

BLAST2 result
TBLASTN 2.2.2 [Dec-14-2001]
Reference: Altschul, Stephen F., Thomas L. Madden, Alejandro A. Schaffer,
Jinghui Zhang, Zheng Zhang, Webb Miller, and David J. Lipman (1997),
"Gapped BLAST and PSI-BLAST: a new generation of protein database search
programs", Nucleic Acids Res. 25:3389-3402.
Query= AC144931.14 - phase: 0 /pseudo
(122 letters)
Database: GMGI
63,676 sequences; 37,918,896 total letters
Searching..................................................done
Score E
Sequences producing significant alignments: (bits) Value
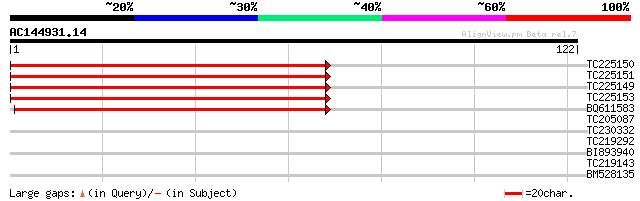
TC225150 homologue to UP|Q9M5L1 (Q9M5L1) 40S ribosomal protein S... 136 1e-33
TC225151 homologue to UP|Q9M5L1 (Q9M5L1) 40S ribosomal protein S... 136 1e-33
TC225149 homologue to UP|Q9M5L1 (Q9M5L1) 40S ribosomal protein S... 132 2e-32
TC225153 homologue to UP|Q9M5L1 (Q9M5L1) 40S ribosomal protein S... 132 2e-32
BQ611583 weakly similar to GP|6984222|gb| 40S ribosomal protein ... 92 3e-20
TC205087 weakly similar to UP|Q8LB56 (Q8LB56) Nuclear RNA bindin... 27 1.0
TC230332 similar to UP|Q9FIL5 (Q9FIL5) Gb|AAB82637.1 (AT5g58960/... 25 5.0
TC219292 similar to UP|Q8S0V9 (Q8S0V9) DnaJ-like protein, partia... 25 5.0
BI893940 25 6.5
TC219143 similar to UP|Q9FJJ8 (Q9FJJ8) Arabidopsis thaliana geno... 25 6.5
BM528135 weakly similar to GP|15408766|dbj NPK1-related protein ... 24 8.5
>TC225150 homologue to UP|Q9M5L1 (Q9M5L1) 40S ribosomal protein S16
(Fragment), partial (97%)
Length = 910
Score = 136 bits (343), Expect = 1e-33
Identities = 65/69 (94%), Positives = 68/69 (98%)
Frame = +2
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
+EQVQCFGRKK AVAVT+CKRGRGLIKINGCPIELVEPEILRFKA+EPILLLGRHRFAGV
Sbjct: 125 IEQVQCFGRKKTAVAVTYCKRGRGLIKINGCPIELVEPEILRFKAFEPILLLGRHRFAGV 304
Query: 61 DMRIRVKGG 69
DMRIRVKGG
Sbjct: 305 DMRIRVKGG 331
>TC225151 homologue to UP|Q9M5L1 (Q9M5L1) 40S ribosomal protein S16
(Fragment), partial (97%)
Length = 710
Score = 136 bits (343), Expect = 1e-33
Identities = 65/69 (94%), Positives = 68/69 (98%)
Frame = +1
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
+EQVQCFGRKK AVAVT+CKRGRGLIKINGCPIELVEPEILRFKA+EPILLLGRHRFAGV
Sbjct: 118 IEQVQCFGRKKTAVAVTYCKRGRGLIKINGCPIELVEPEILRFKAFEPILLLGRHRFAGV 297
Query: 61 DMRIRVKGG 69
DMRIRVKGG
Sbjct: 298 DMRIRVKGG 324
>TC225149 homologue to UP|Q9M5L1 (Q9M5L1) 40S ribosomal protein S16
(Fragment), partial (97%)
Length = 768
Score = 132 bits (332), Expect = 2e-32
Identities = 63/69 (91%), Positives = 67/69 (96%)
Frame = +2
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
+EQVQCFGRKK AVAVT+CKRGRGLIKINGCPIELVEPEILRFKA+EPILLLG+ RFAGV
Sbjct: 110 LEQVQCFGRKKTAVAVTYCKRGRGLIKINGCPIELVEPEILRFKAFEPILLLGKSRFAGV 289
Query: 61 DMRIRVKGG 69
DMRIRVKGG
Sbjct: 290 DMRIRVKGG 316
>TC225153 homologue to UP|Q9M5L1 (Q9M5L1) 40S ribosomal protein S16
(Fragment), partial (97%)
Length = 778
Score = 132 bits (332), Expect = 2e-32
Identities = 63/69 (91%), Positives = 67/69 (96%)
Frame = +1
Query: 1 MEQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGV 60
+EQVQCFGRKK AVAVT+CKRGRGLIKINGCPIELVEPEILRFKA+EPILLLG+ RFAGV
Sbjct: 127 LEQVQCFGRKKTAVAVTYCKRGRGLIKINGCPIELVEPEILRFKAFEPILLLGKSRFAGV 306
Query: 61 DMRIRVKGG 69
DMRIRVKGG
Sbjct: 307 DMRIRVKGG 333
