = 528
Score = 28.9 bits (63), Expect = 3.7
Identities = 14/36 (38%), Positives = 21/36 (57%)
Frame = +3
Query: 269 PPKERTKAFYNTPPSKYEIIDQGREIFYRVIRMGFE 304
PP E+TK F+ TPP +G++ FYR + F+
Sbjct: 165 PPPEKTKNFFKTPPPG---APRGQKSFYRPPKRVFQ 263
>TC203388 homologue to UP|H2B_GOSHI (O22582) Histone H2B, complete
Length = 675
Score = 28.5 bits (62), Expect = 4.8
Identities = 22/81 (27%), Positives = 35/81 (43%)
Frame = -3
Query: 131 VKDVDYDAIDALQVELISTNRFFDMTYIVSLYPVSMATAVIVKNKSPKPVTLTNAILSHF 190
++DVD + AL V L+ L P S +++ K S + L +++SH
Sbjct: 283 LEDVDLVCLHALLVSLL-------------LLPFSAGASLLRKLLSGLWLLLRRSLVSHR 143
Query: 191 RFKRRGGAAIKGLQTCSYCSH 211
RF G + L C +C H
Sbjct: 142 RFLLLRGLLLCWLLLCLWCHH 80
>TC232080 homologue to UP|O64851 (O64851) Expressed protein
(At2g26190/T1D16.17), partial (8%)
Length = 745
Score = 28.5 bits (62), Expect = 4.8
Identities = 16/58 (27%), Positives = 31/58 (52%)
Frame = -2
Query: 187 LSHFRFKRRGGAAIKGLQTCSYCSHPPLDSPFQILTPSEAMKSESQRLISFGAEPEMK 244
L H RF R GLQ+ + +++PF++++ S +K ++I+ EP++K
Sbjct: 534 LMHERFGSR--TMCSGLQSVNLLEKLVVEAPFEVVSDSLILKLPDLKIIAPSLEPKVK 367
>TC226588
Length = 501
Score = 28.5 bits (62), Expect = 4.8
Identities = 26/76 (34%), Positives = 37/76 (48%)
Frame = +2
Query: 16 KASSAATTTTPLSTPEALNDKFGRKGIKFLESNSIPIVELTVRNGSSLSLRLPDAHVTSY 75
K+ S+A+ + S+ L+ KGI N + V L GSS + L AHVTS
Sbjct: 164 KSPSSASASASSSSSYELS-----KGI-----NGLDKVILRDPRGSSAEVYLYGAHVTS- 310
Query: 76 KPKVFWKDDGLEELLY 91
WK++ EELL+
Sbjct: 311 -----WKNEQAEELLF 343
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.316 0.134 0.388
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 14,922,491
Number of Sequences: 63676
Number of extensions: 198901
Number of successful extensions: 1062
Number of sequences better than 10.0: 32
Number of HSP's better than 10.0 without gapping: 1053
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 1058
length of query: 355
length of database: 12,639,632
effective HSP length: 98
effective length of query: 257
effective length of database: 6,399,384
effective search space: 1644641688
effective search space used: 1644641688
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 16 ( 7.3 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (21.6 bits)
S2: 59 (27.3 bits)
Medicago: description of AC145024.2