>BQ611583 weakly similar to GP|6984222|gb| 40S ribosomal protein S16
{Euphorbia esula}, partial (92%)
Length = 446
Score = 92.4 bits (228), Expect = 3e-20
Identities = 41/68 (60%), Positives = 53/68 (77%)
Frame = +2
Query: 2 EQVQCFGRKKNAVAVTHCKRGRGLIKINGCPIELVEPEILRFKAYEPILLLGRHRFAGVD 61
E VQ FGRKKNA+AV CK G GLI++NG PI+++EP LR K YEP+LLLG F+ +D
Sbjct: 41 EYVQTFGRKKNALAVATCKPGNGLIRVNGAPIDVLEPSTLRIKVYEPVLLLGAKTFSNLD 220
Query: 62 MRIRVKGG 69
+R+RV+GG
Sbjct: 221 IRVRVRGG 244
>TC205087 weakly similar to UP|Q8LB56 (Q8LB56) Nuclear RNA binding protein
A-like protein, partial (21%)
Length = 564
Score = 27.3 bits (59), Expect = 1.0
Identities = 10/14 (71%), Positives = 12/14 (85%)
Frame = +1
Query: 108 GSGGGKRYGRTRGG 121
G GGG+R+GR RGG
Sbjct: 502 GRGGGRRFGRGRGG 543
>TC230332 similar to UP|Q9FIL5 (Q9FIL5) Gb|AAB82637.1 (AT5g58960/k19m22_160),
partial (36%)
Length = 827
Score = 25.0 bits (53), Expect = 5.0
Identities = 9/14 (64%), Positives = 11/14 (78%)
Frame = +3
Query: 108 GSGGGKRYGRTRGG 121
G GGG++ GR RGG
Sbjct: 309 GGGGGRKKGRRRGG 350
>TC219292 similar to UP|Q8S0V9 (Q8S0V9) DnaJ-like protein, partial (37%)
Length = 901
Score = 25.0 bits (53), Expect = 5.0
Identities = 10/21 (47%), Positives = 13/21 (61%)
Frame = +3
Query: 2 EQVQCFGRKKNAVAVTHCKRG 22
E+V CFGR + + HCK G
Sbjct: 18 EEV*CFGRG*SDLRAAHCKAG 80
>BI893940
Length = 406
Score = 24.6 bits (52), Expect = 6.5
Identities = 16/45 (35%), Positives = 23/45 (50%), Gaps = 4/45 (8%)
Frame = +1
Query: 19 CKRGRGL---IKINGCPIEL-VEPEILRFKAYEPILLLGRHRFAG 59
C++ G IK+ PIE+ + EIL K + PI+ L H G
Sbjct: 172 CRKNTGALTAIKLVMLPIEVKYKSEILLIKRHLPIIFLNHHNQKG 306
>TC219143 similar to UP|Q9FJJ8 (Q9FJJ8) Arabidopsis thaliana genomic DNA,
chromosome 5, TAC clone:K19B1, partial (88%)
Length = 689
Score = 24.6 bits (52), Expect = 6.5
Identities = 9/14 (64%), Positives = 10/14 (71%)
Frame = +3
Query: 108 GSGGGKRYGRTRGG 121
G GGGK YG +GG
Sbjct: 456 GGGGGKHYGGGKGG 497
>BM528135 weakly similar to GP|15408766|dbj NPK1-related protein kinase-like
protein {Oryza sativa (japonica cultivar-group)},
partial (6%)
Length = 425
Score = 24.3 bits (51), Expect = 8.5
Identities = 10/15 (66%), Positives = 11/15 (72%)
Frame = +1
Query: 41 LRFKAYEPILLLGRH 55
LRF+ Y PILL RH
Sbjct: 115 LRFRTYLPILLFQRH 159
Database: GMGI
Posted date: Oct 22, 2004 4:58 PM
Number of letters in database: 37,918,896
Number of sequences in database: 63,676
Lambda K H
0.325 0.147 0.448
Gapped
Lambda K H
0.267 0.0410 0.140
Matrix: BLOSUM62
Gap Penalties: Existence: 11, Extension: 1
Number of Hits to DB: 3,820,736
Number of Sequences: 63676
Number of extensions: 36605
Number of successful extensions: 203
Number of sequences better than 10.0: 22
Number of HSP's better than 10.0 without gapping: 194
Number of HSP's successfully gapped in prelim test: 0
Number of HSP's that attempted gapping in prelim test: 0
Number of HSP's gapped (non-prelim): 203
length of query: 122
length of database: 12,639,632
effective HSP length: 98
effective length of query: 24
effective length of database: 6,399,384
effective search space: 153585216
effective search space used: 153585216
frameshift window, decay const: 50, 0.1
T: 13
A: 40
X1: 15 ( 7.0 bits)
X2: 38 (14.6 bits)
X3: 64 (24.7 bits)
S1: 41 (22.0 bits)
S2: 51 (24.3 bits)
Medicago: description of AC144931.14


